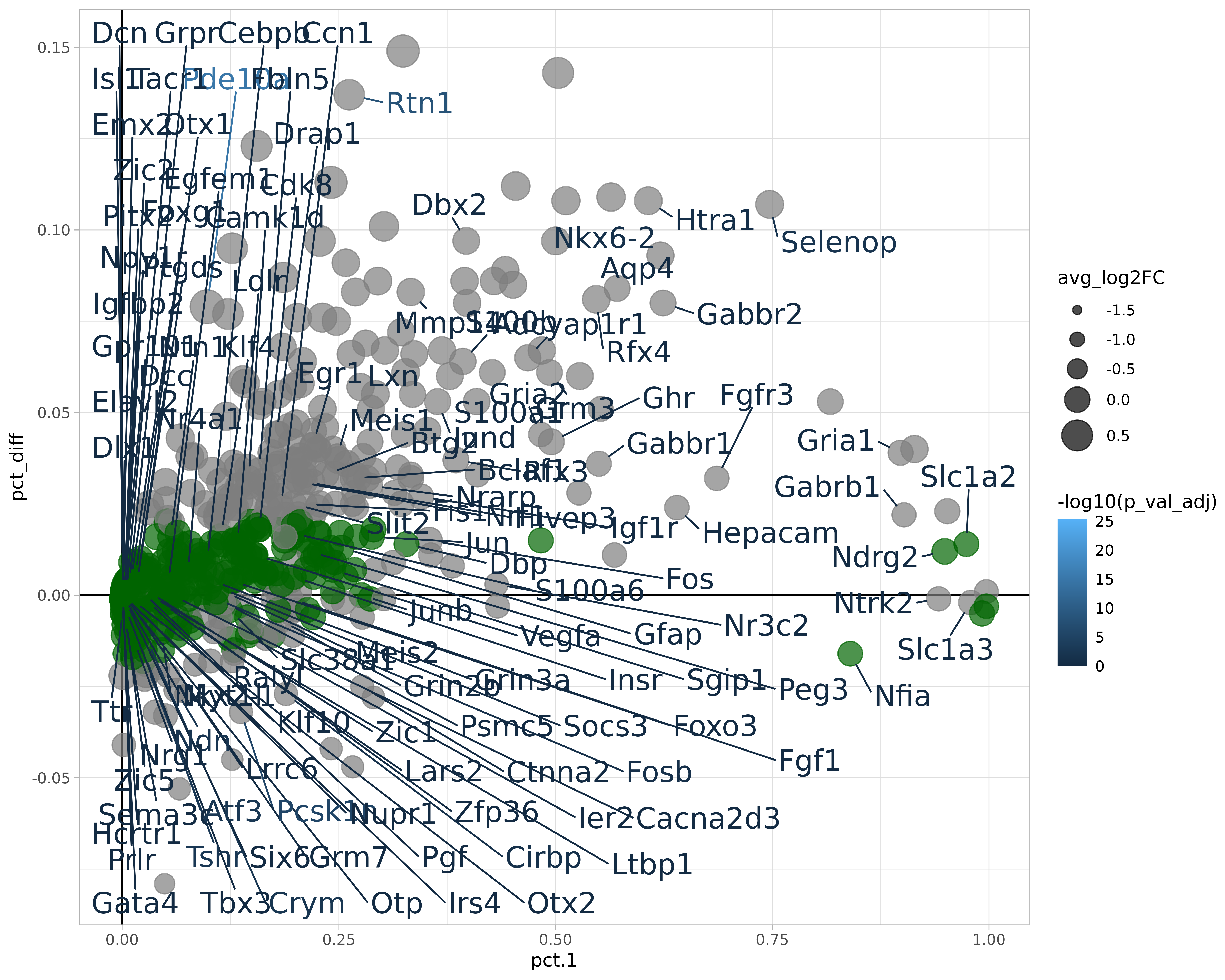
markers_region <- markers_region %>%
filter(pct.2 < 0.3, pct_diff > 0.1) %>%
group_by(cluster)
at_top_50 <- Extract_Top_Markers(marker_dataframe = markers_region, num_genes = 50, group_by = "cluster", gene_column = "gene", rank_by = "avg_log2FC", make_unique = TRUE, named_vector = FALSE)
markers_region2 <- markers_region %>%
filter(gene %in% c(nmr, npr, transcription_factors, mitochondrial)) %>%
group_by(cluster)
at_top_50s <- Extract_Top_Markers(marker_dataframe = markers_region2, num_genes = 50, group_by = "cluster", gene_column = "gene", rank_by = "avg_log2FC", make_unique = TRUE, named_vector = FALSE)
selected_genes <- c(
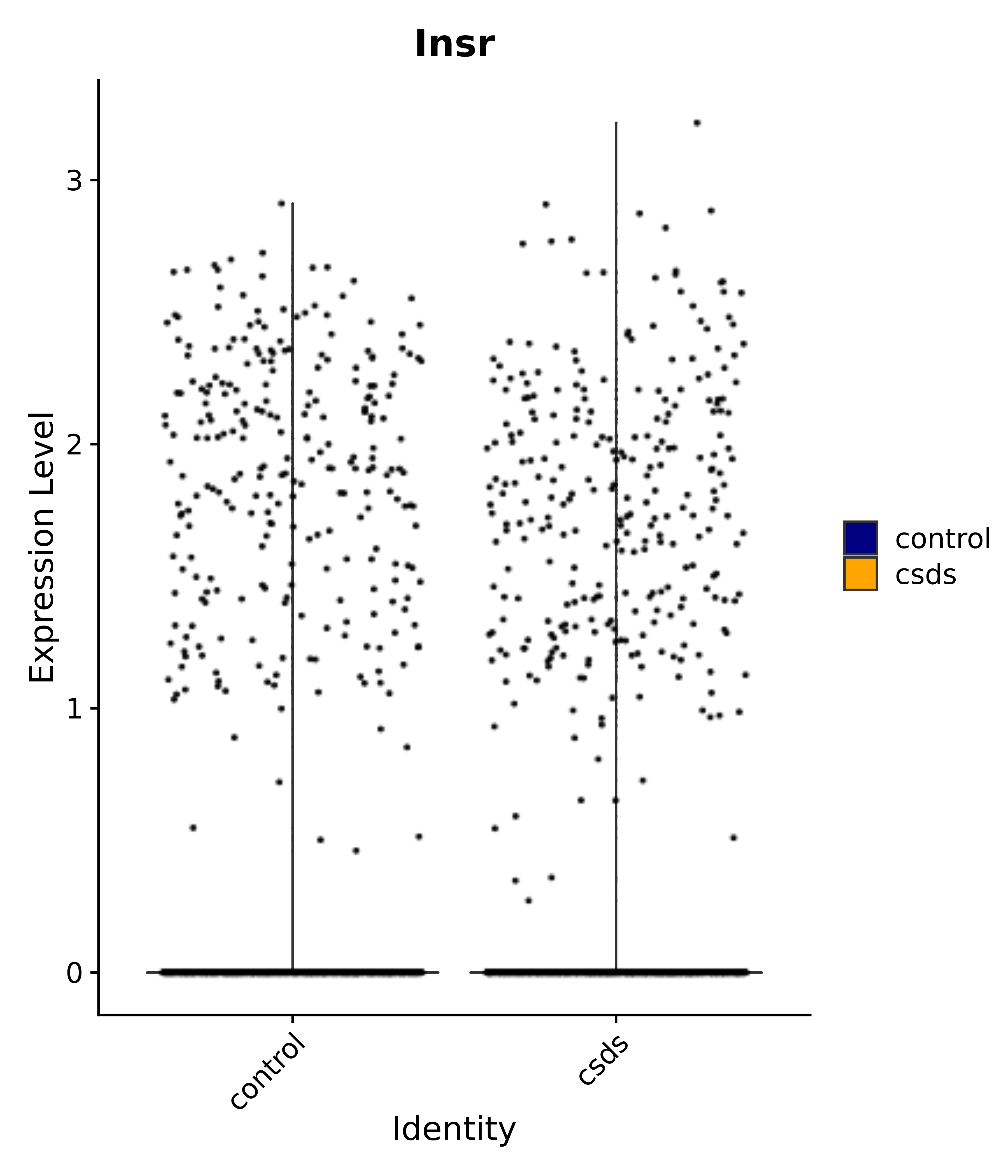
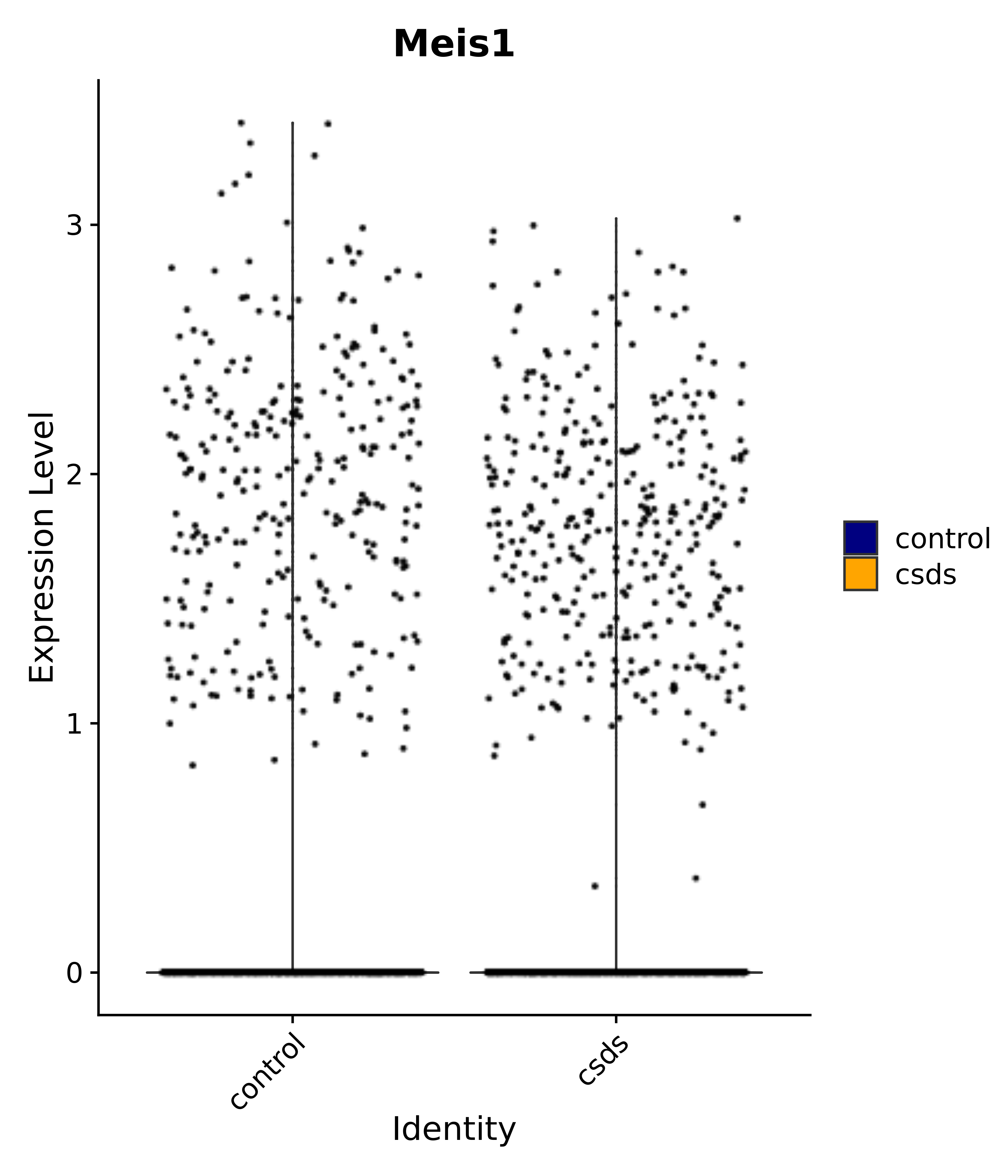
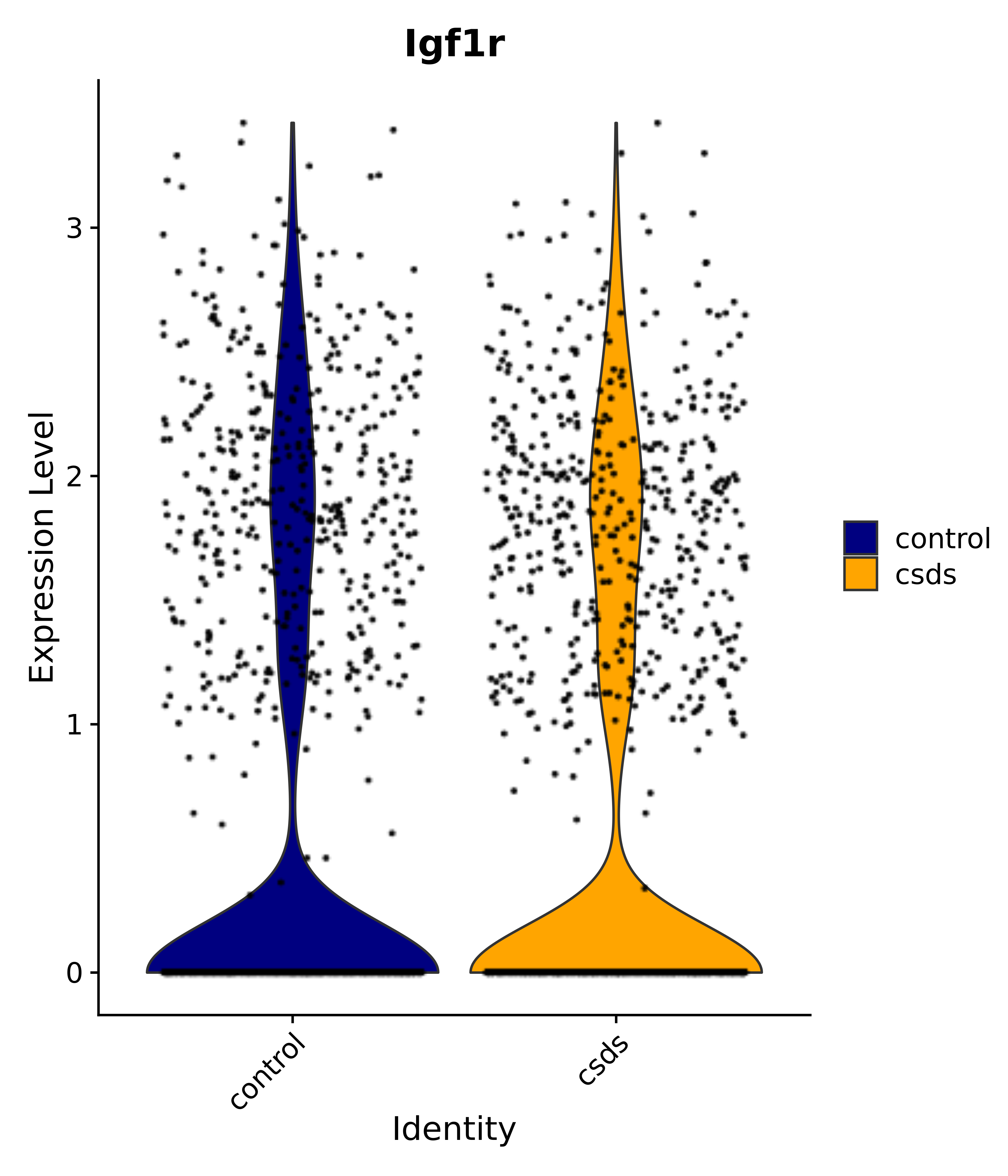
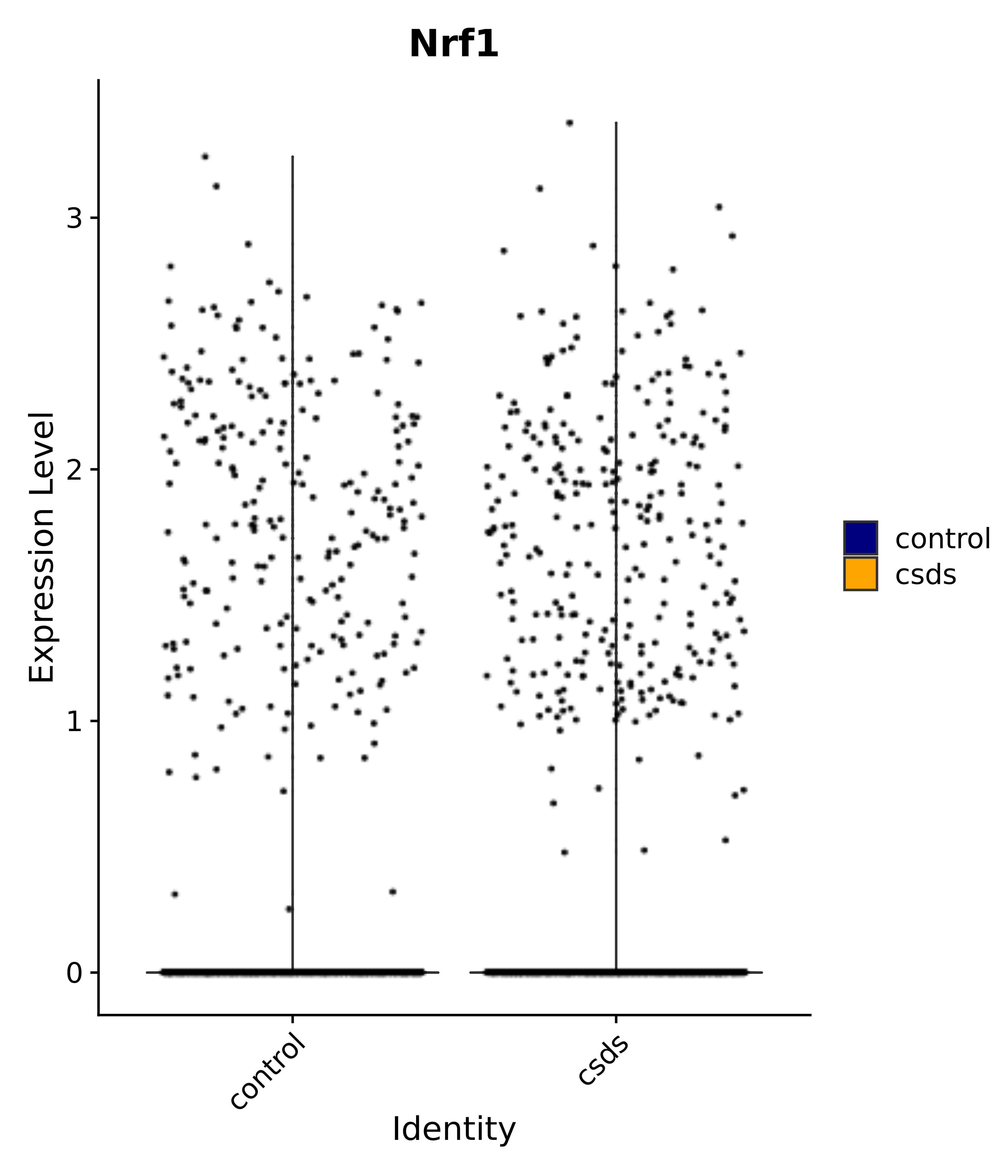
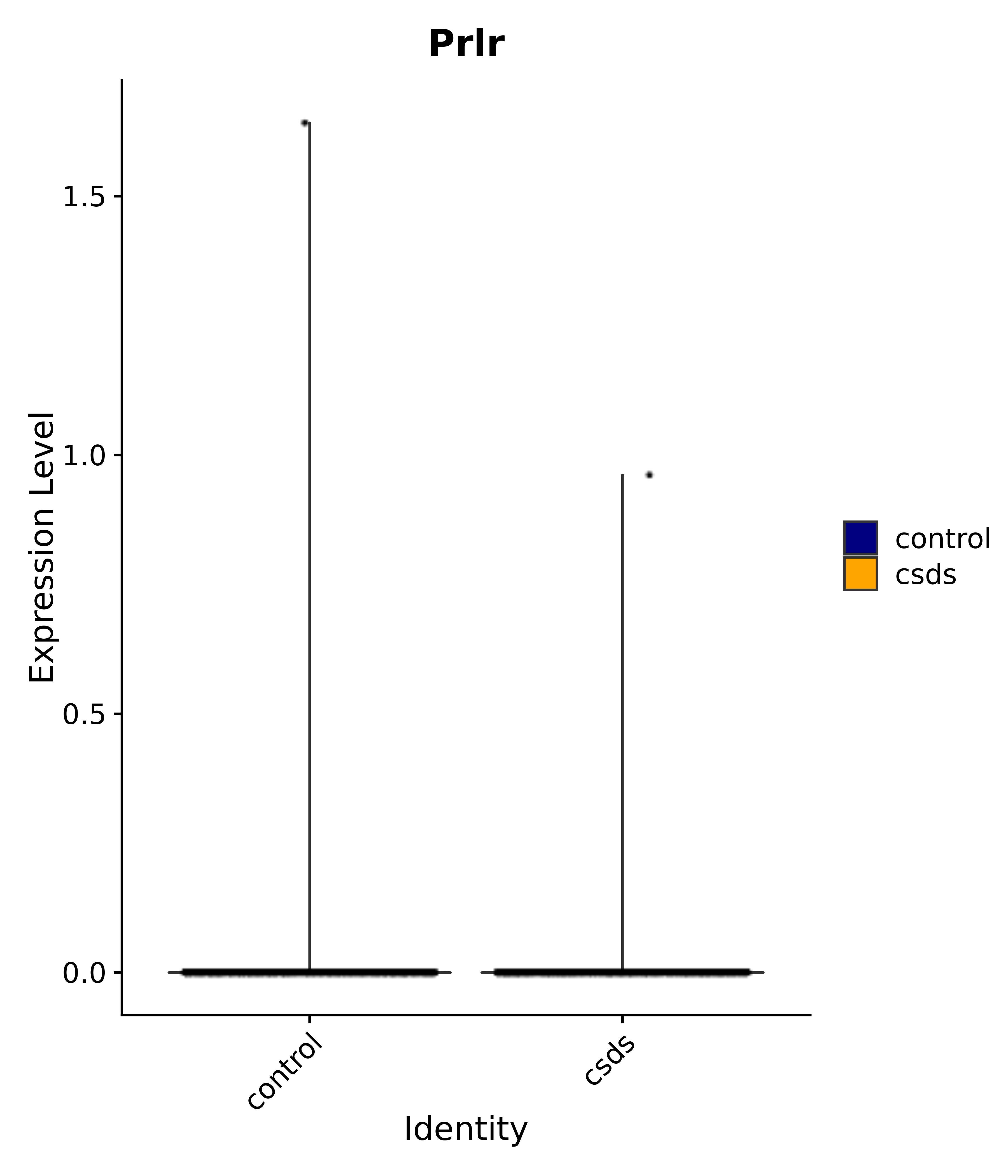
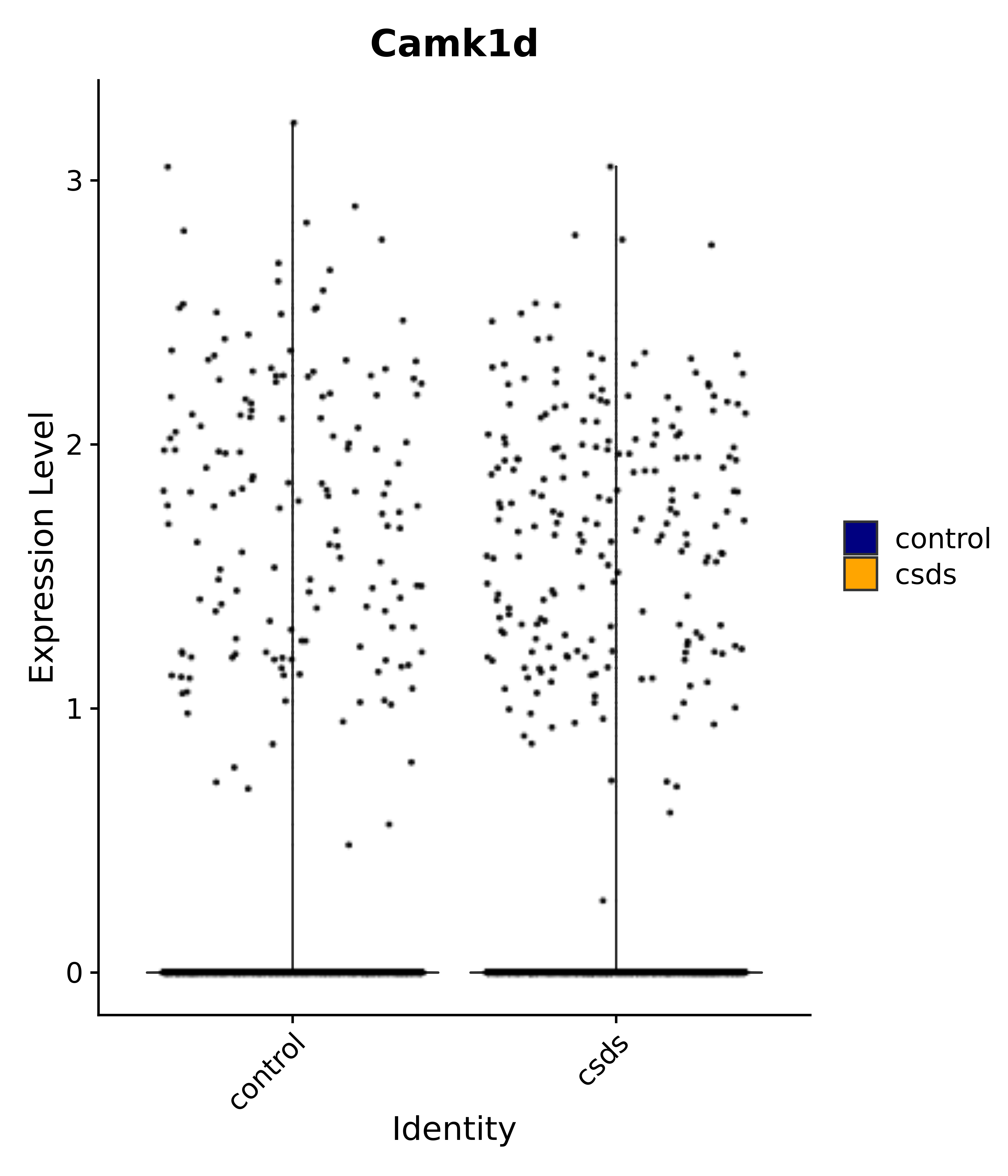
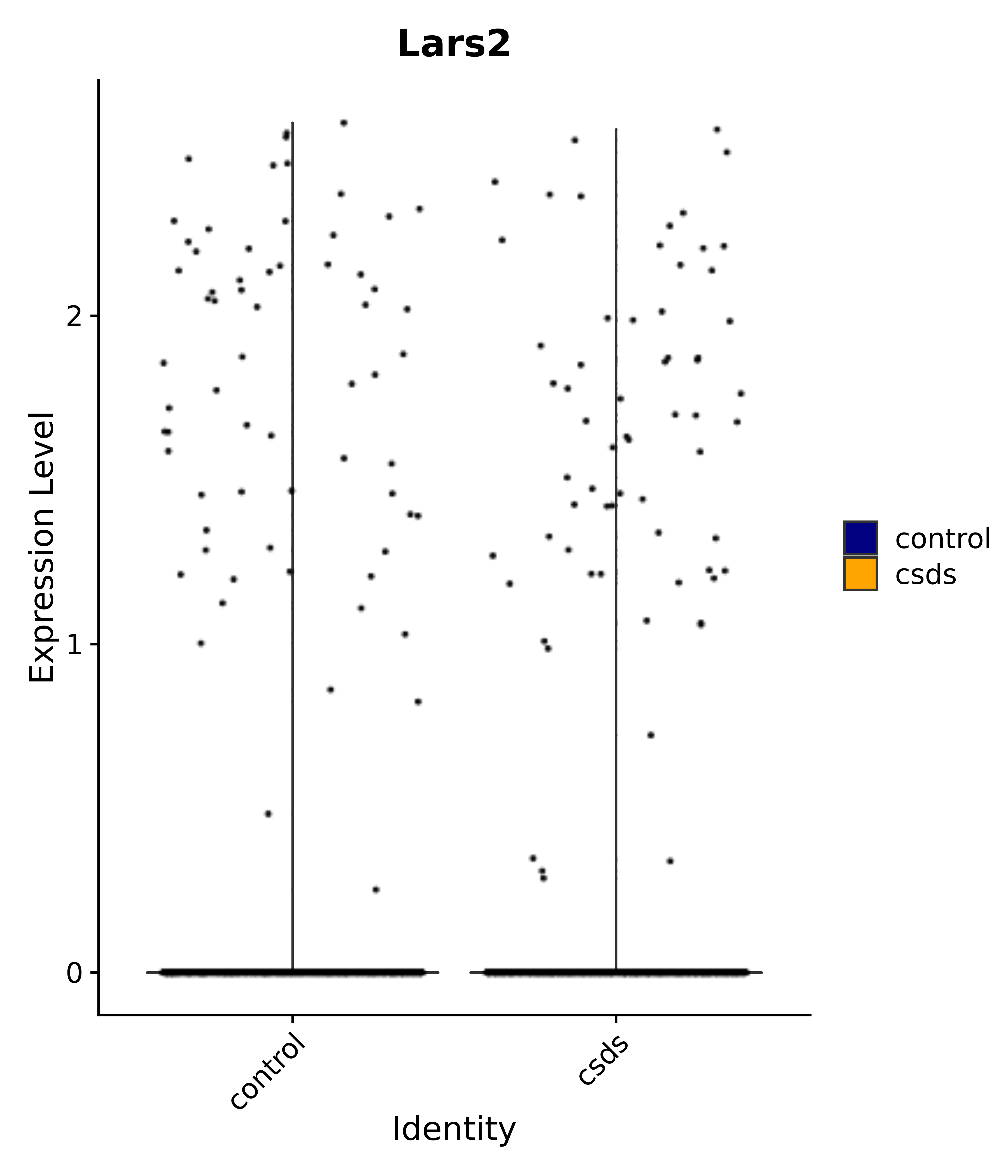
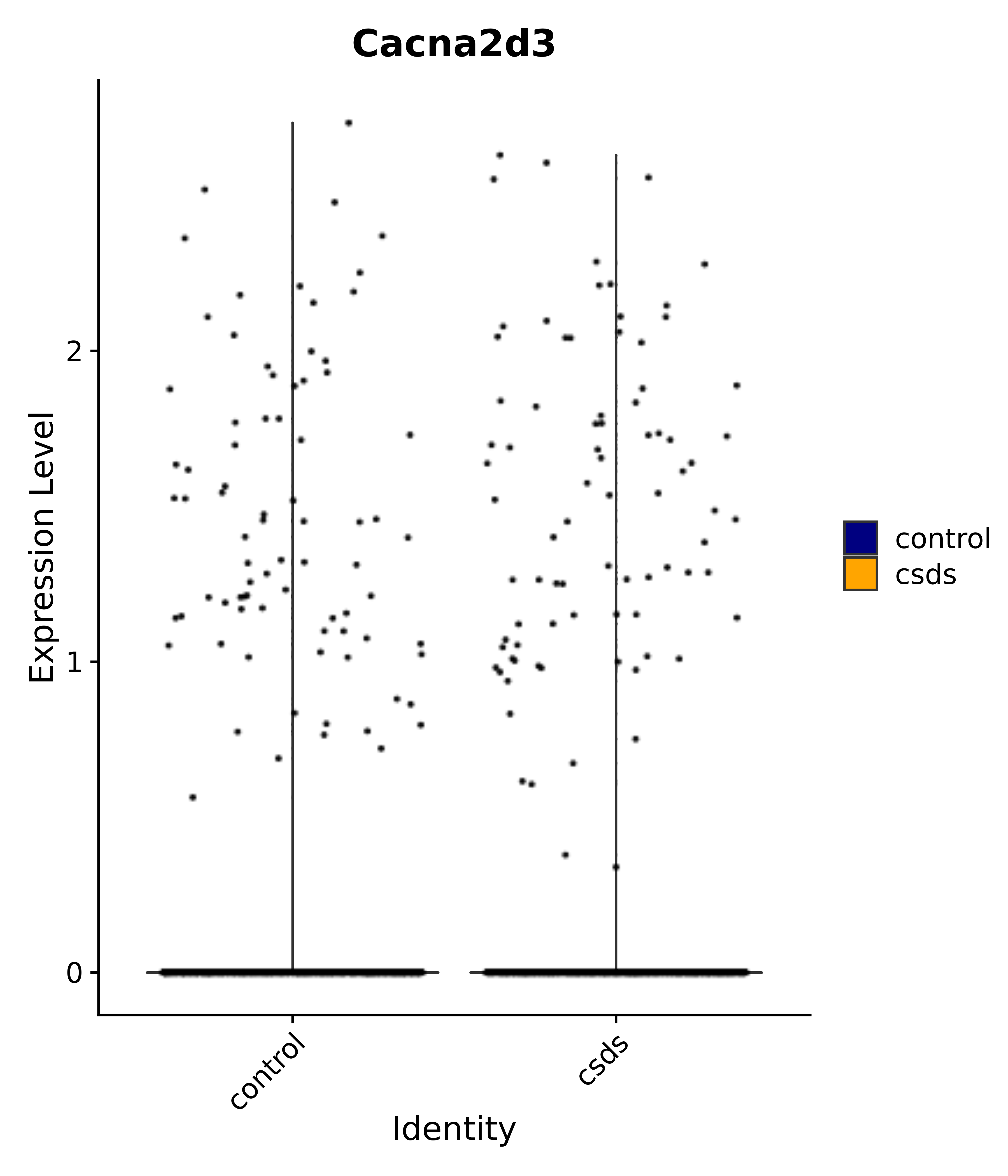
"Insr", "Meis1", "Igf1r", "Nrf1", "Prlr", "Camk1d", "Lars2", "Cacna2d3", # 0 ARC
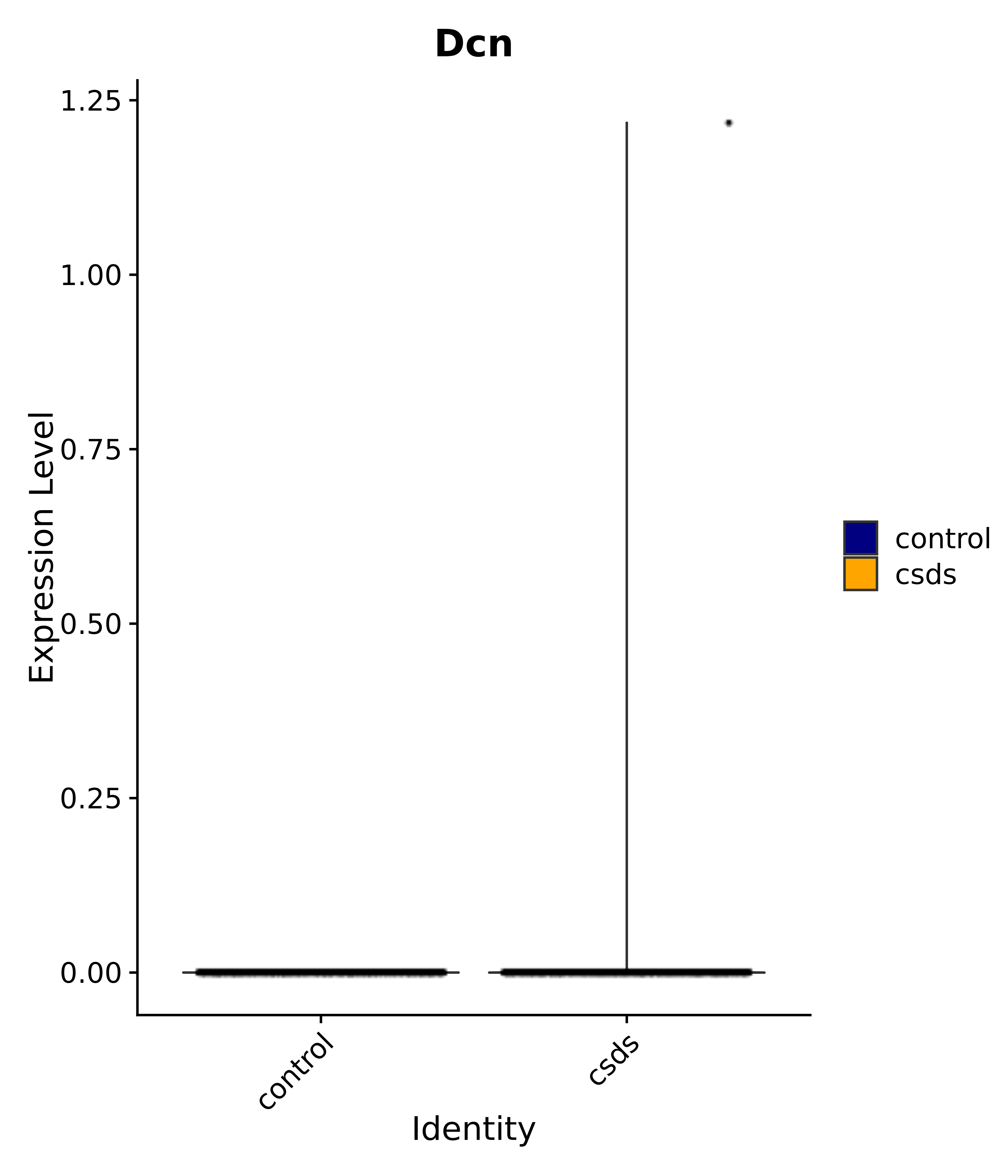
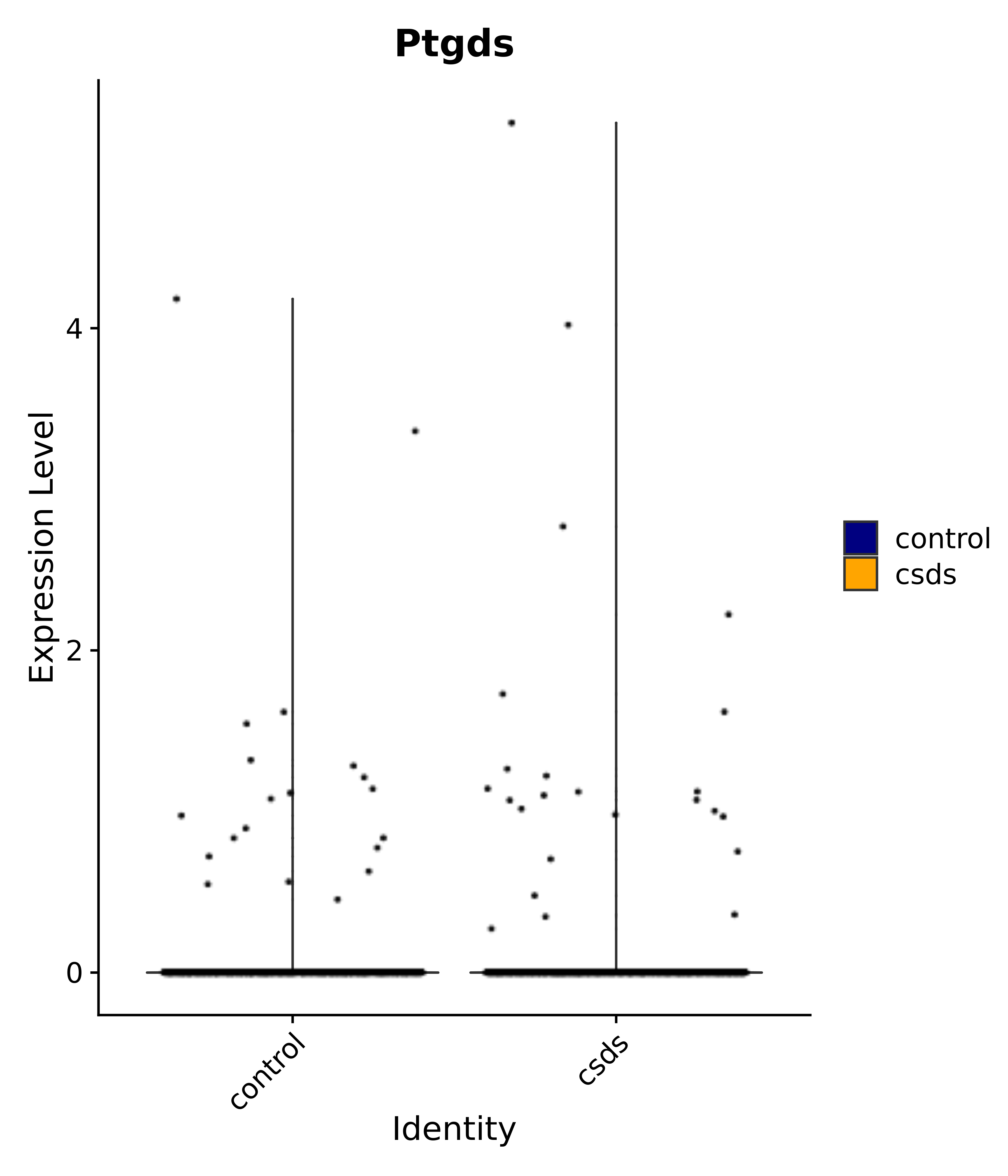
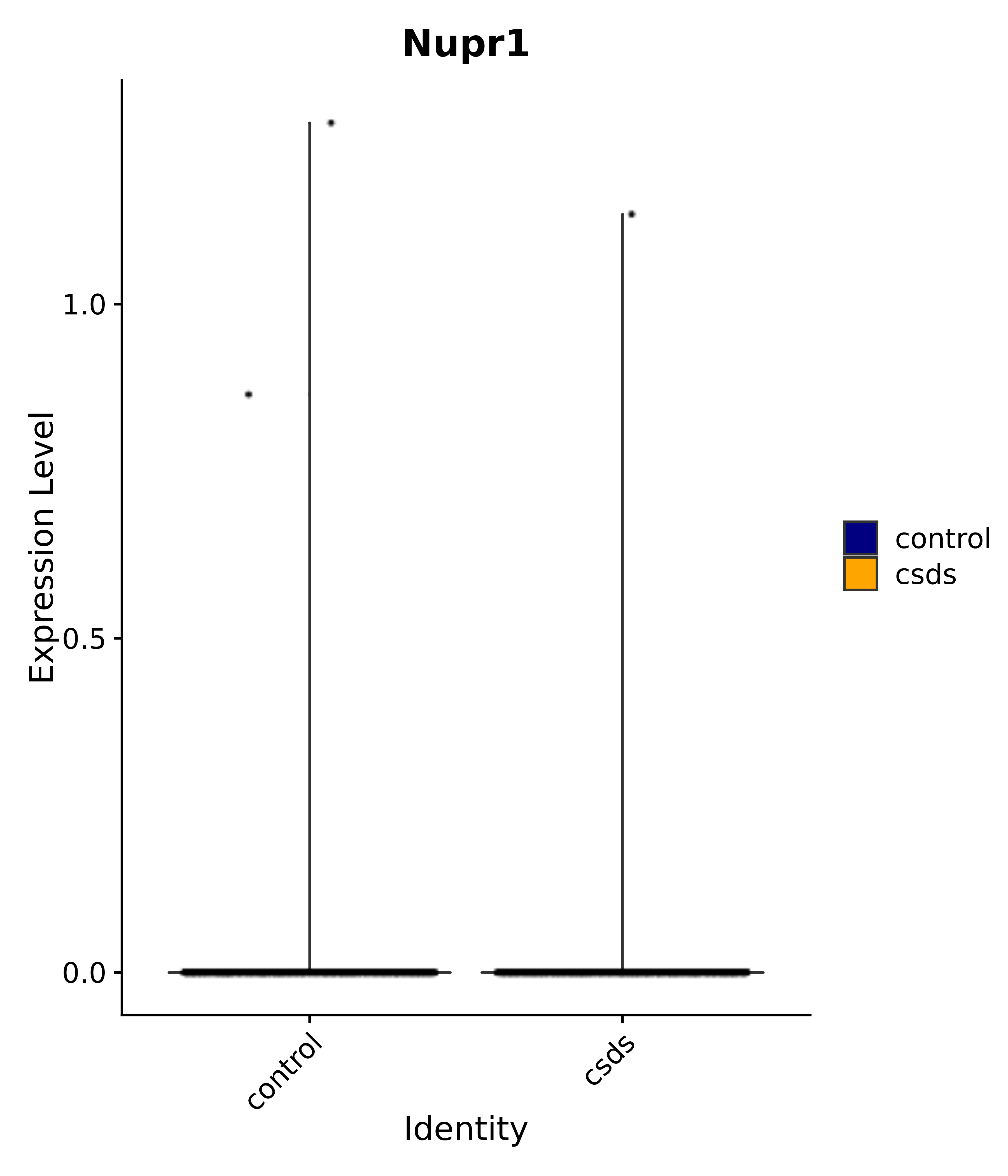
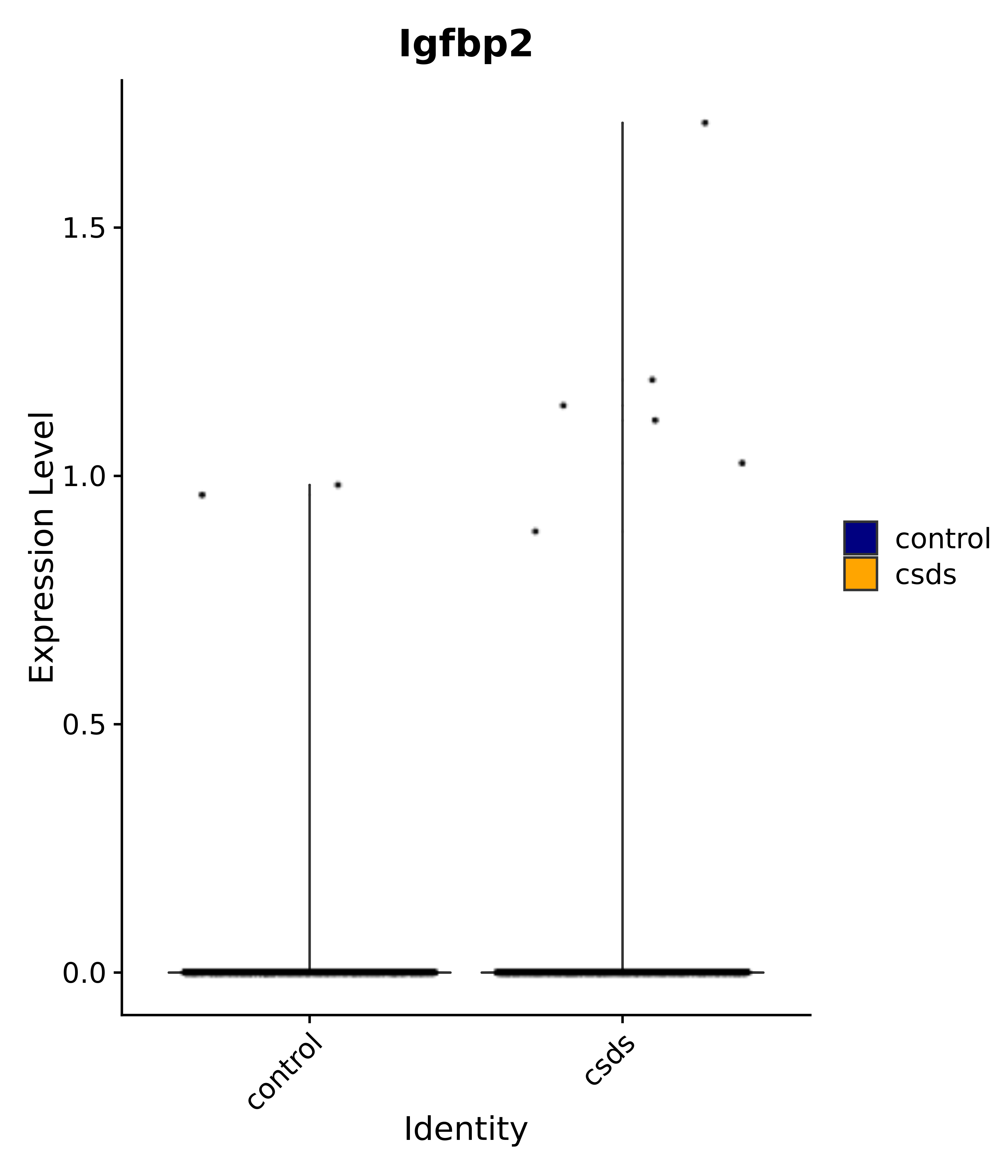
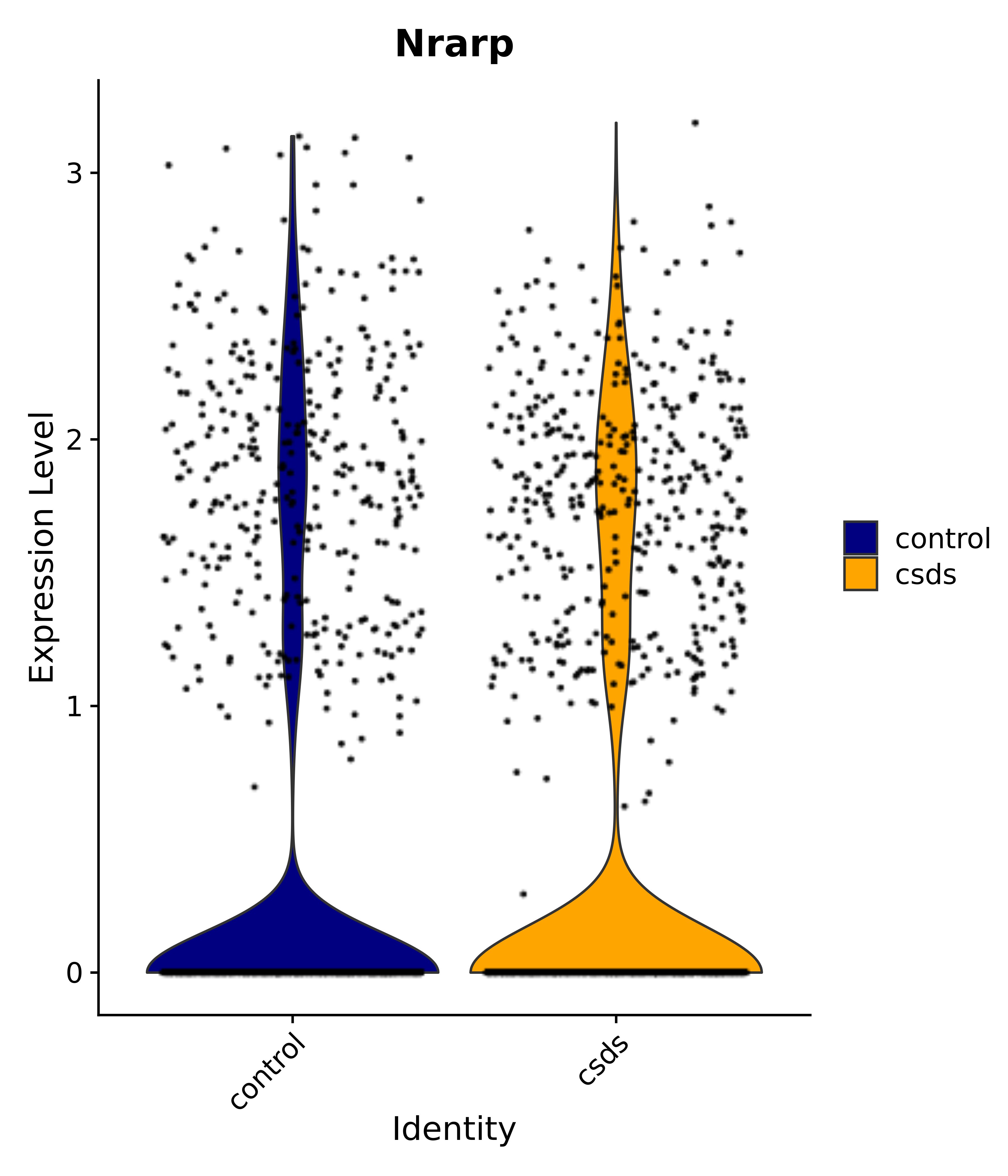
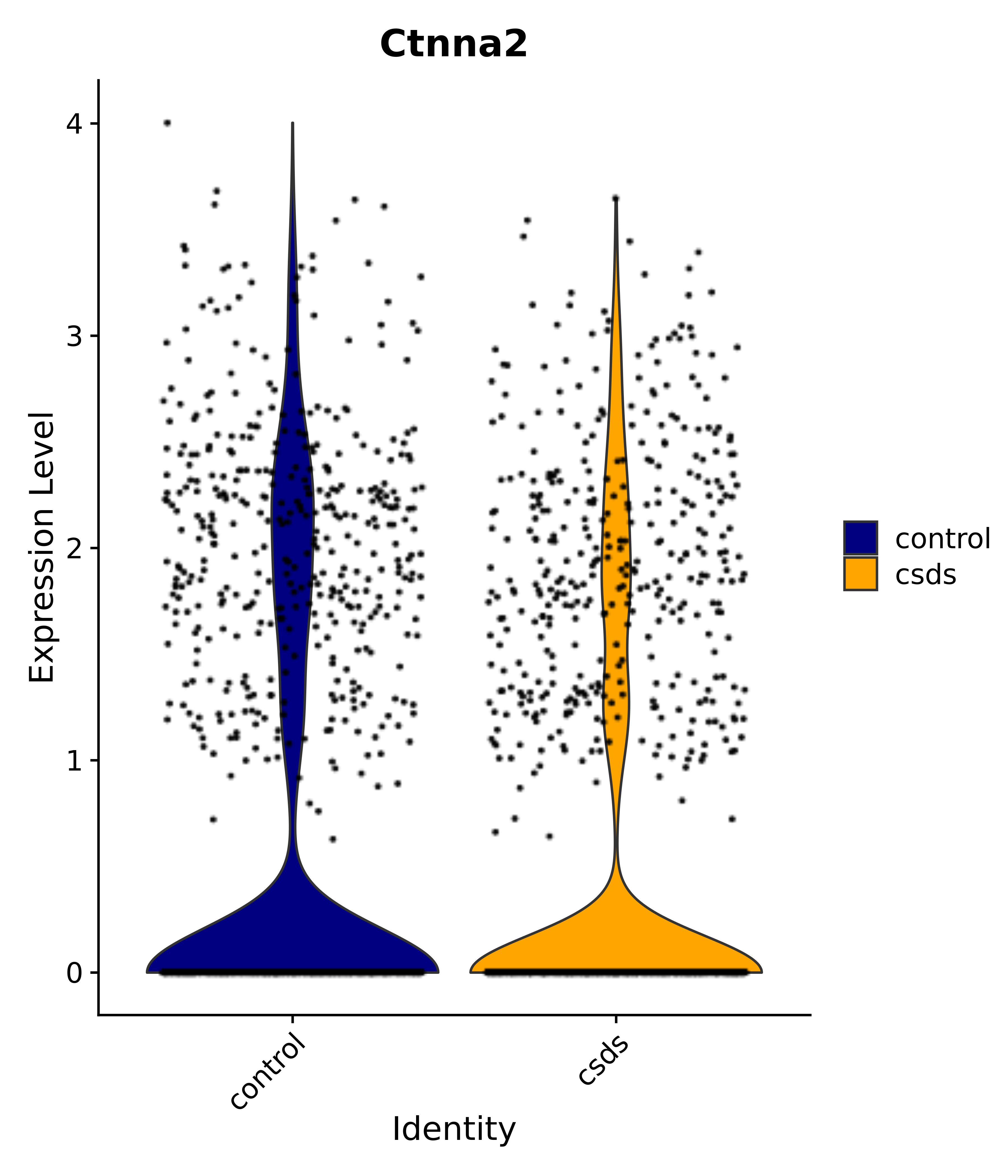
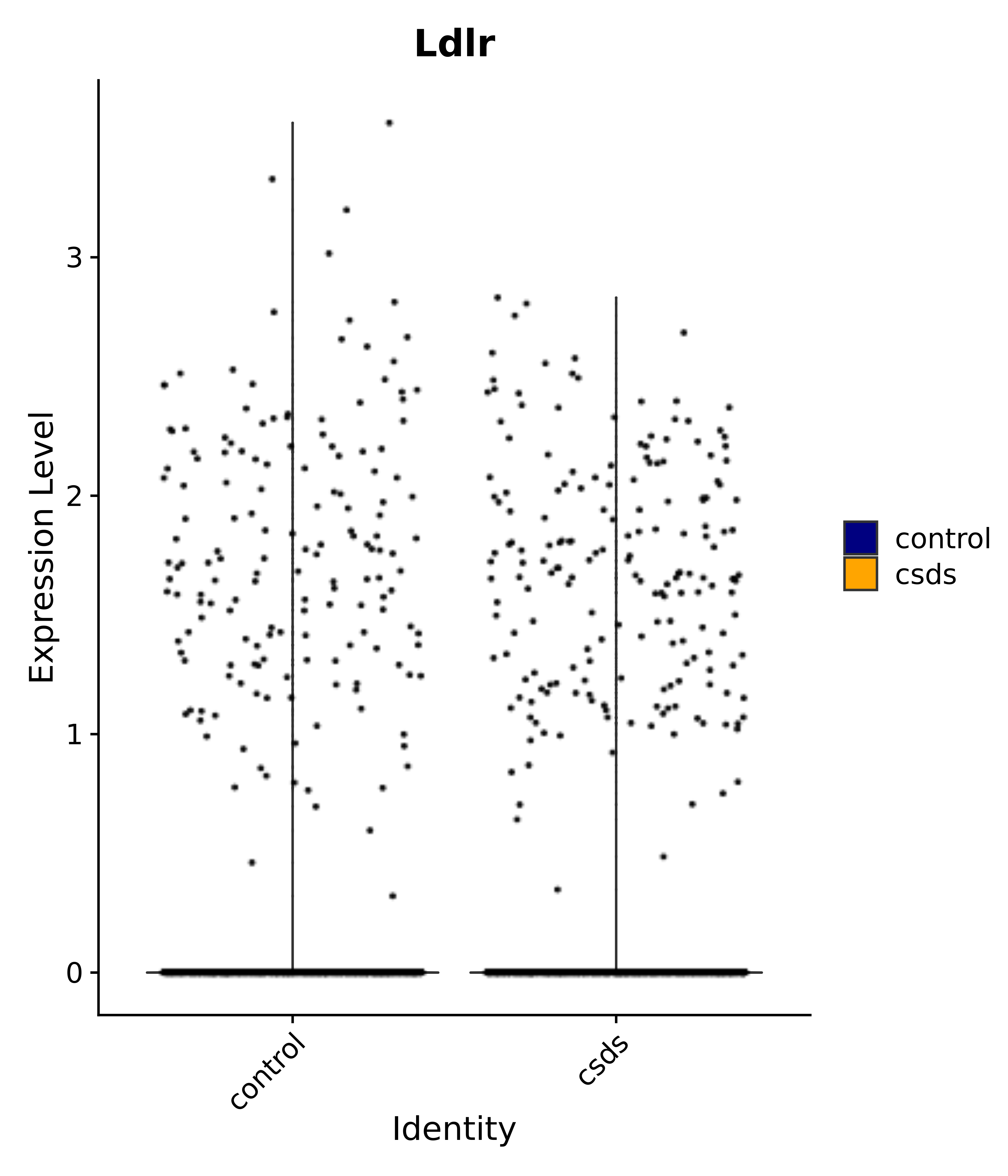
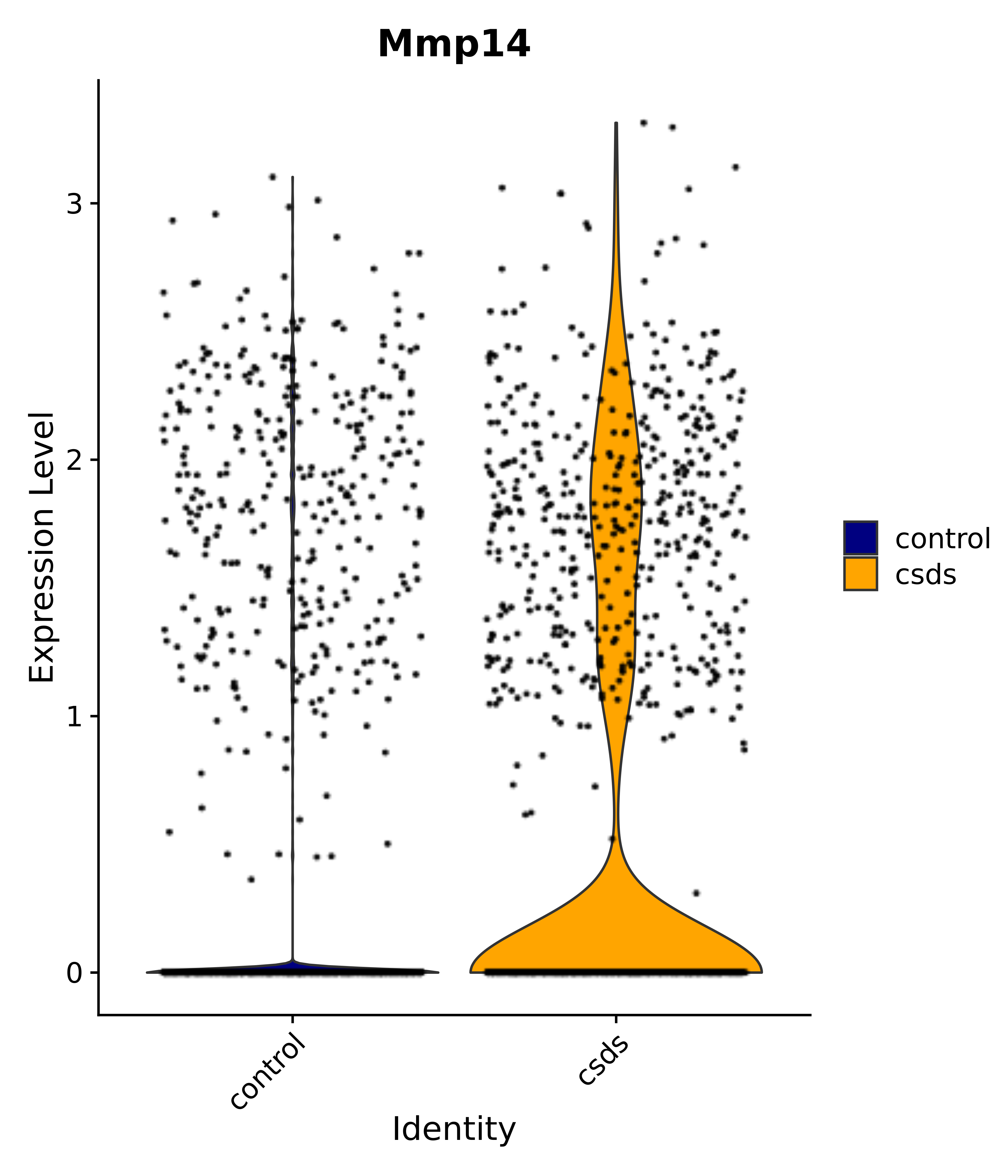
"Dcn", "Ptgds", "Nupr1", "Igfbp2", "Nrarp", "Ctnna2", "Ldlr", "Mmp14", # 1 LHA
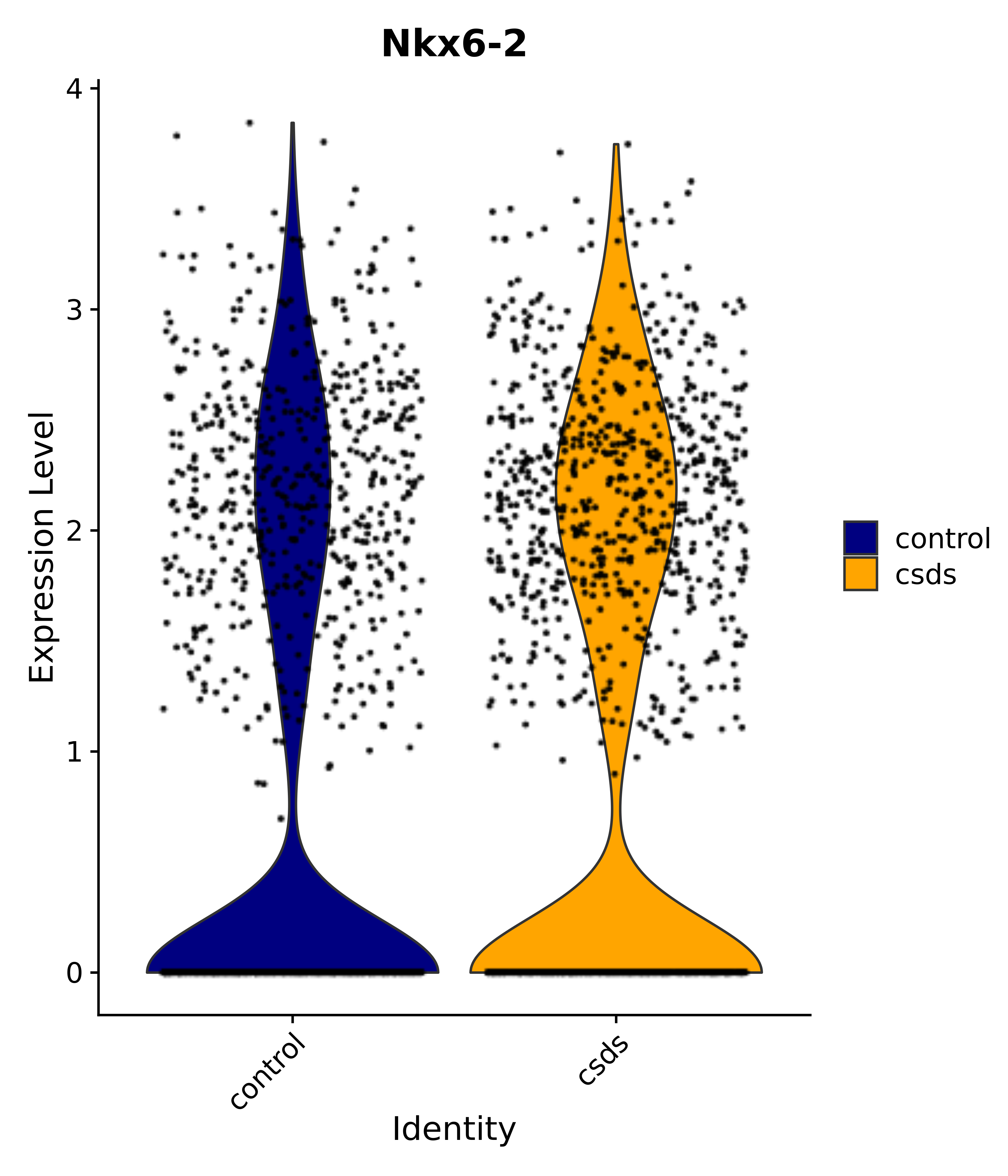
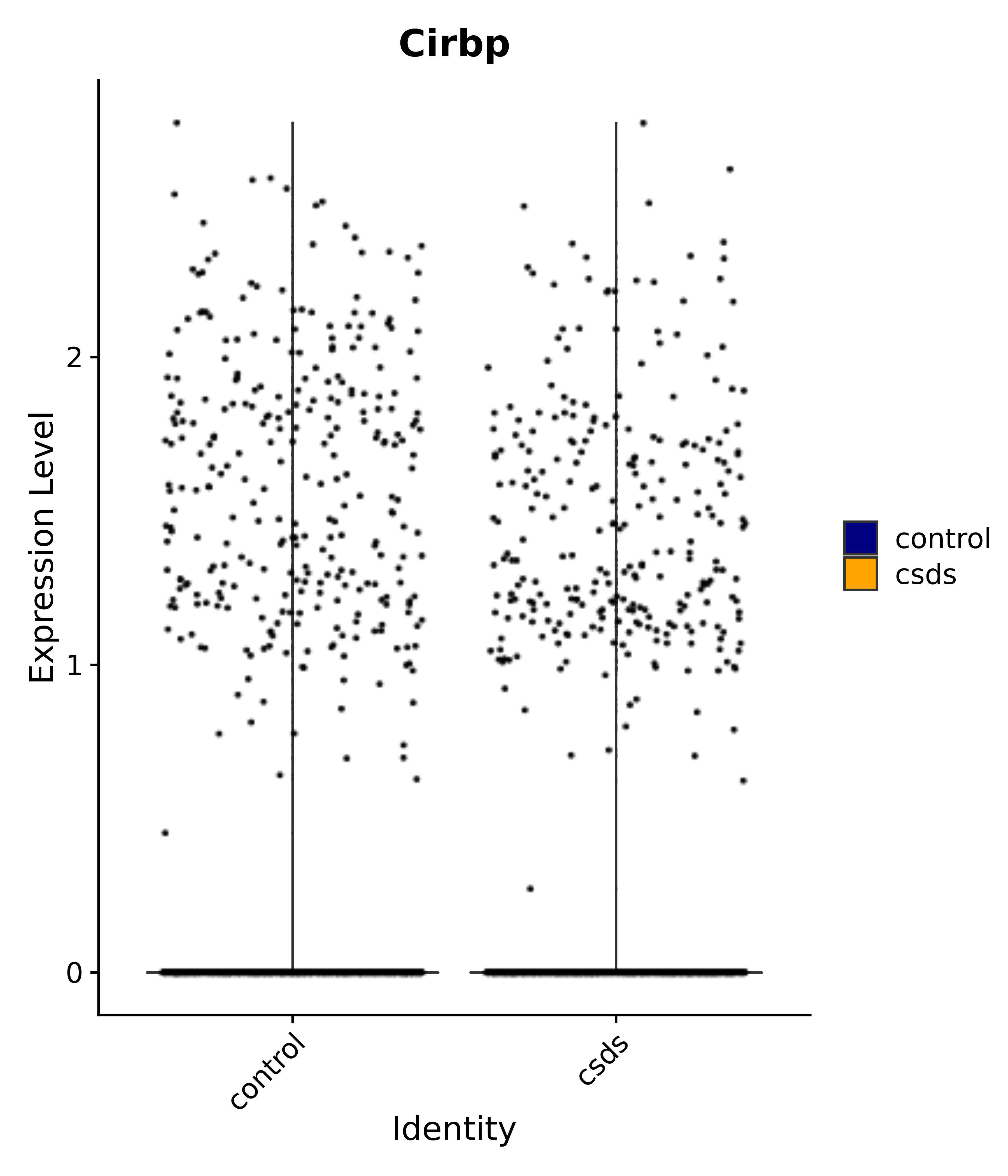
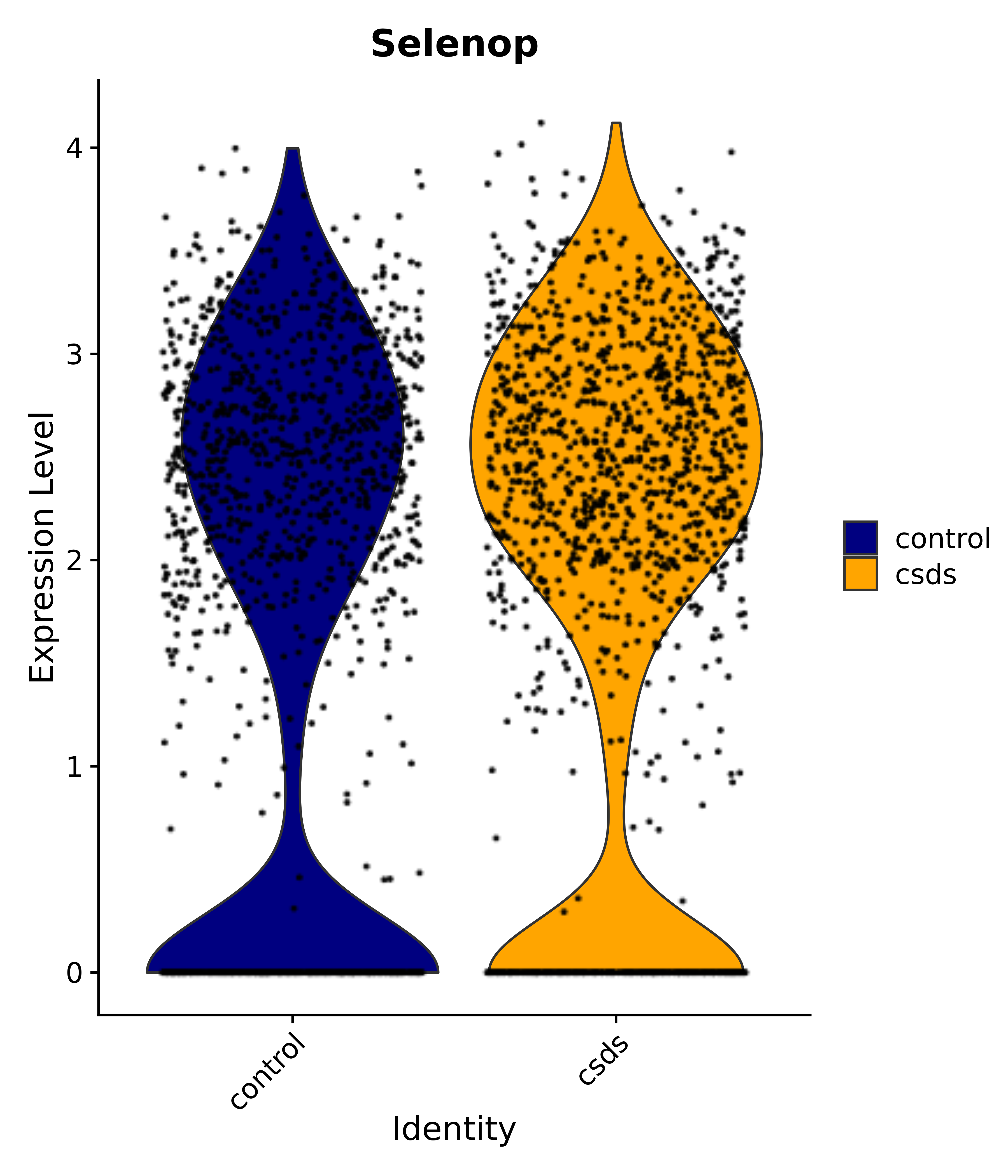
"Nkx6-2", "Cirbp", "Selenop", # 2 MnPO
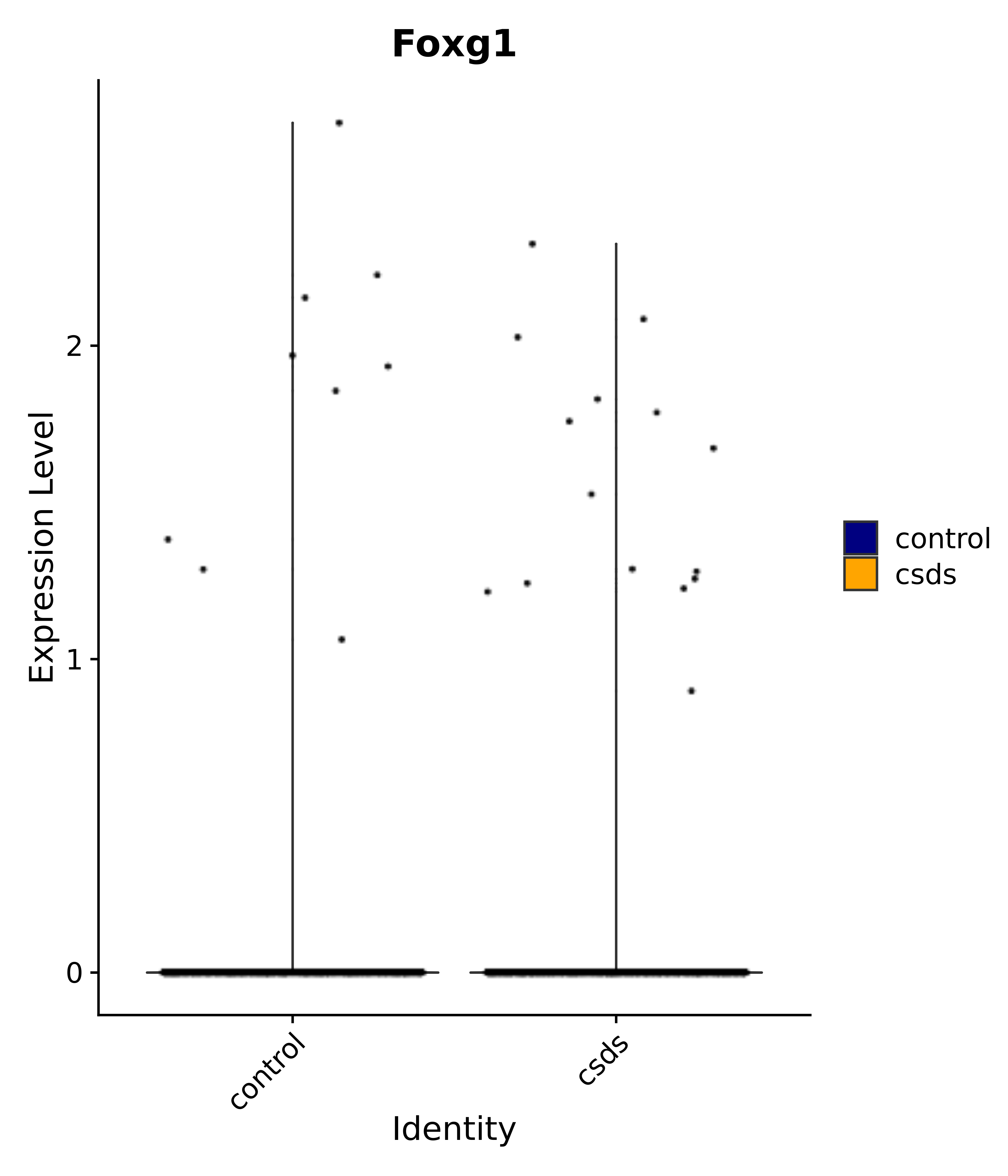
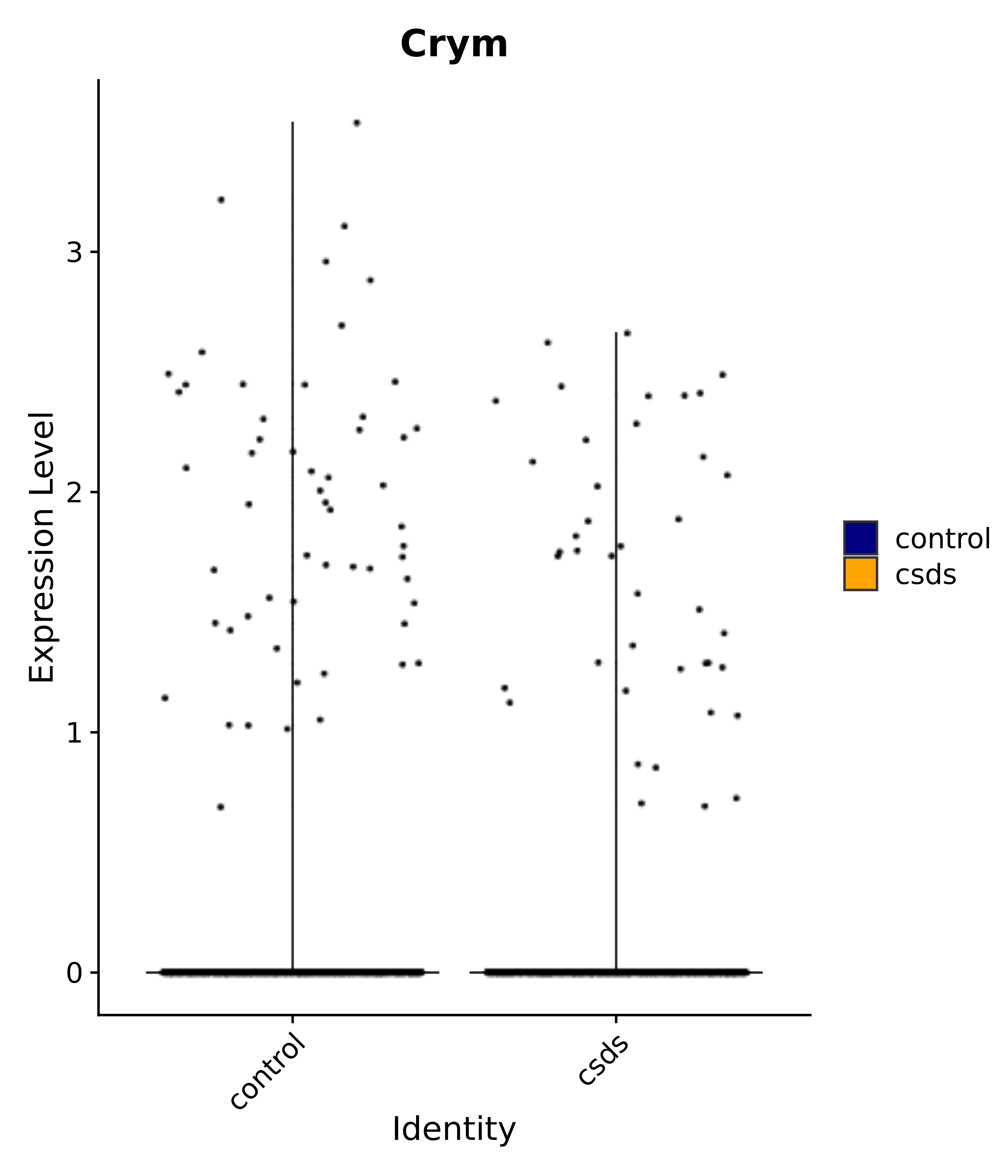
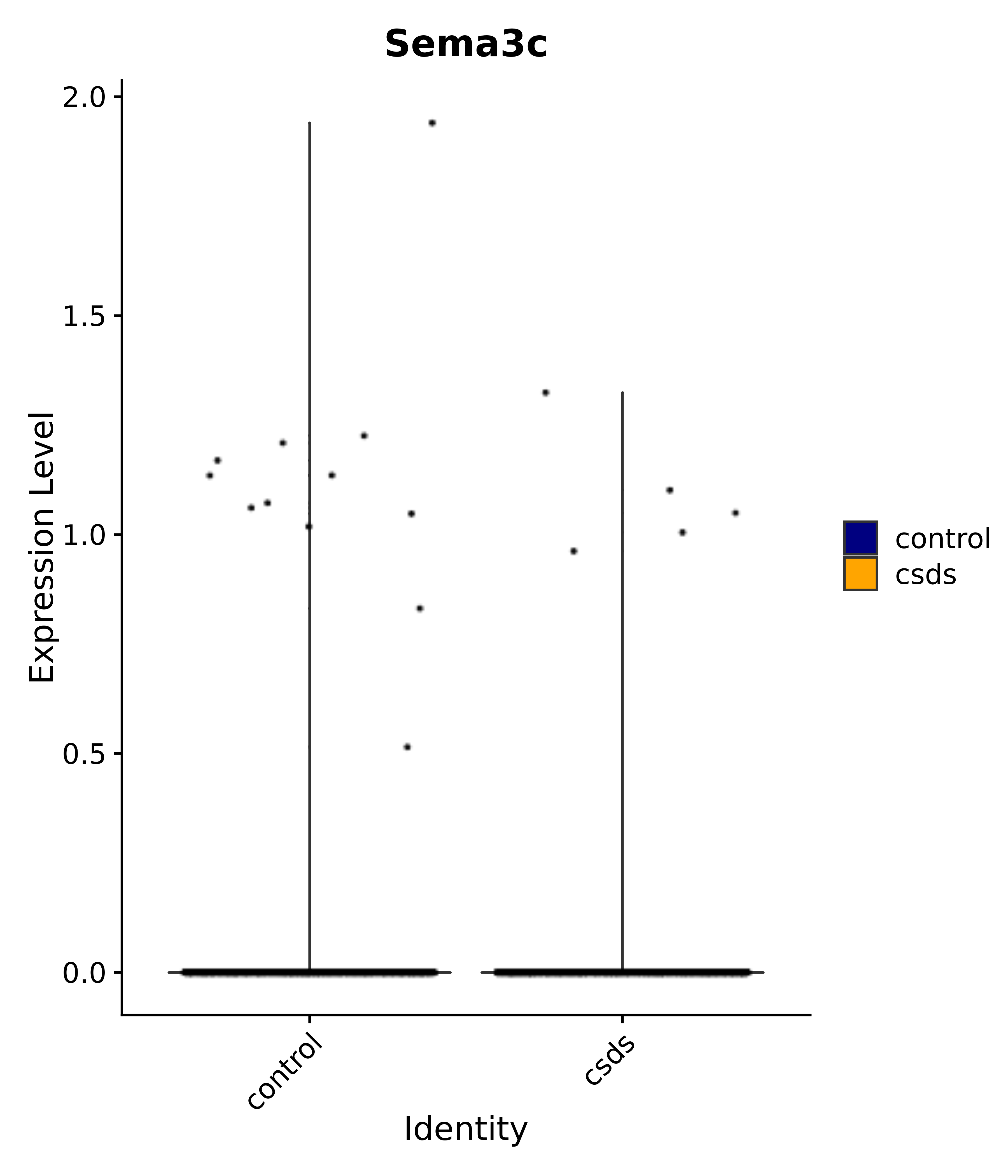
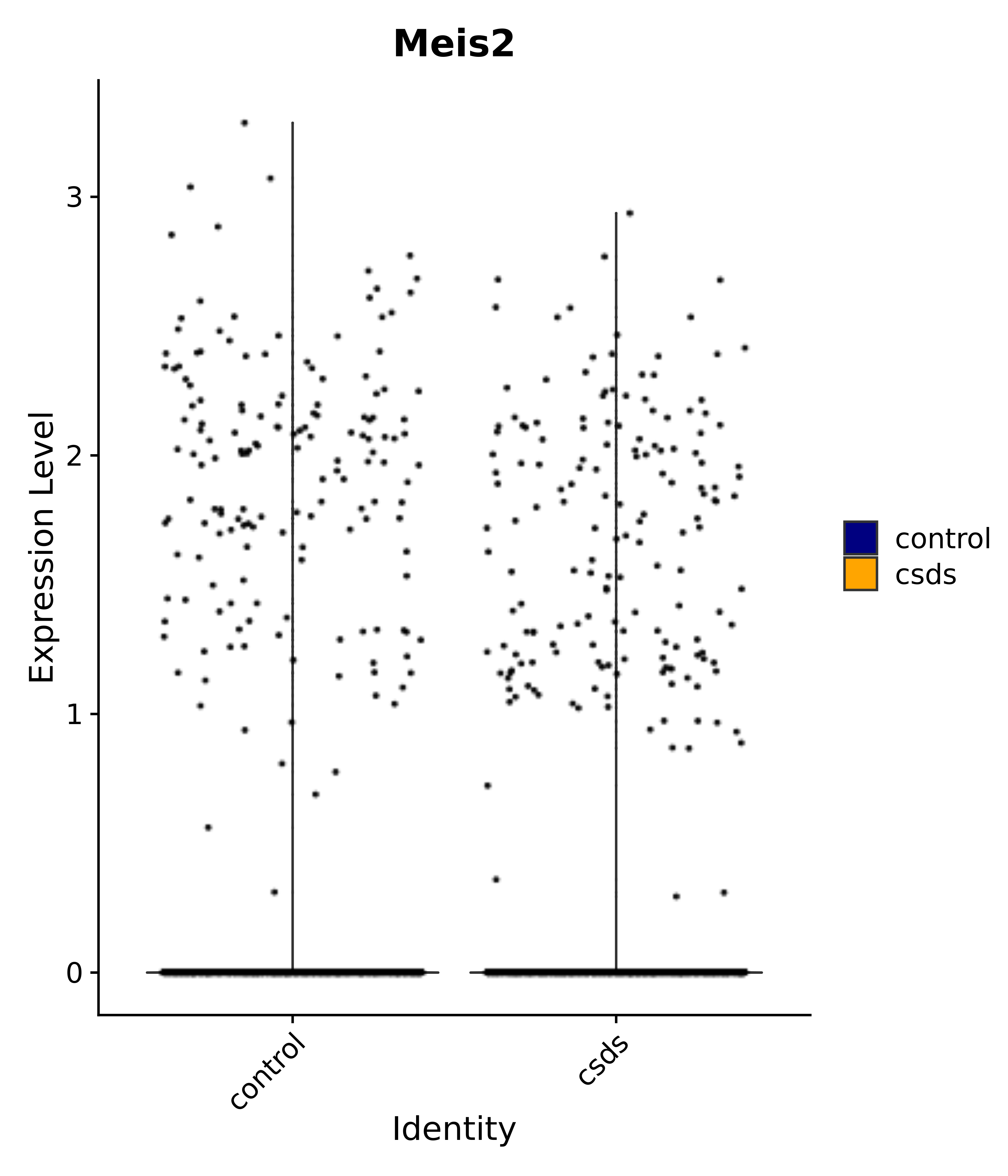
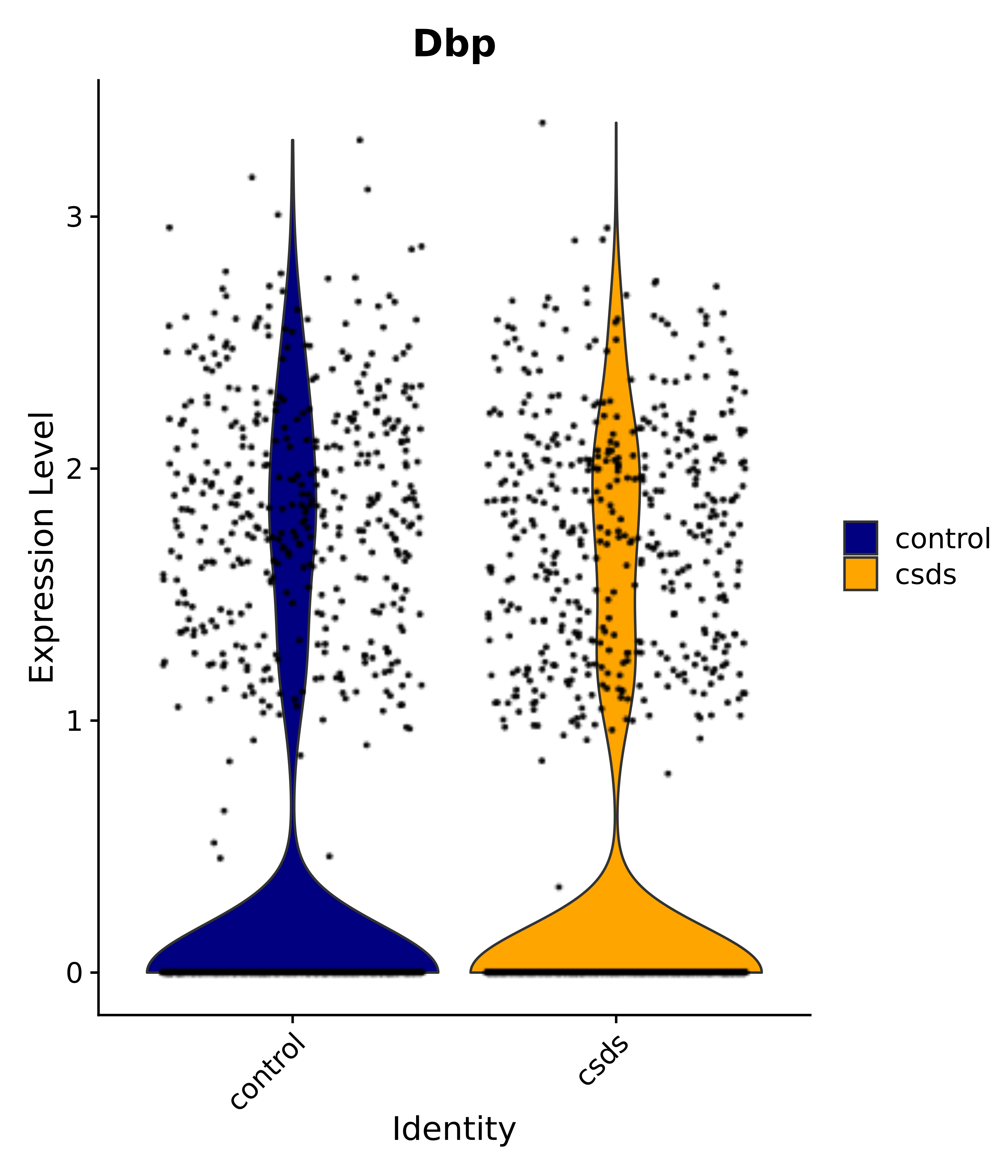
"Foxg1", "Crym", "Sema3c", "Meis2", "Dbp", # 3 POA
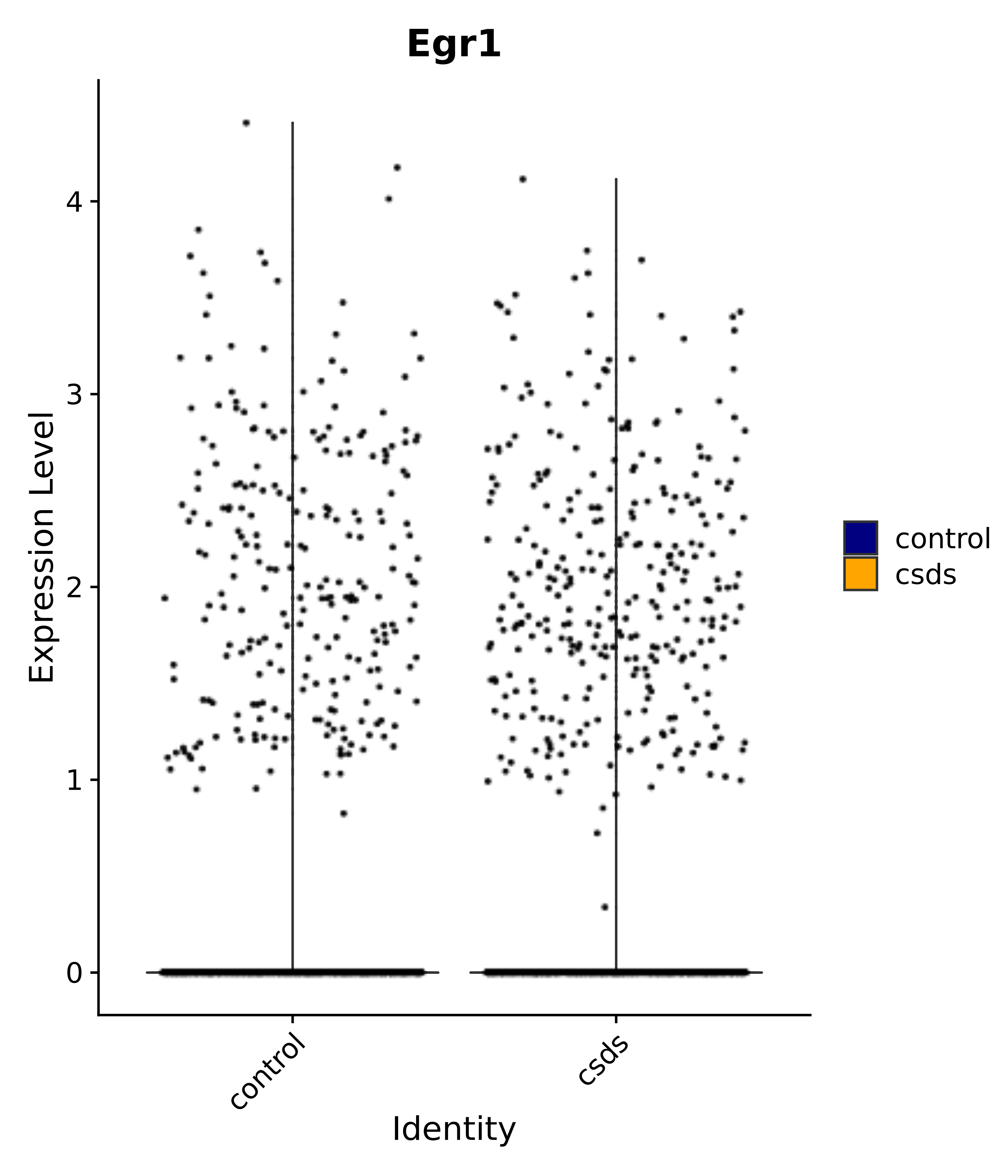
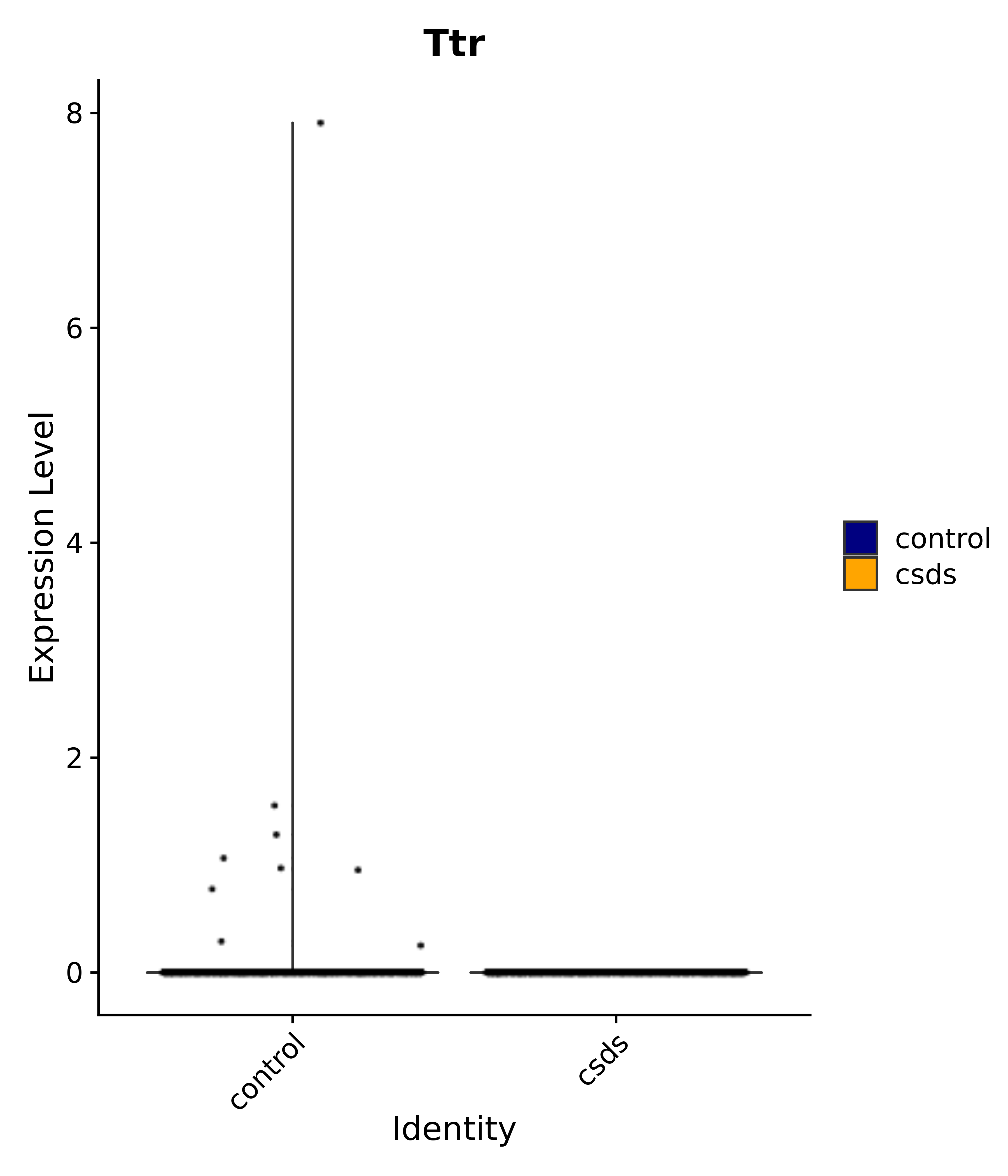
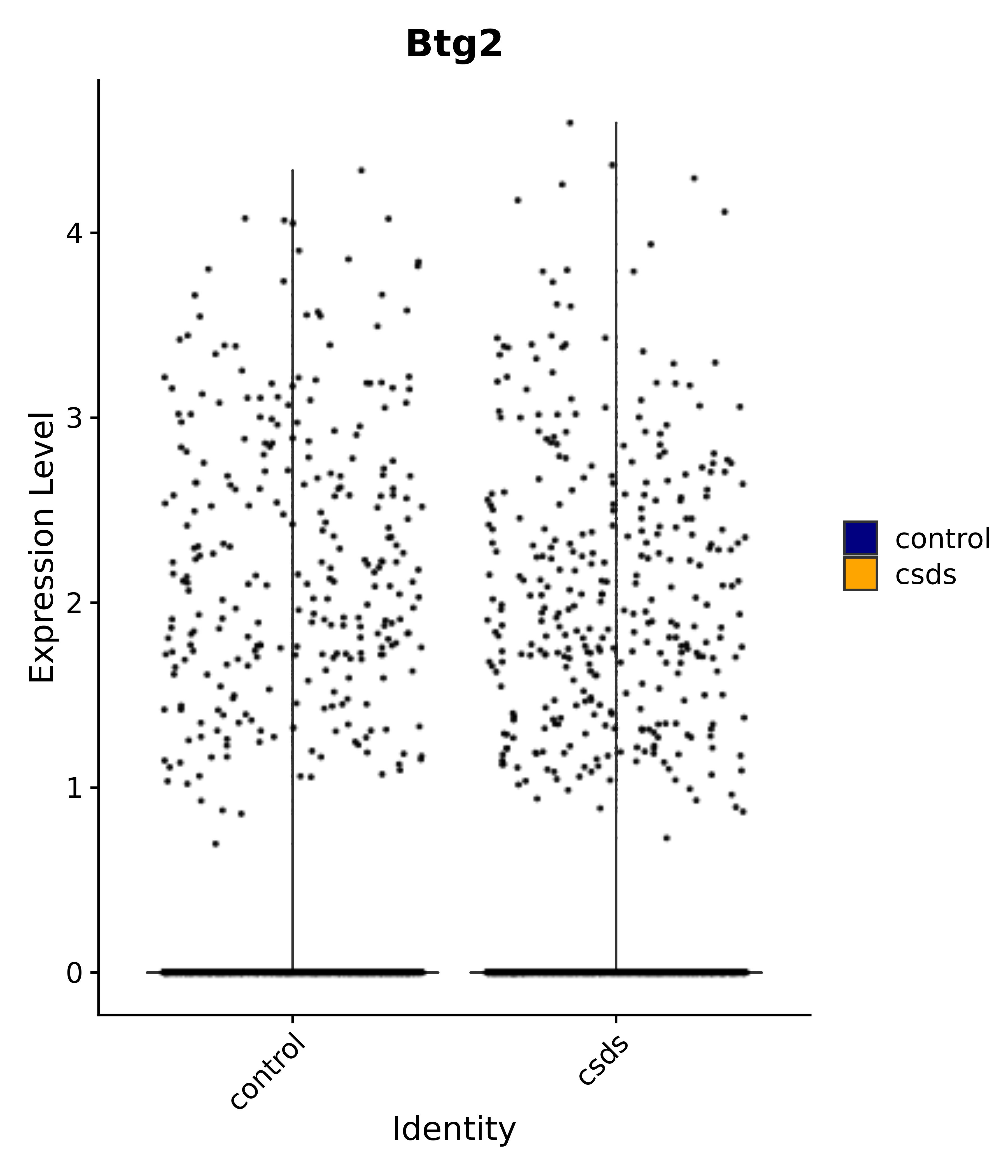
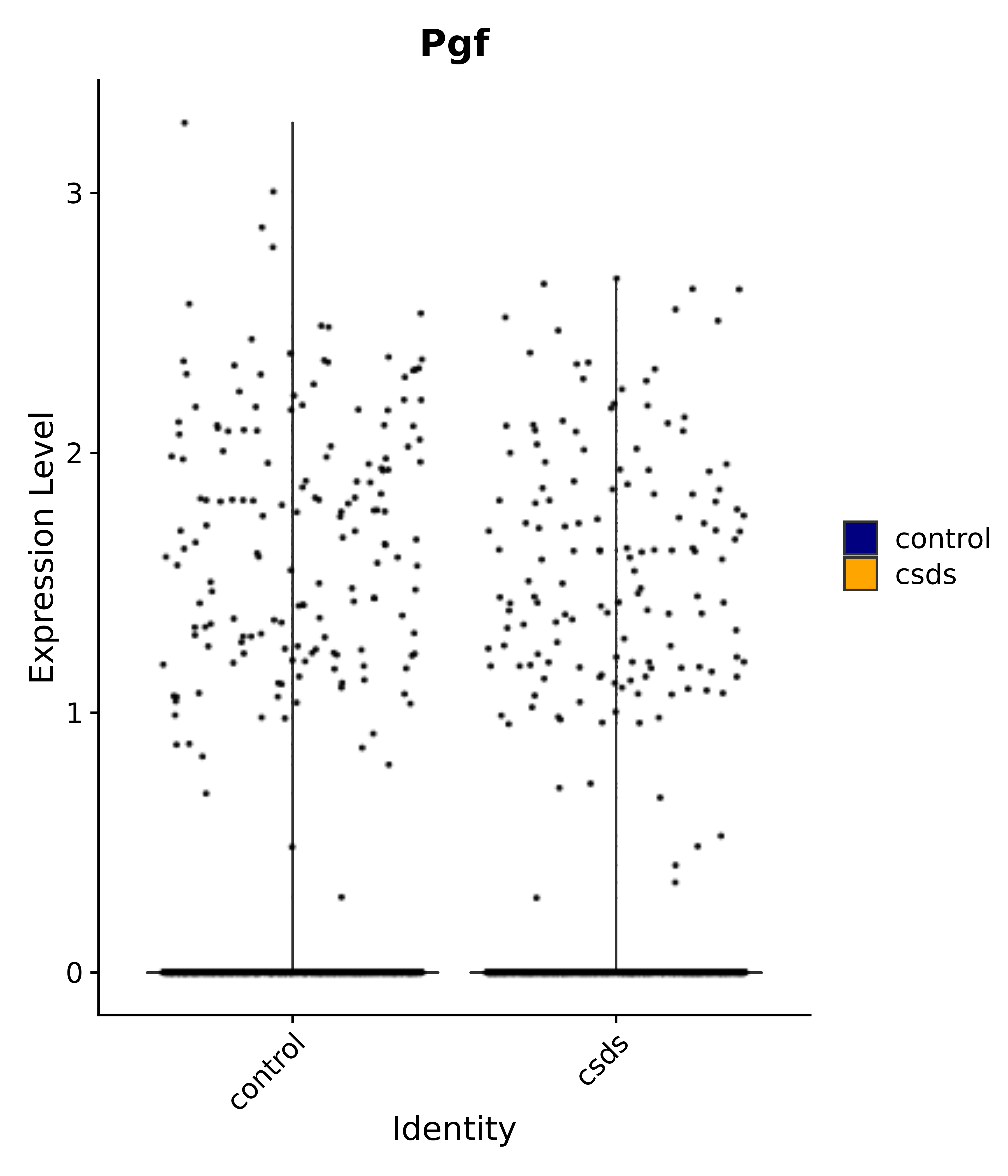
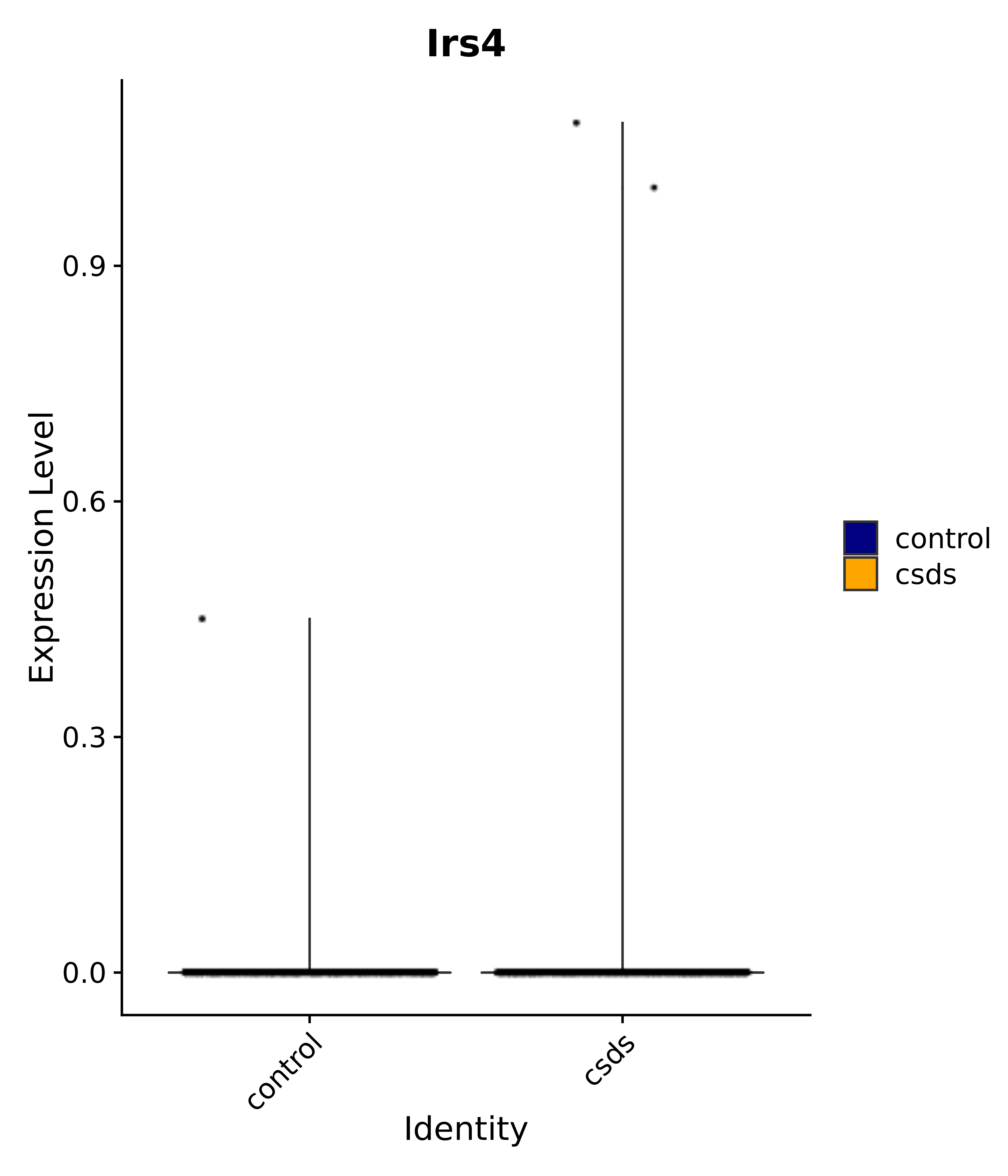
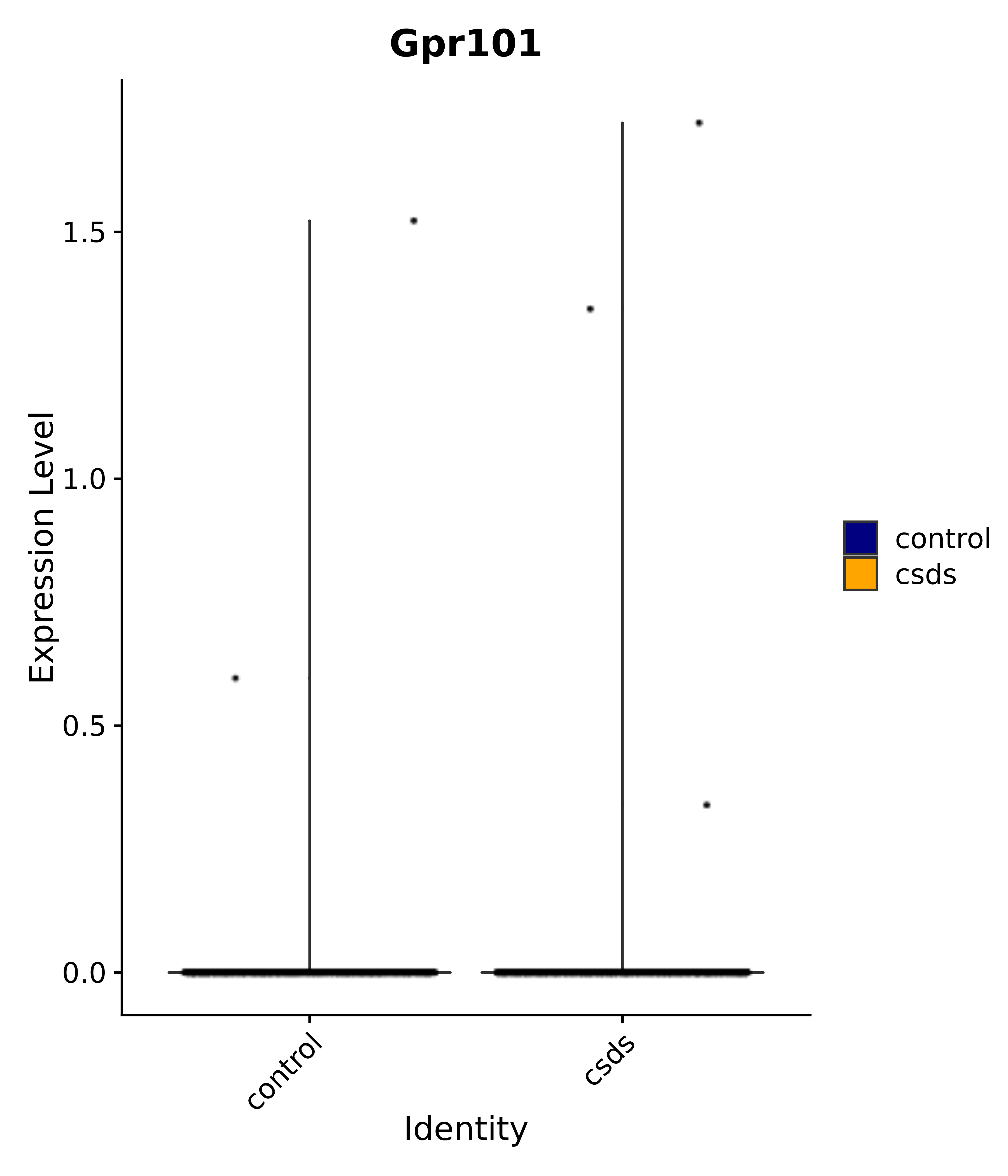
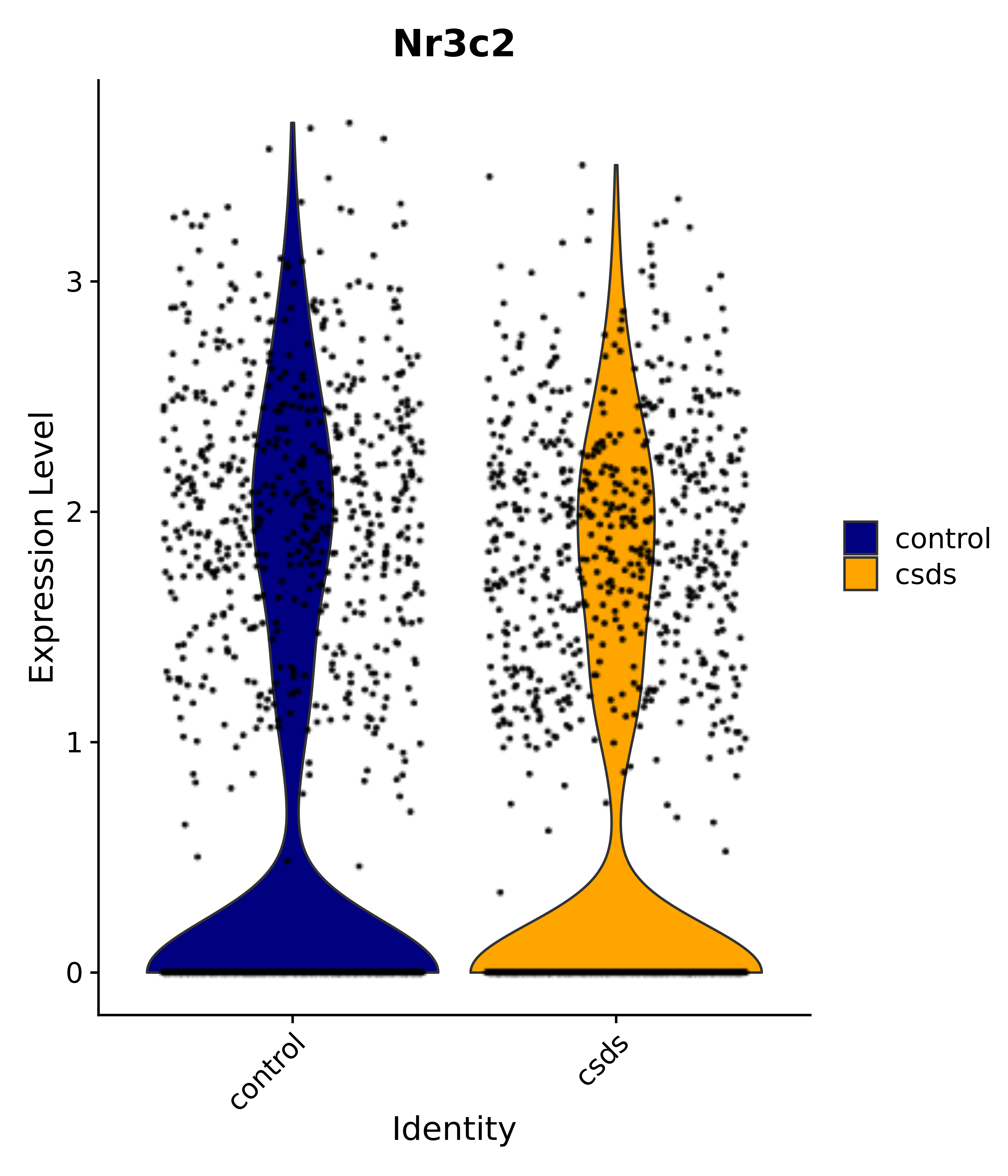
"Egr1", "Ttr", "Btg2", "Mbnl3", "Pgf", "Irs4", "Gpr101", "Nr3c2", "Agtr1", # 4 PVN
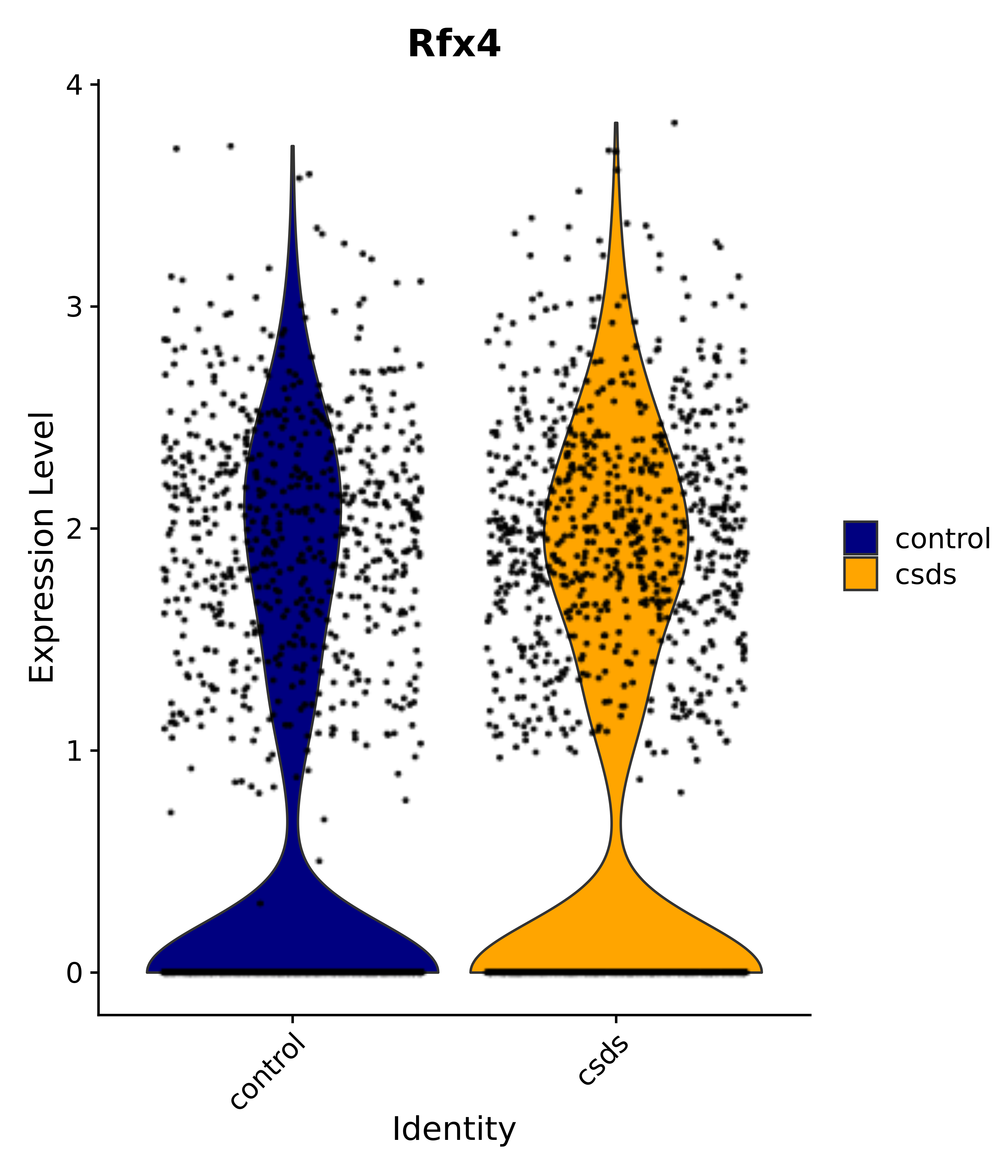
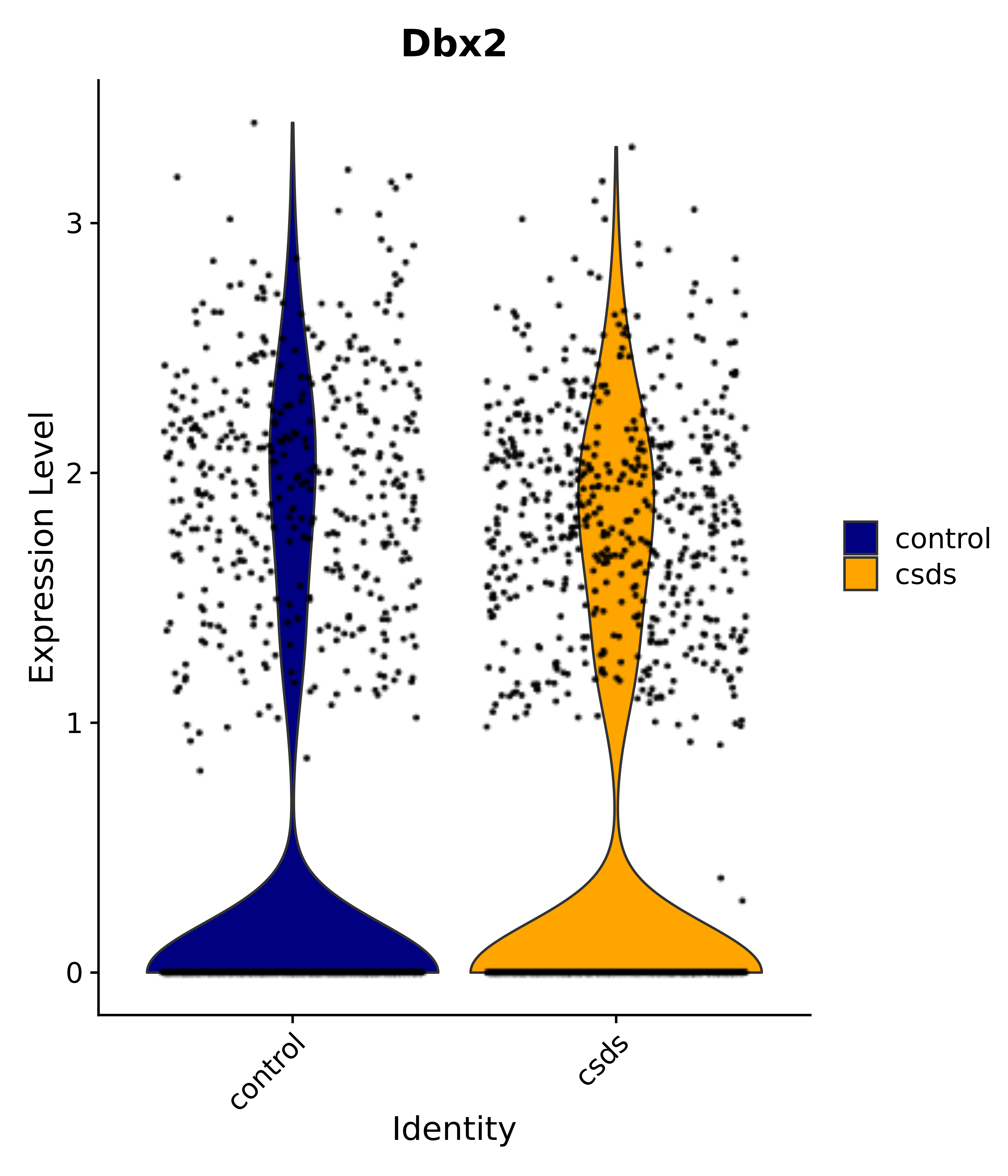
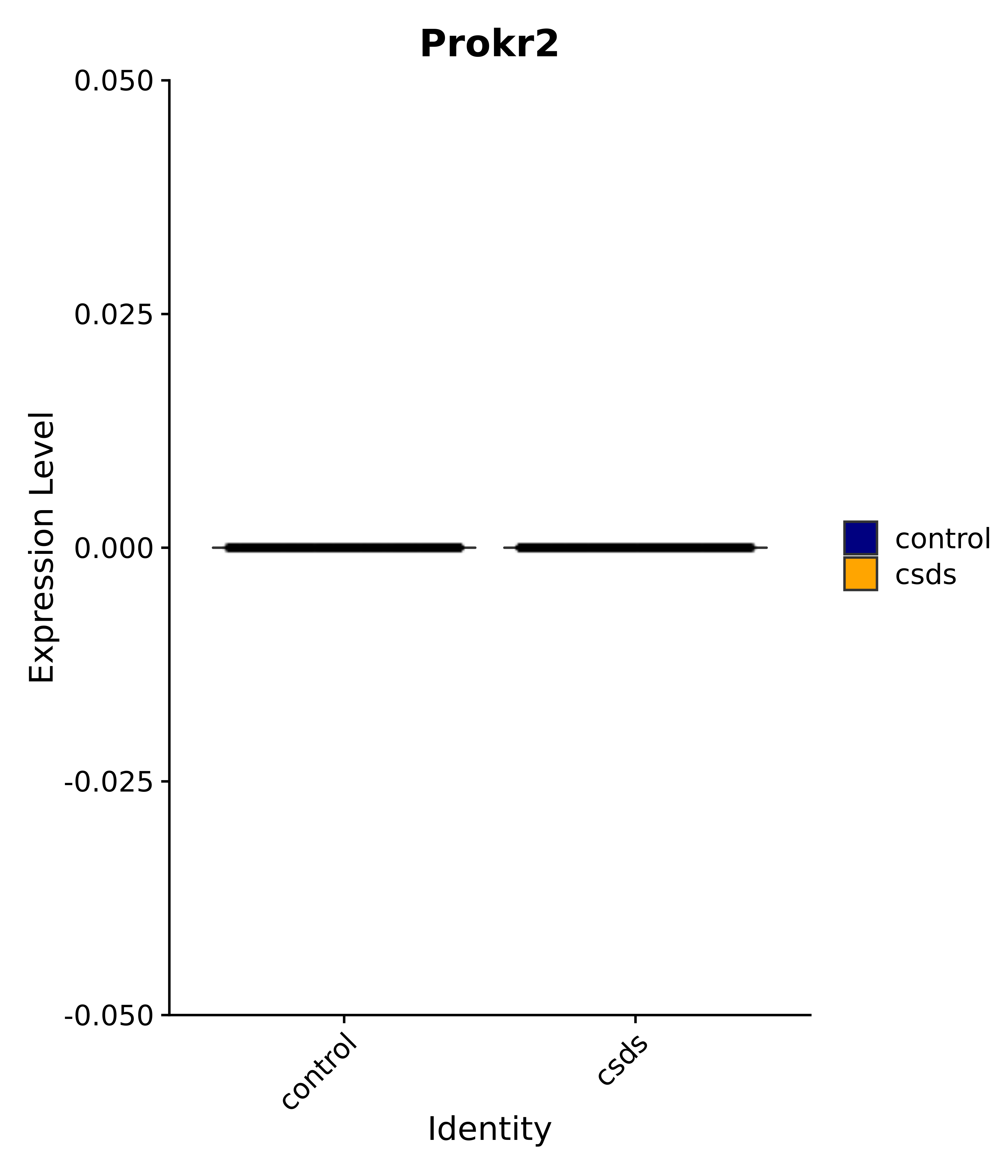
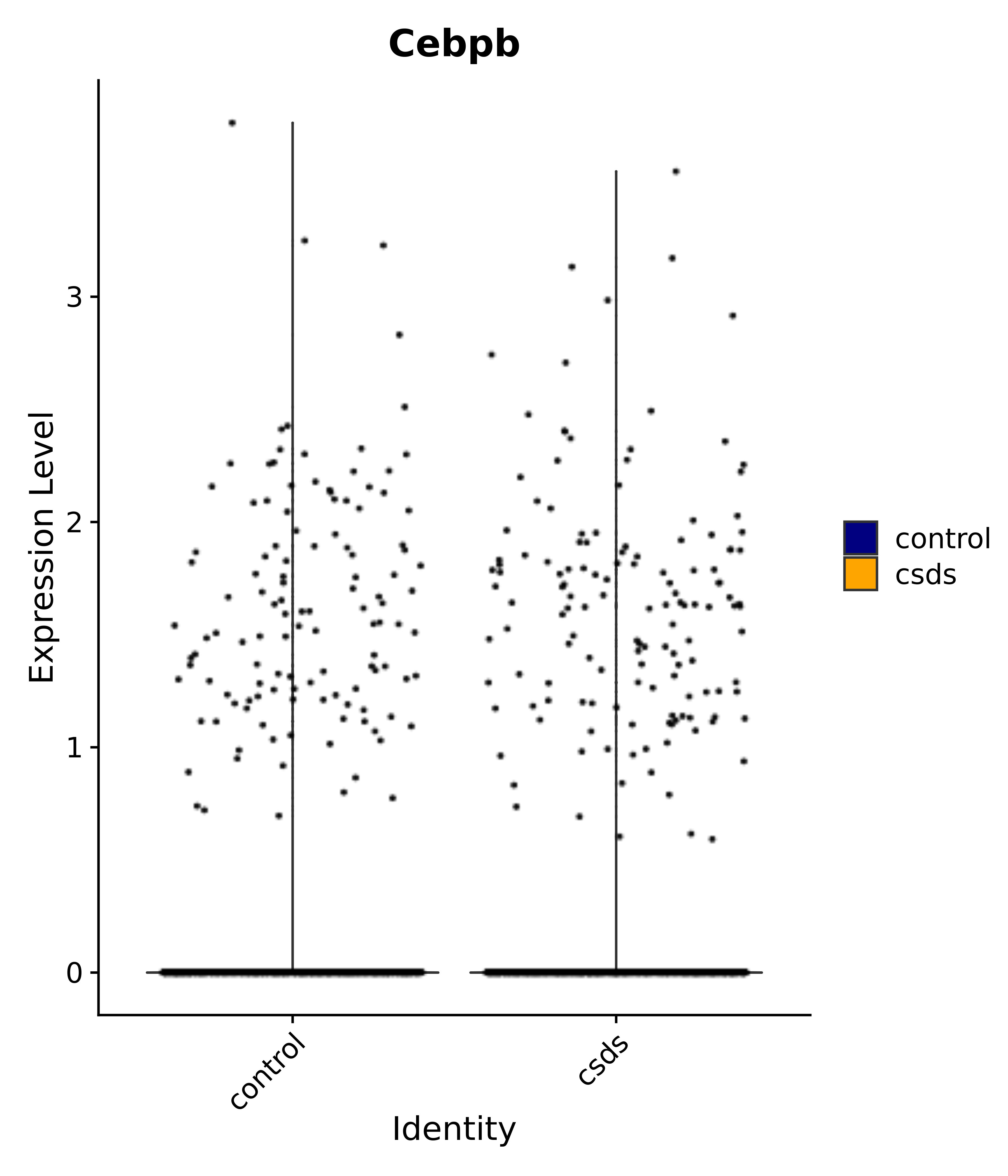
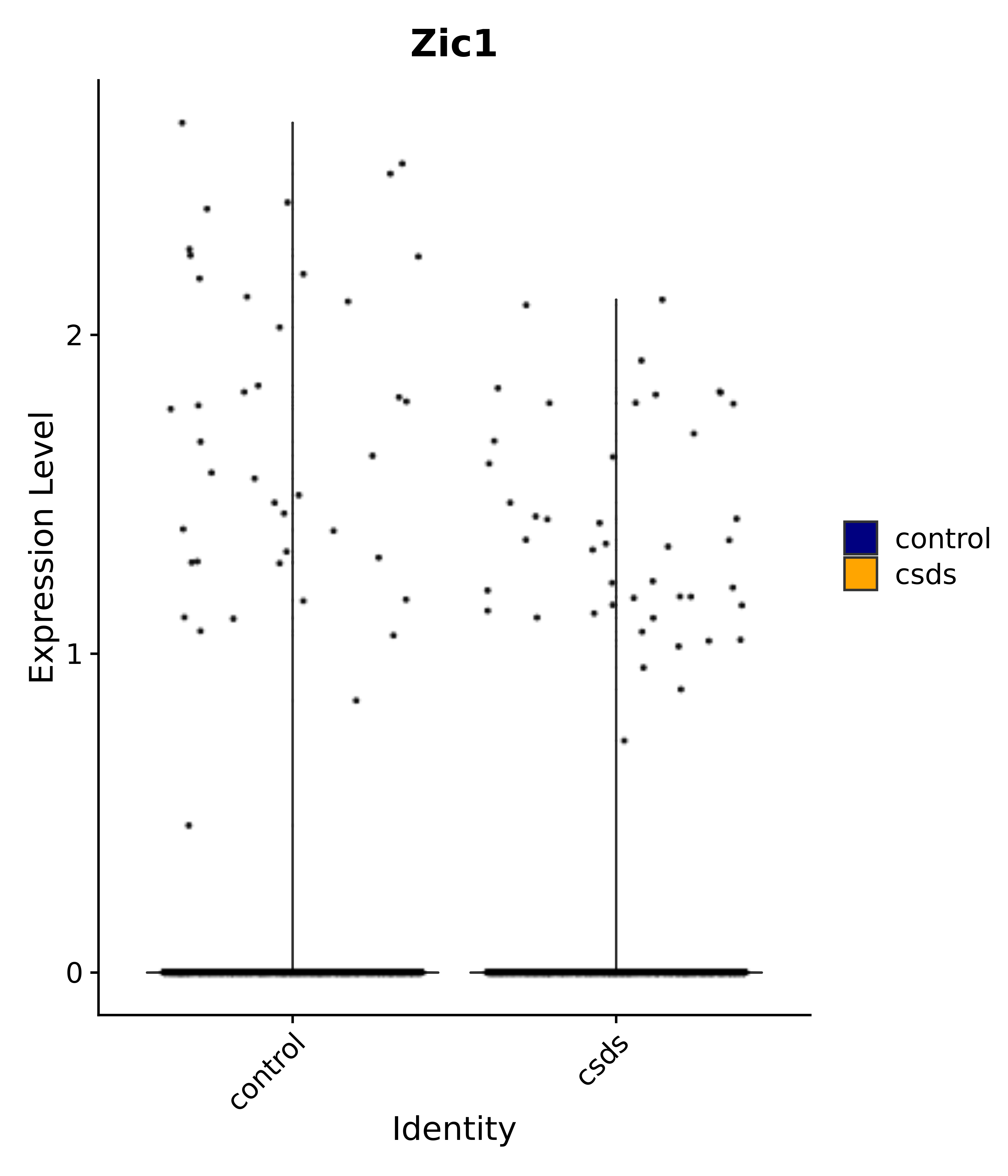
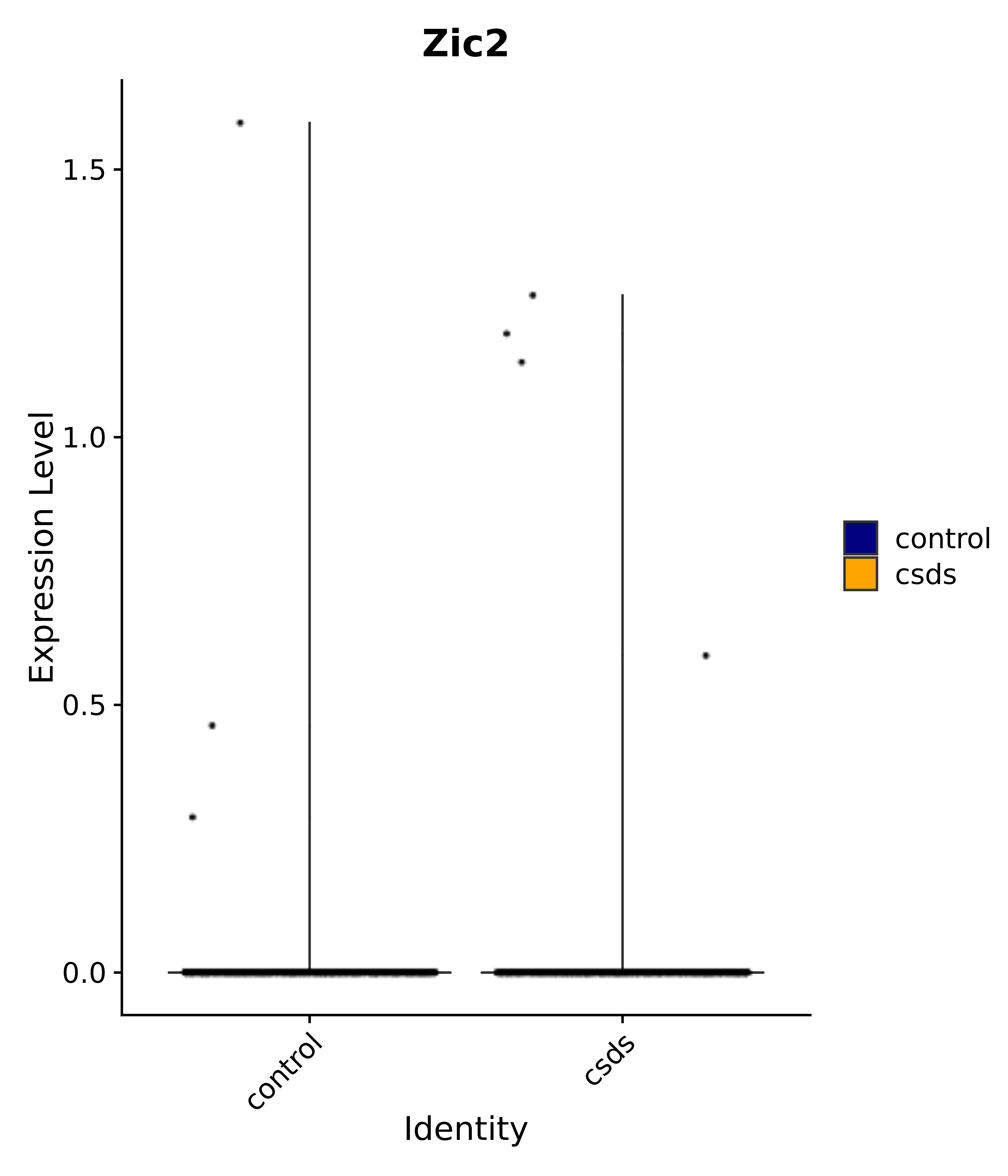
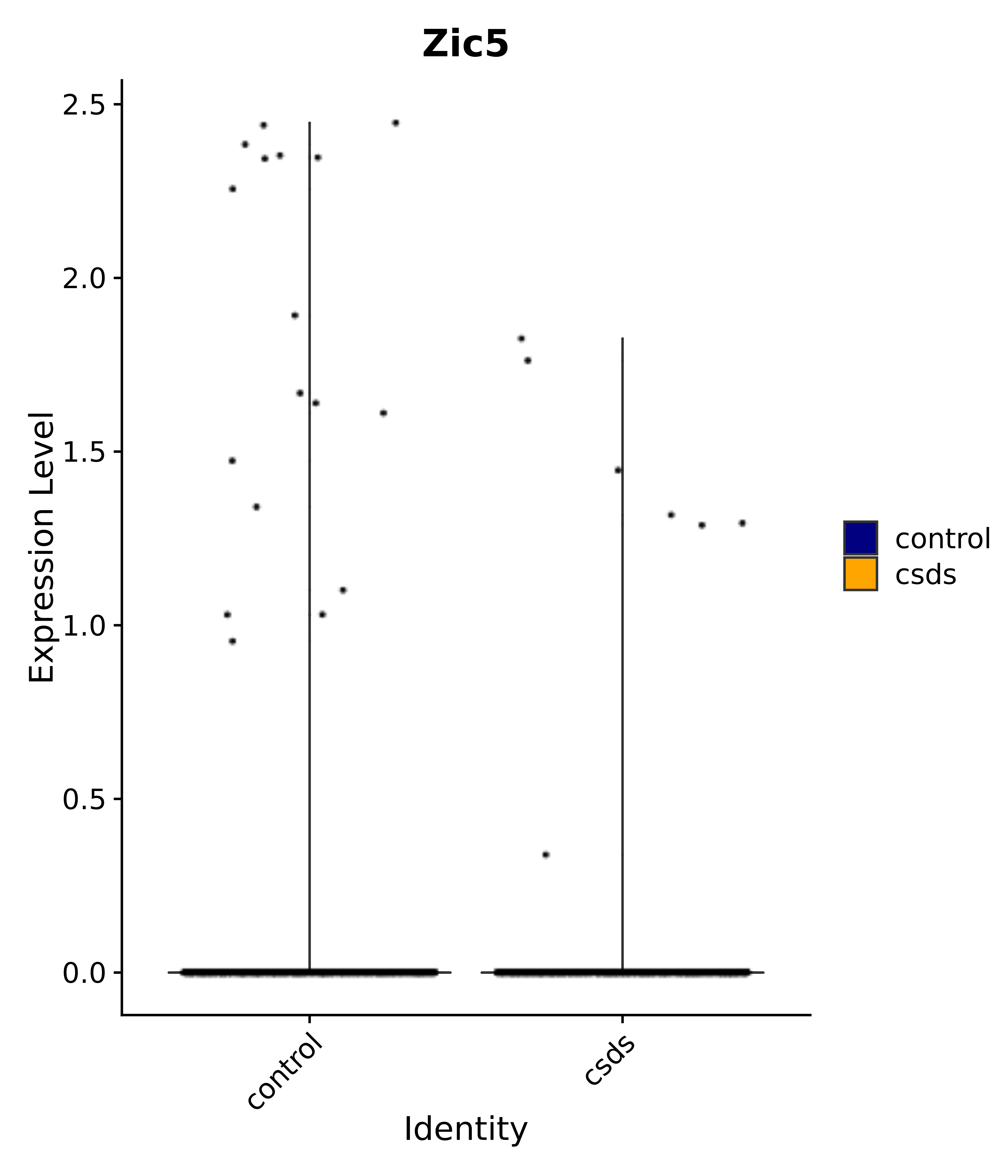
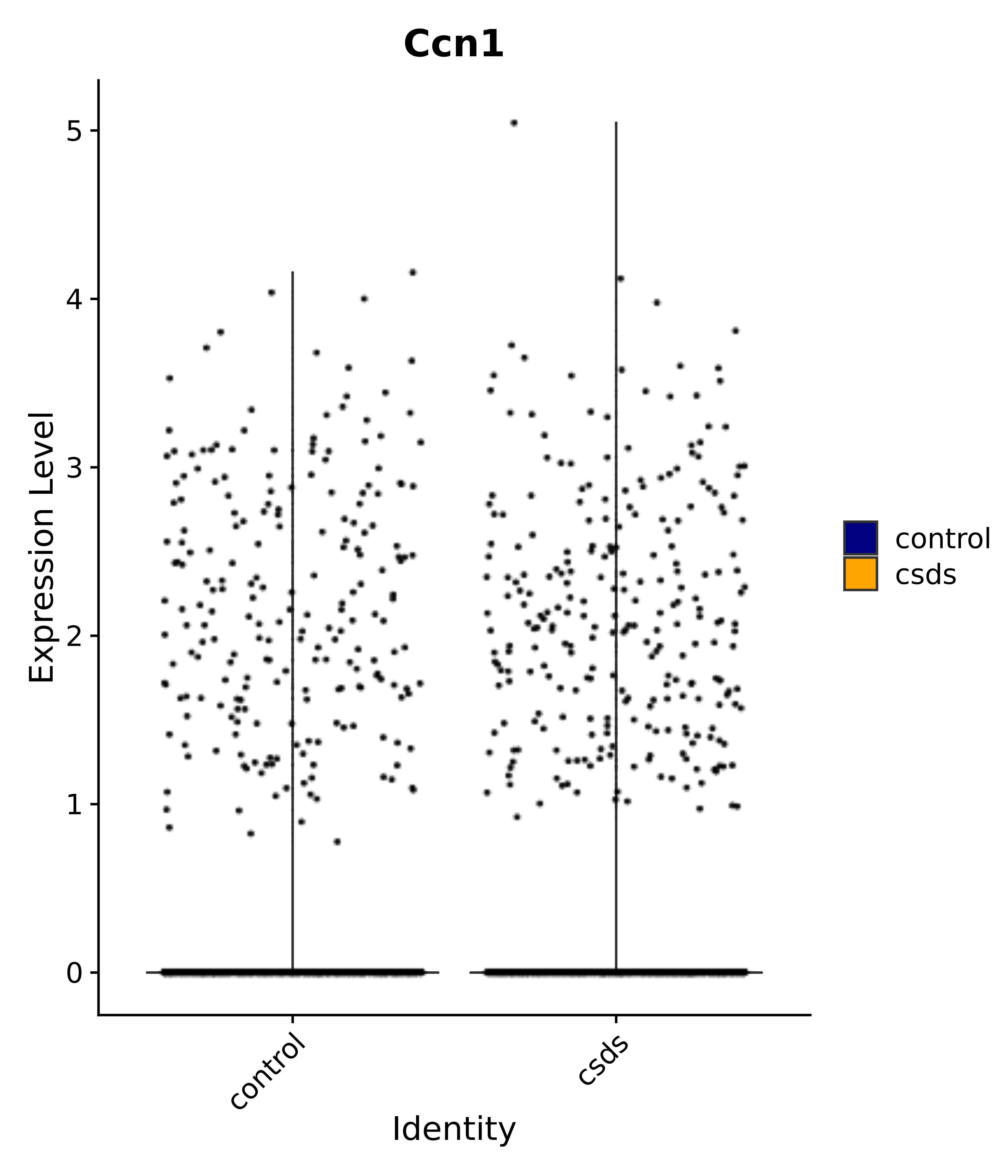
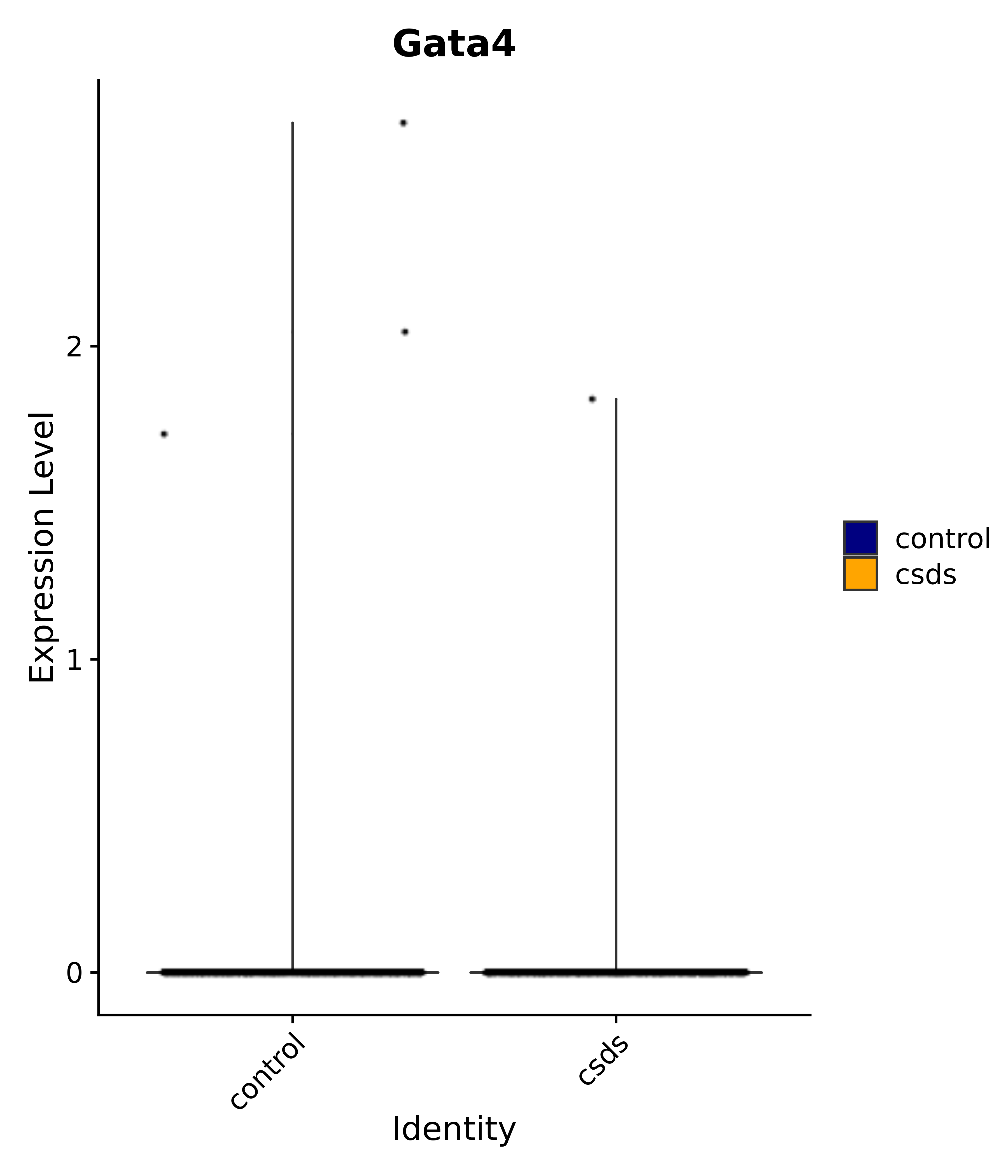
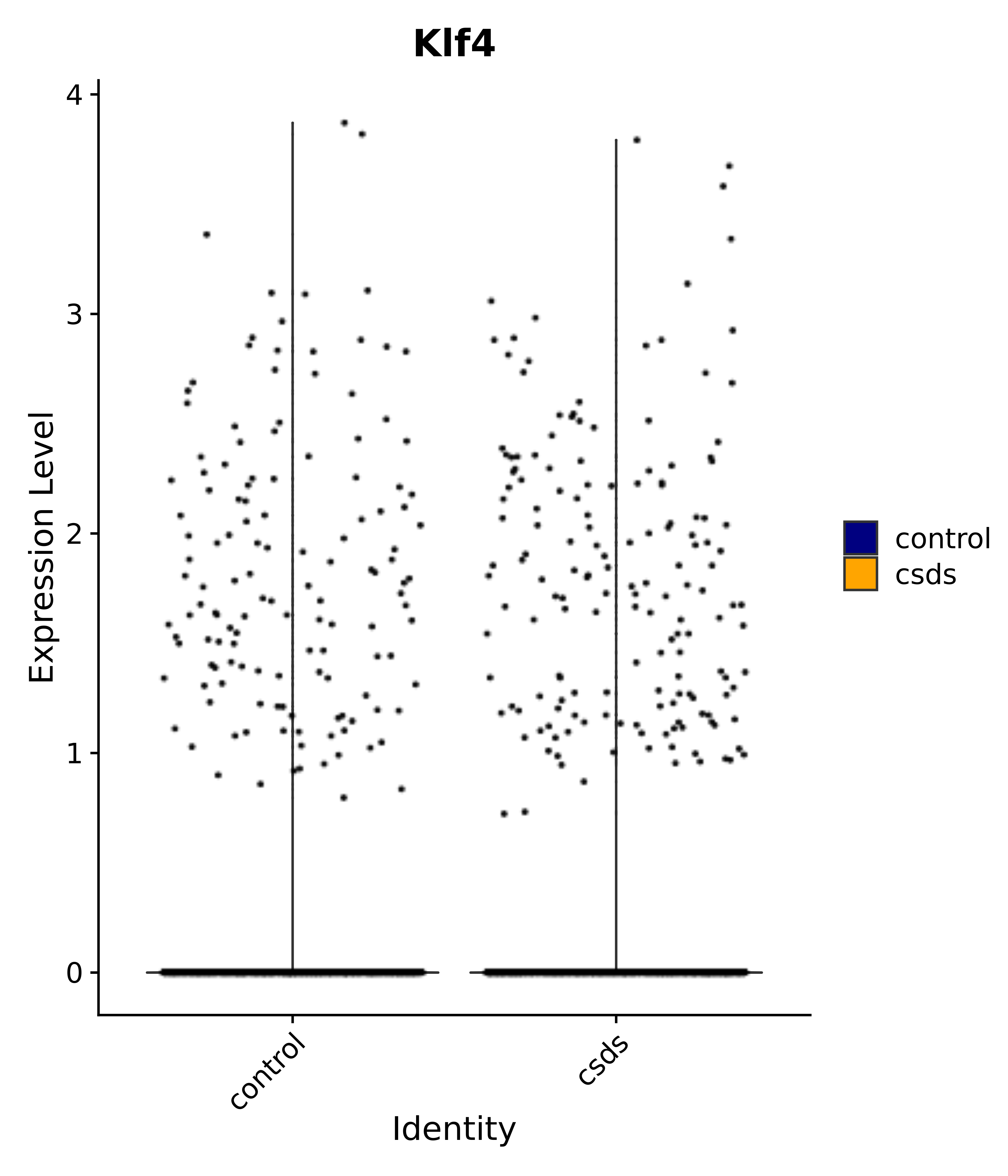
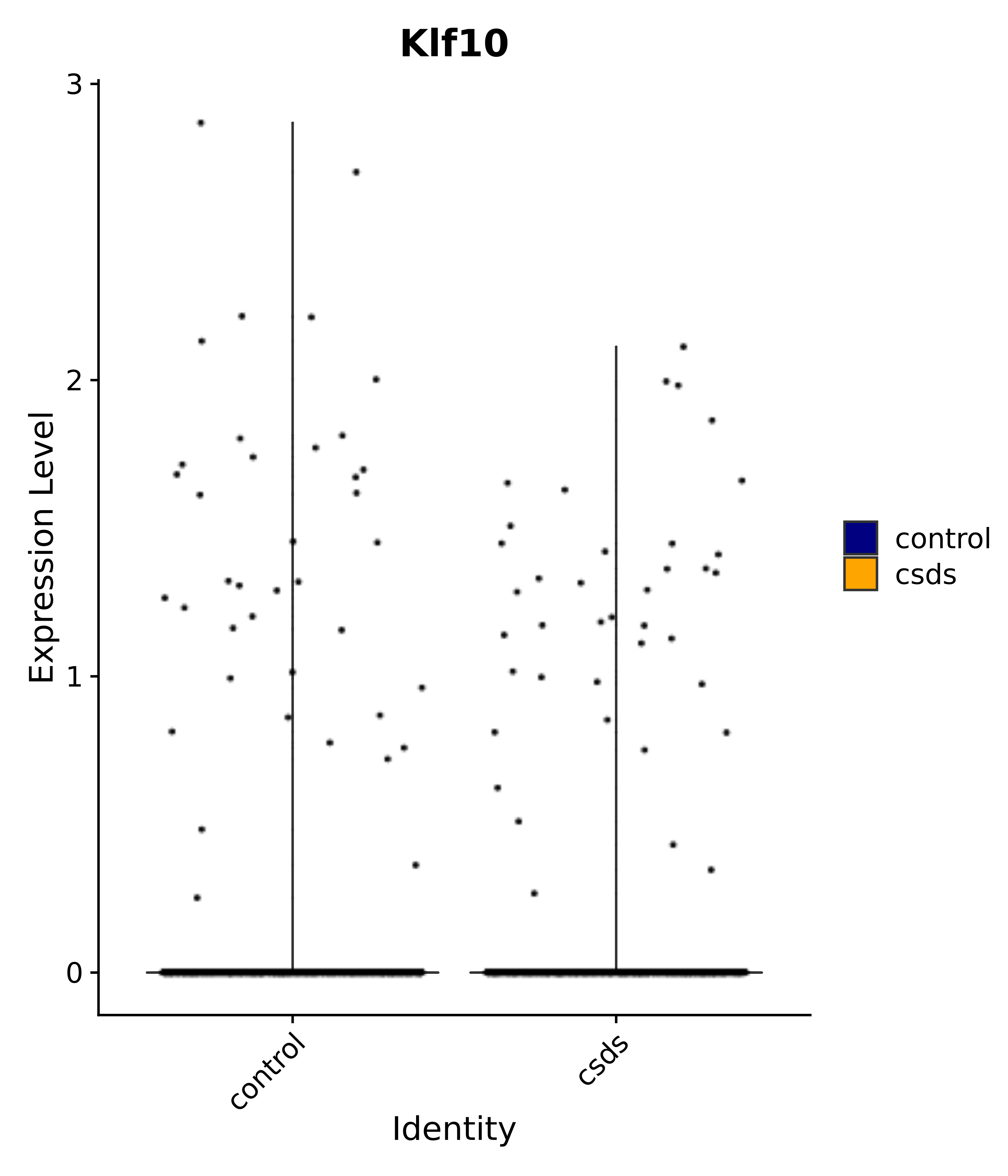
"Rfx4", "Dbx2", "Prokr2", "Cebpb", "Zic1", "Zic2", "Zic5", "Ccn1", "Gata4", "Klf4", "Klf10", # 5 SCN
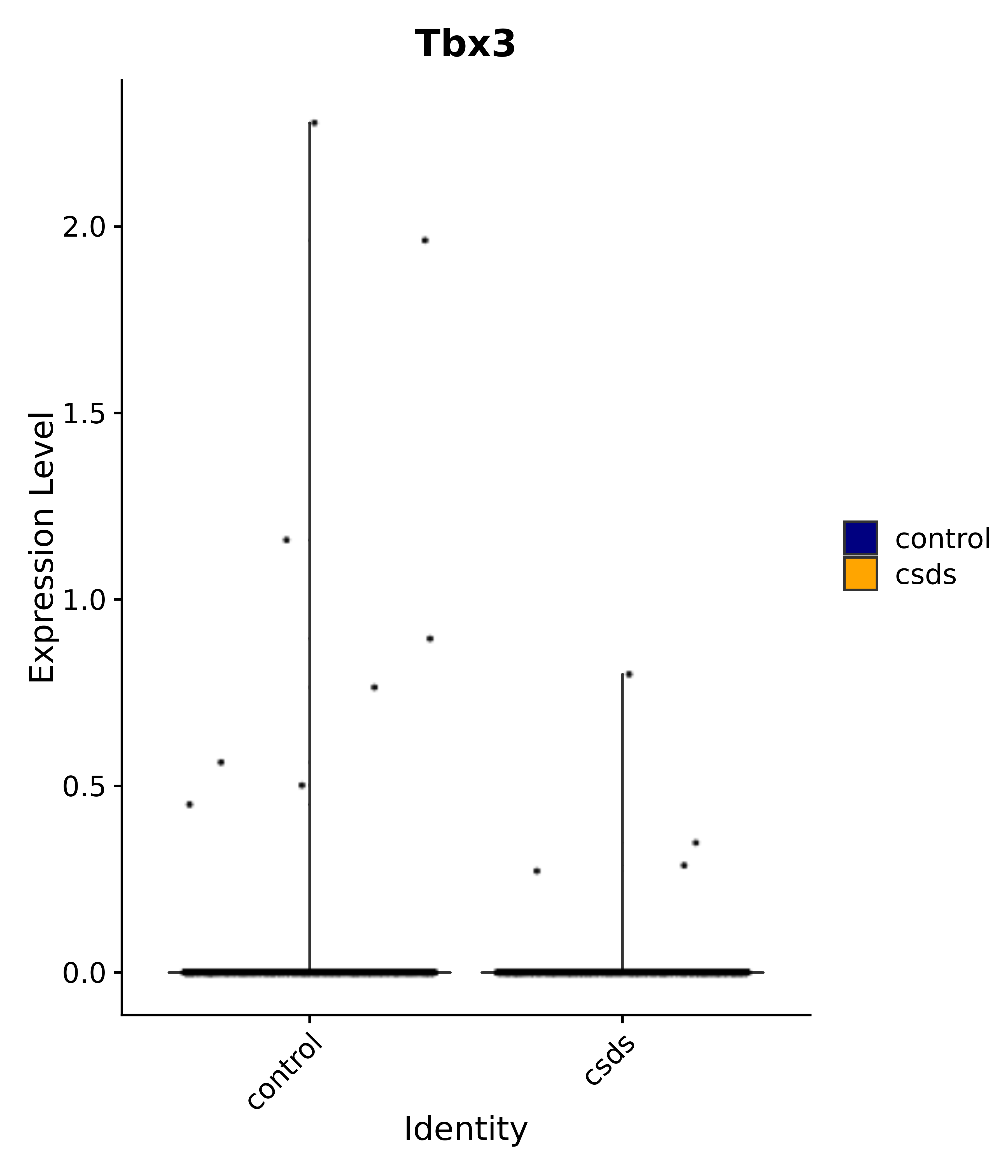
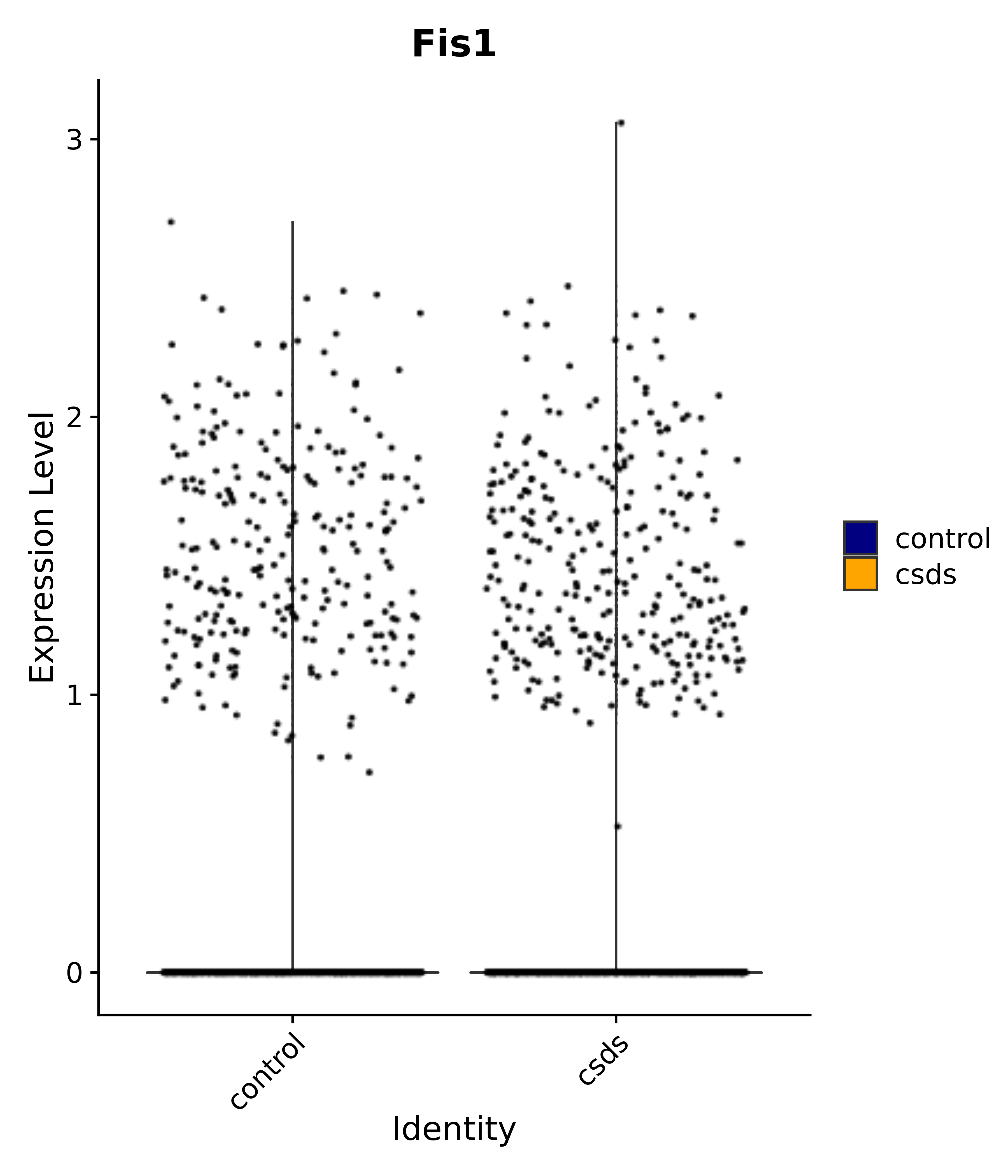
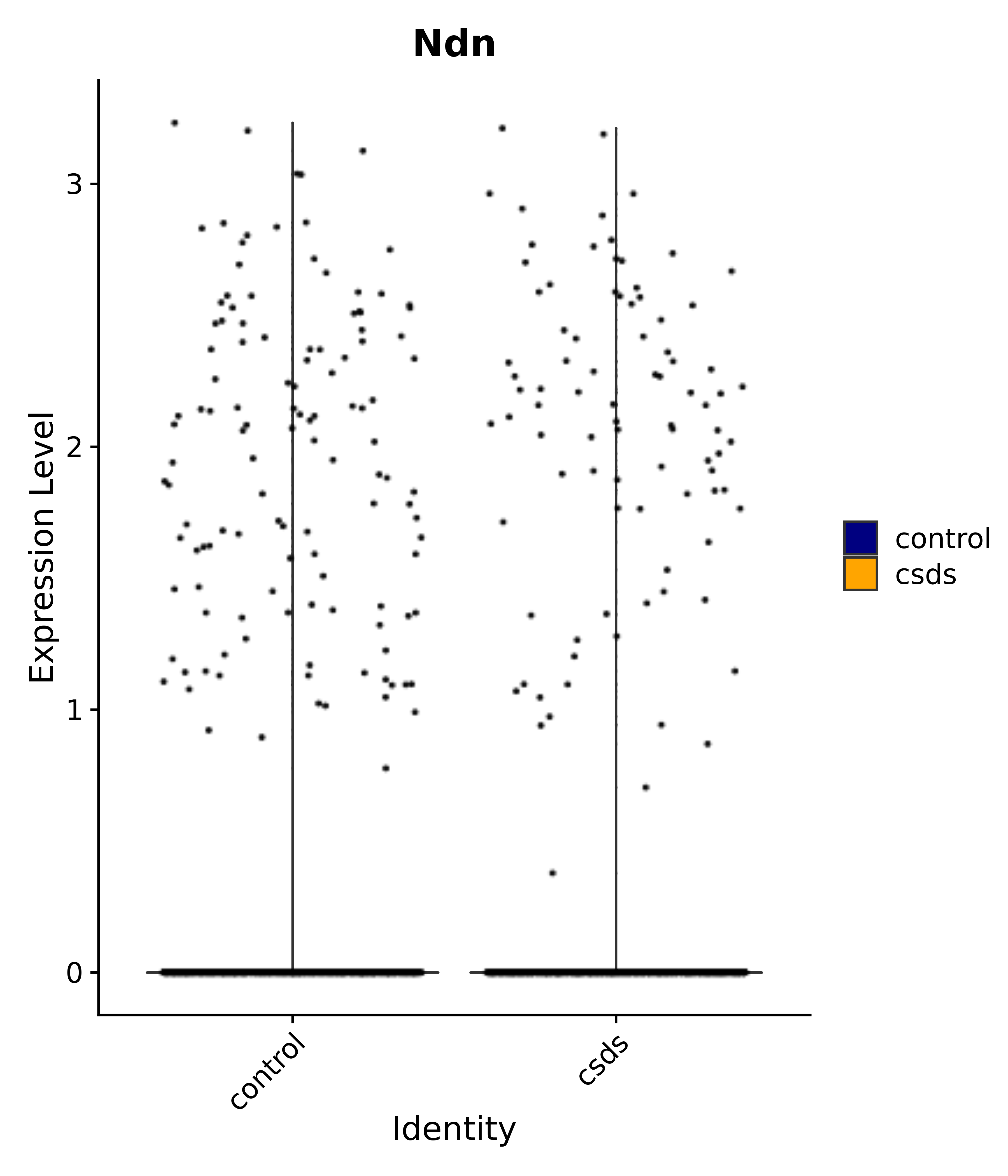
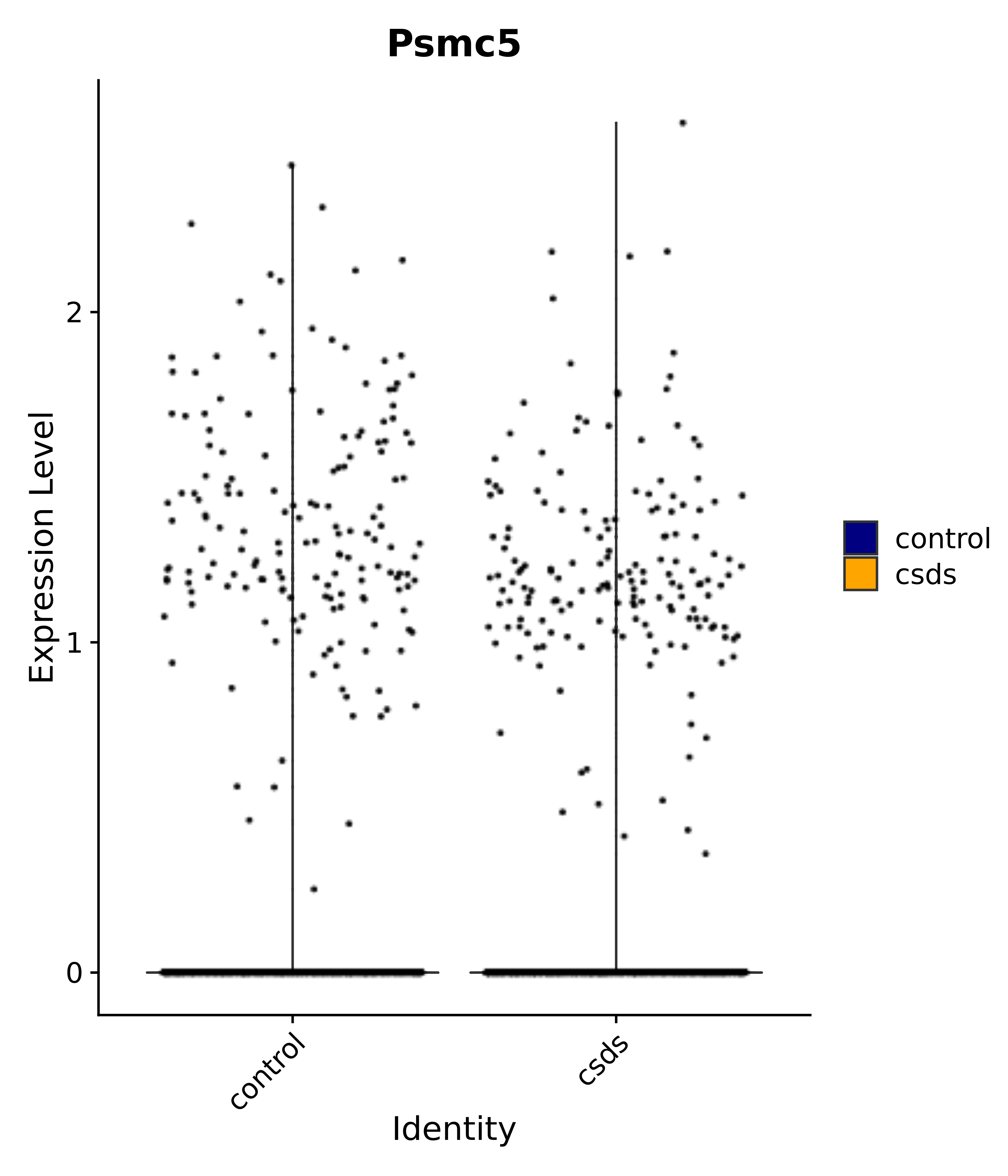
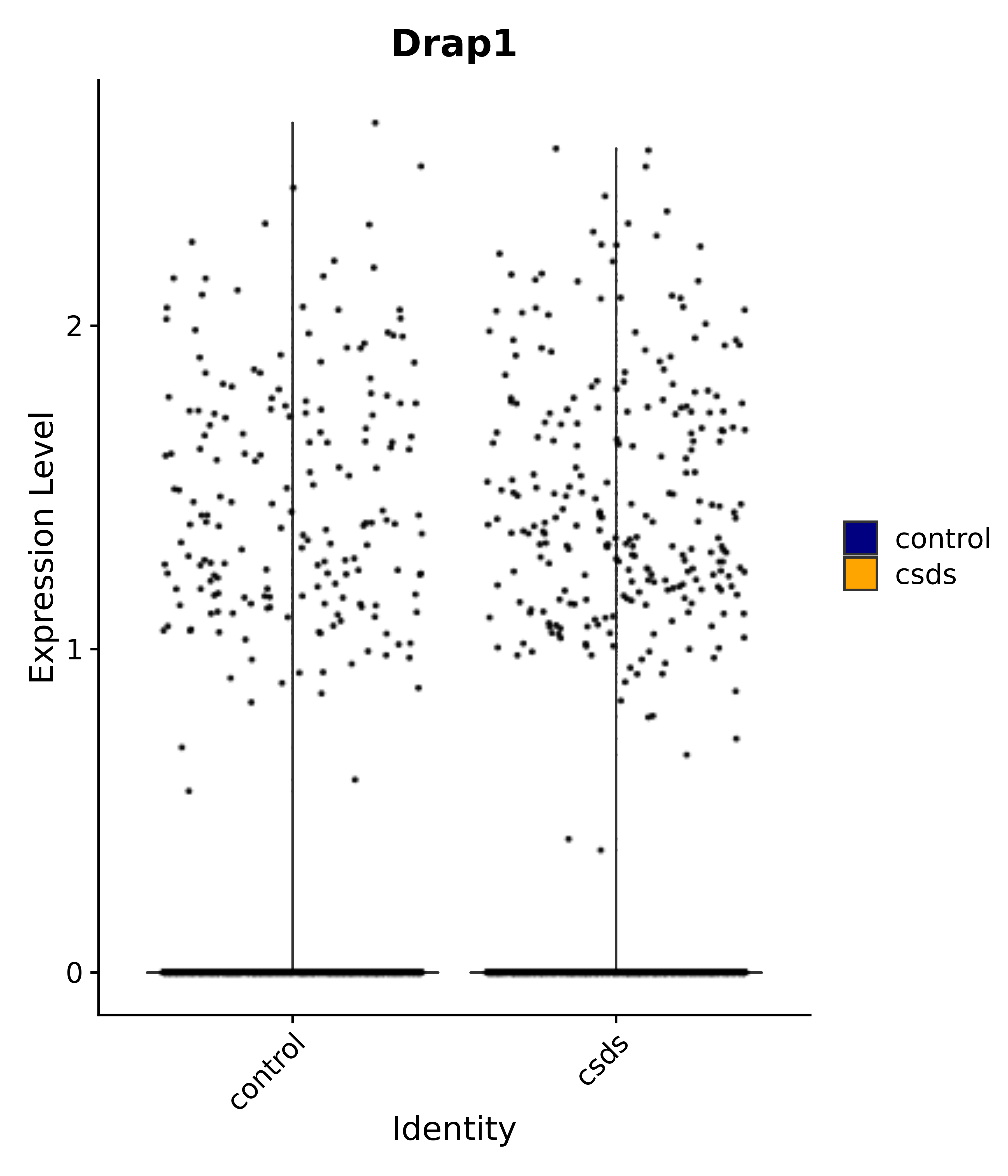
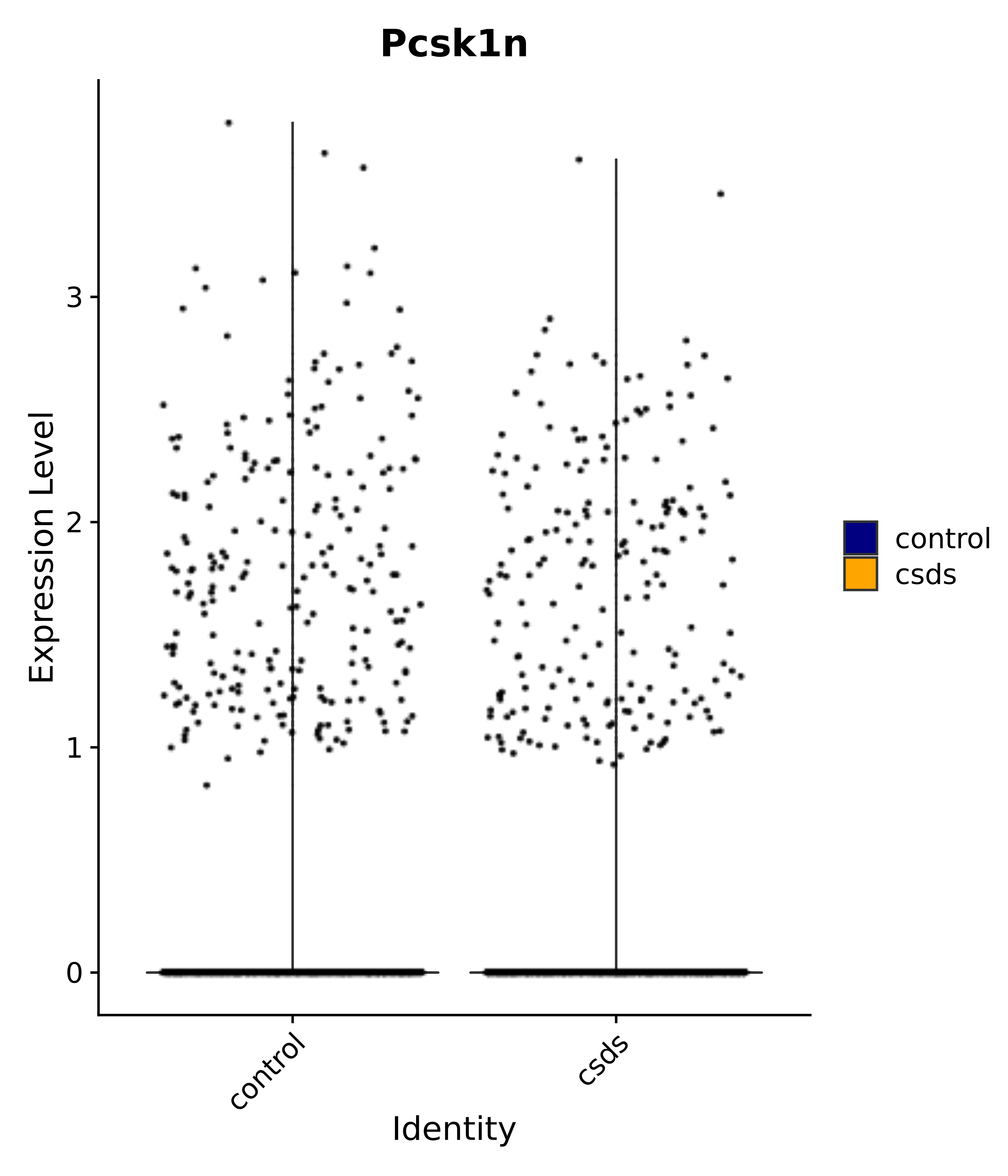
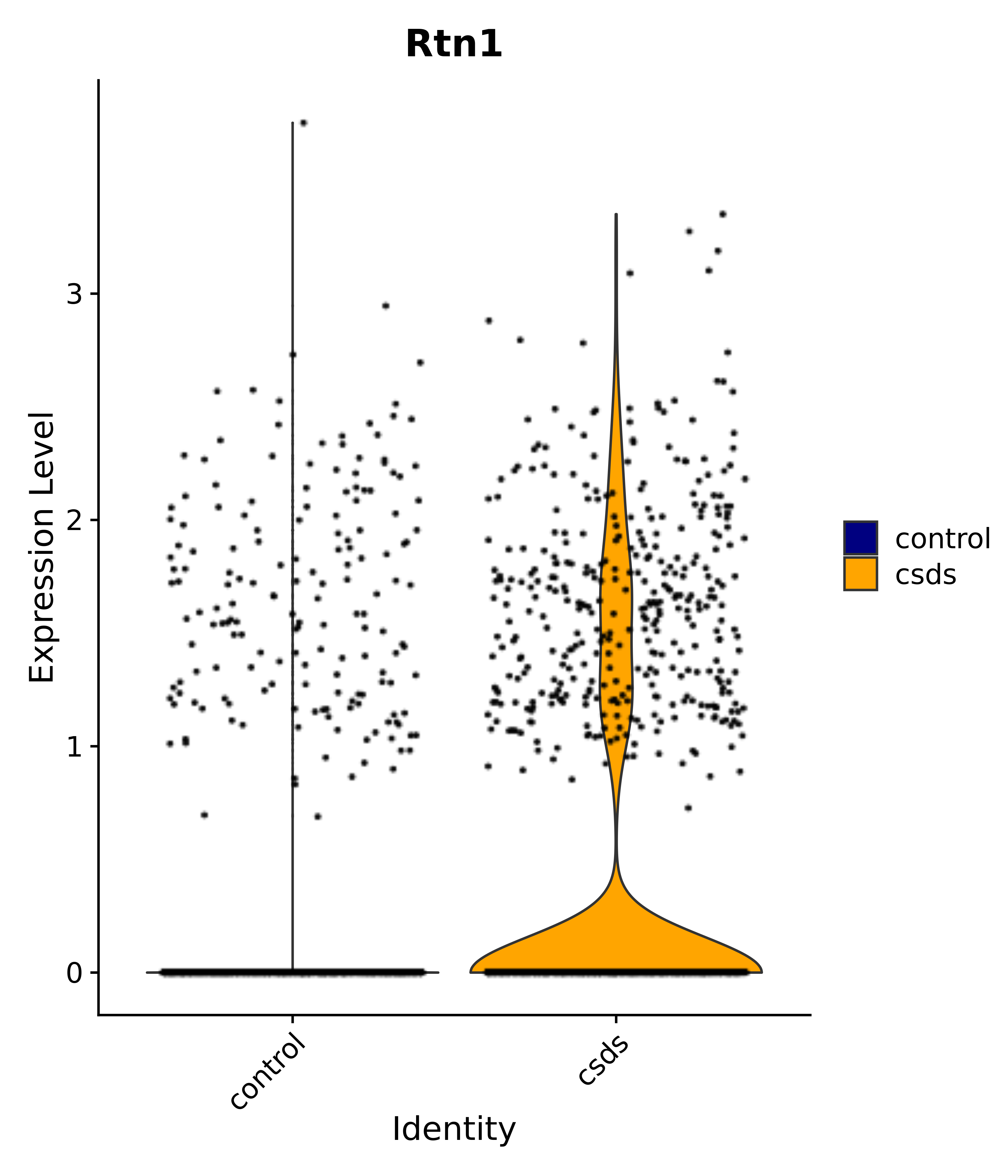
"Tbx3", "Fis1", "Ndn", "Psmc5", "Drap1", "Pcsk1n", "Rtn1", # 6 VMH
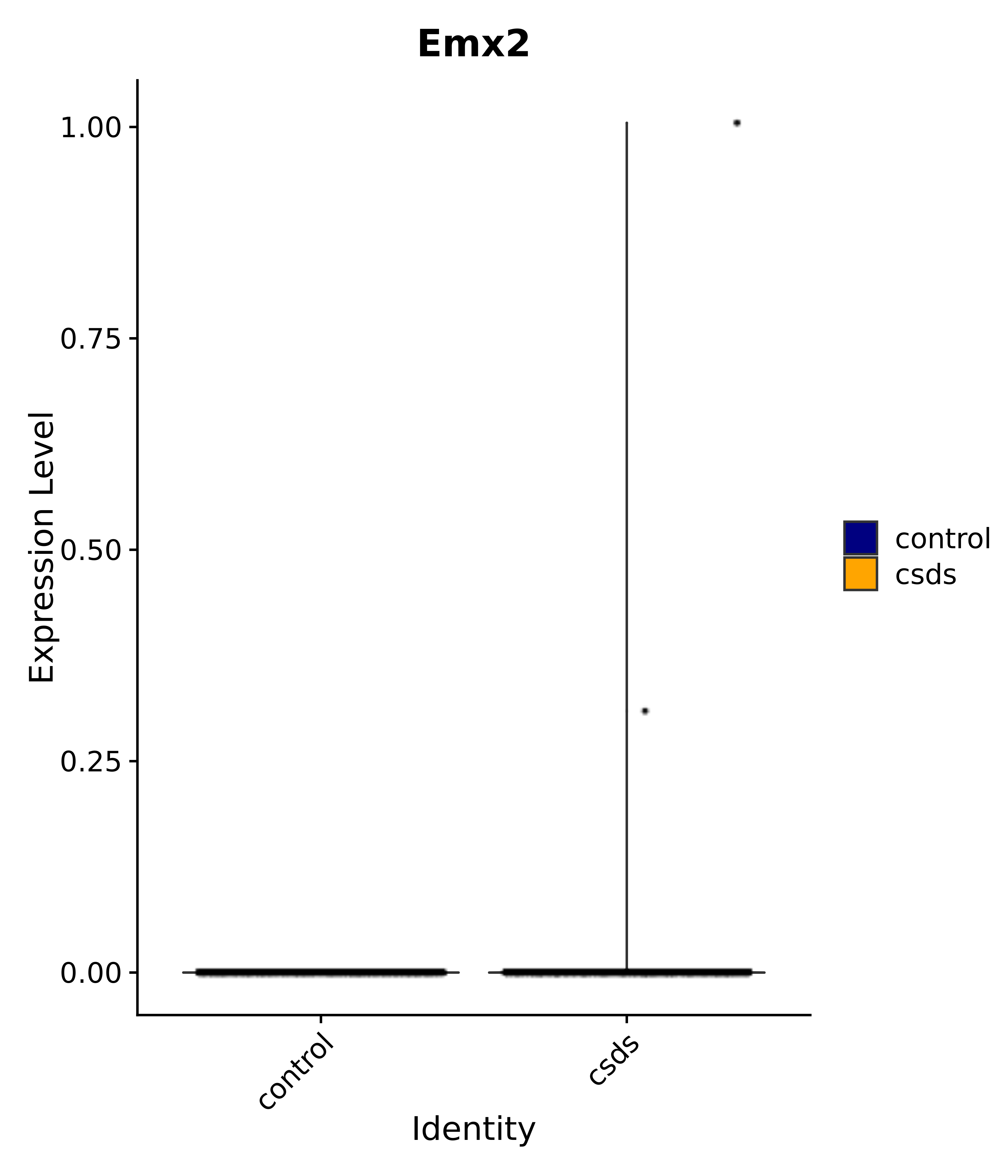
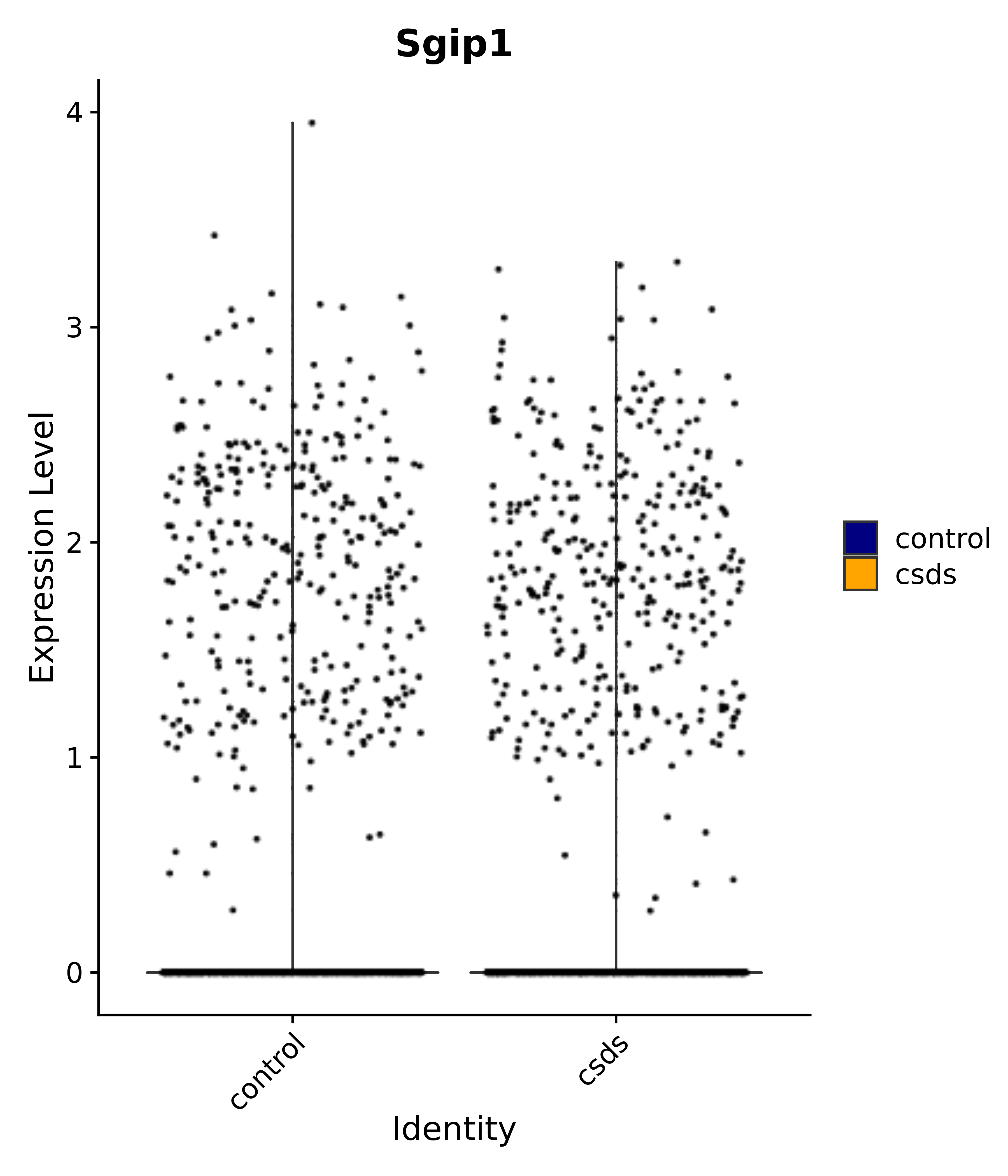
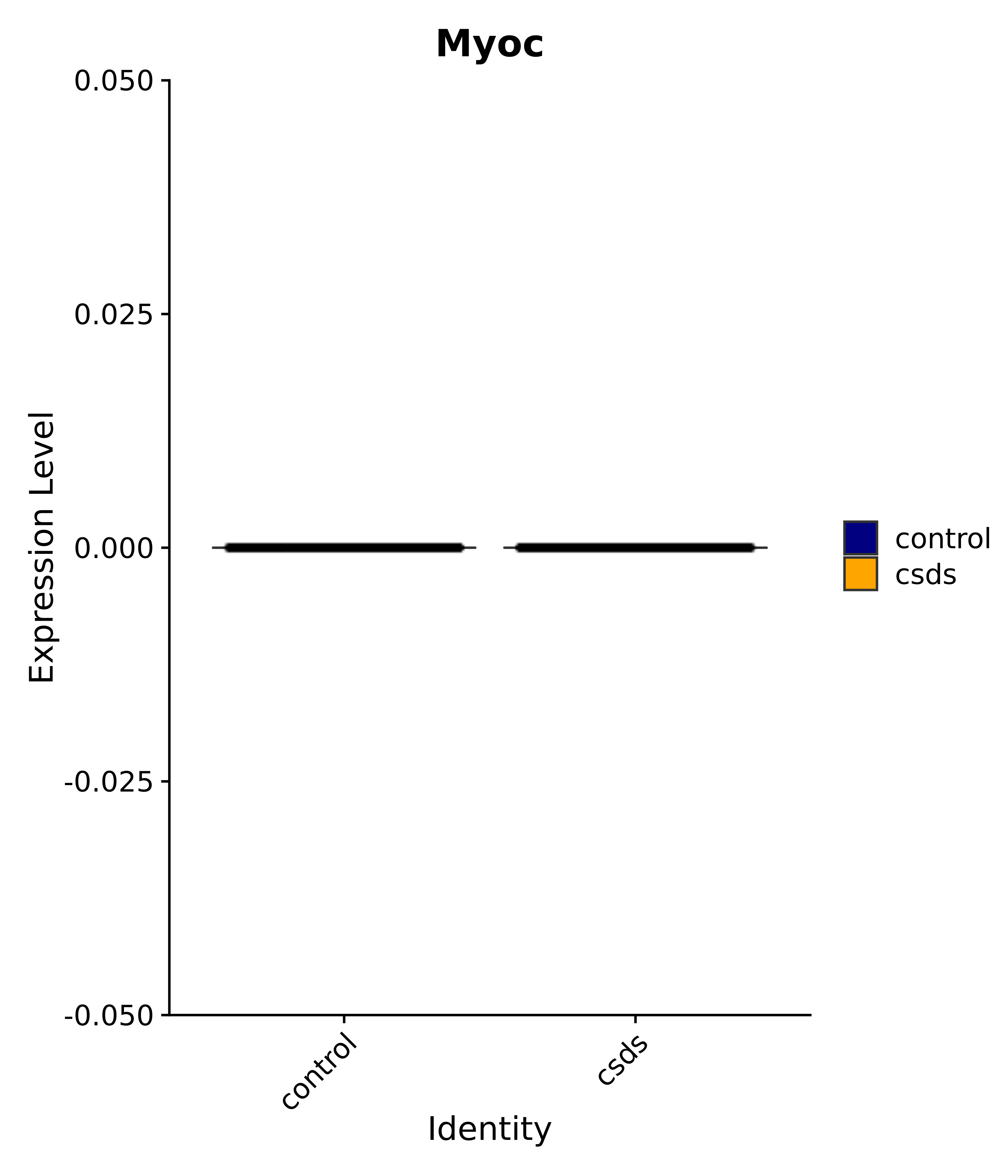
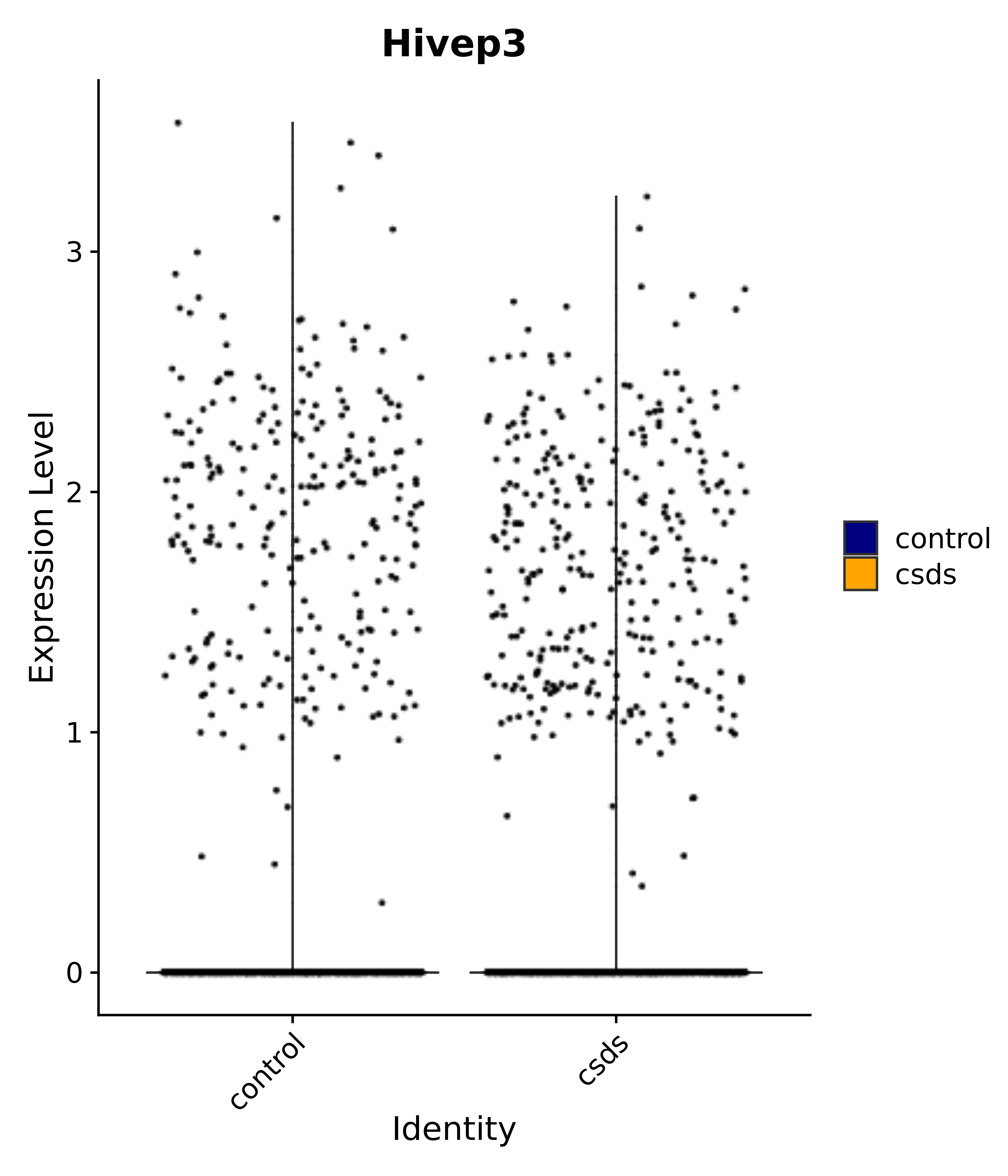
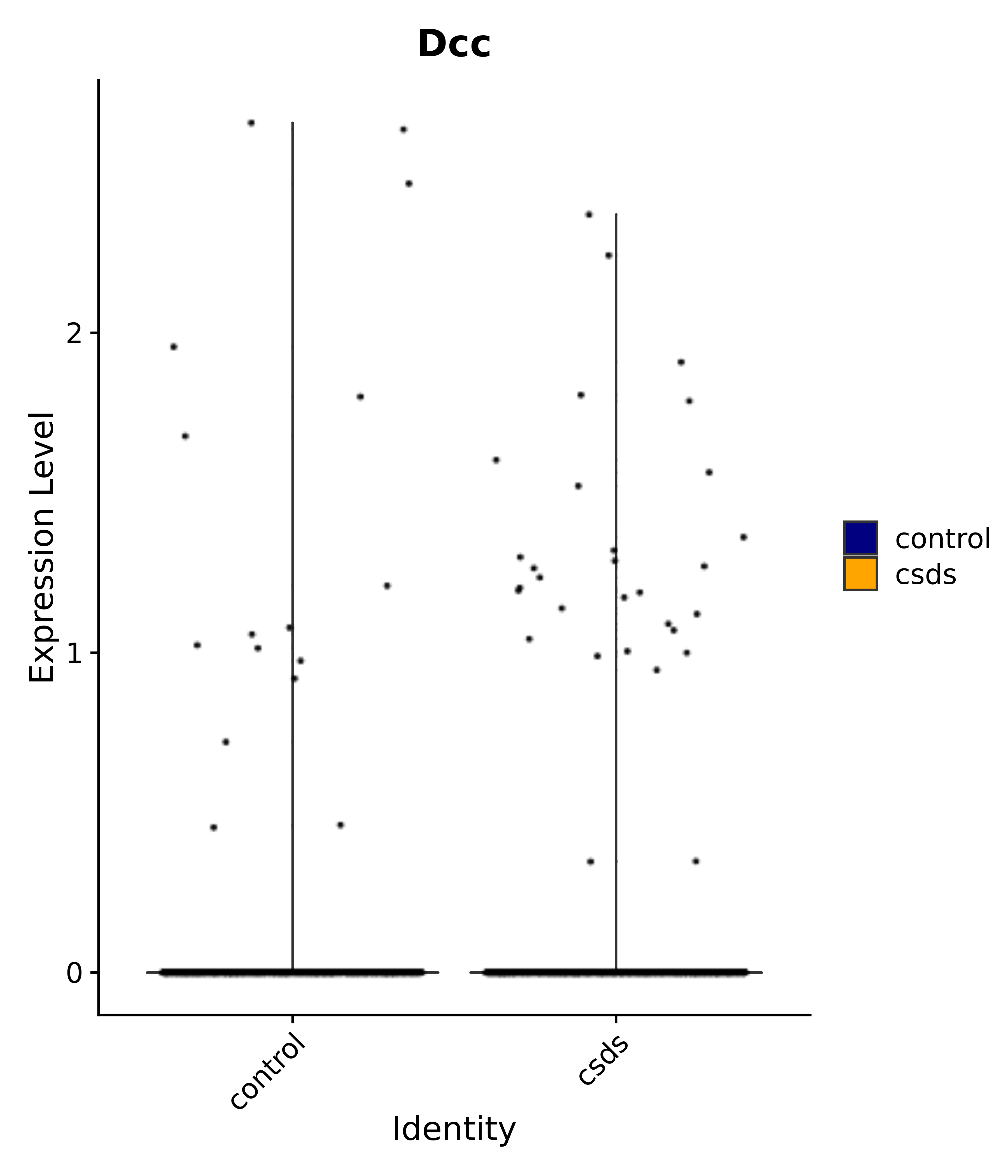
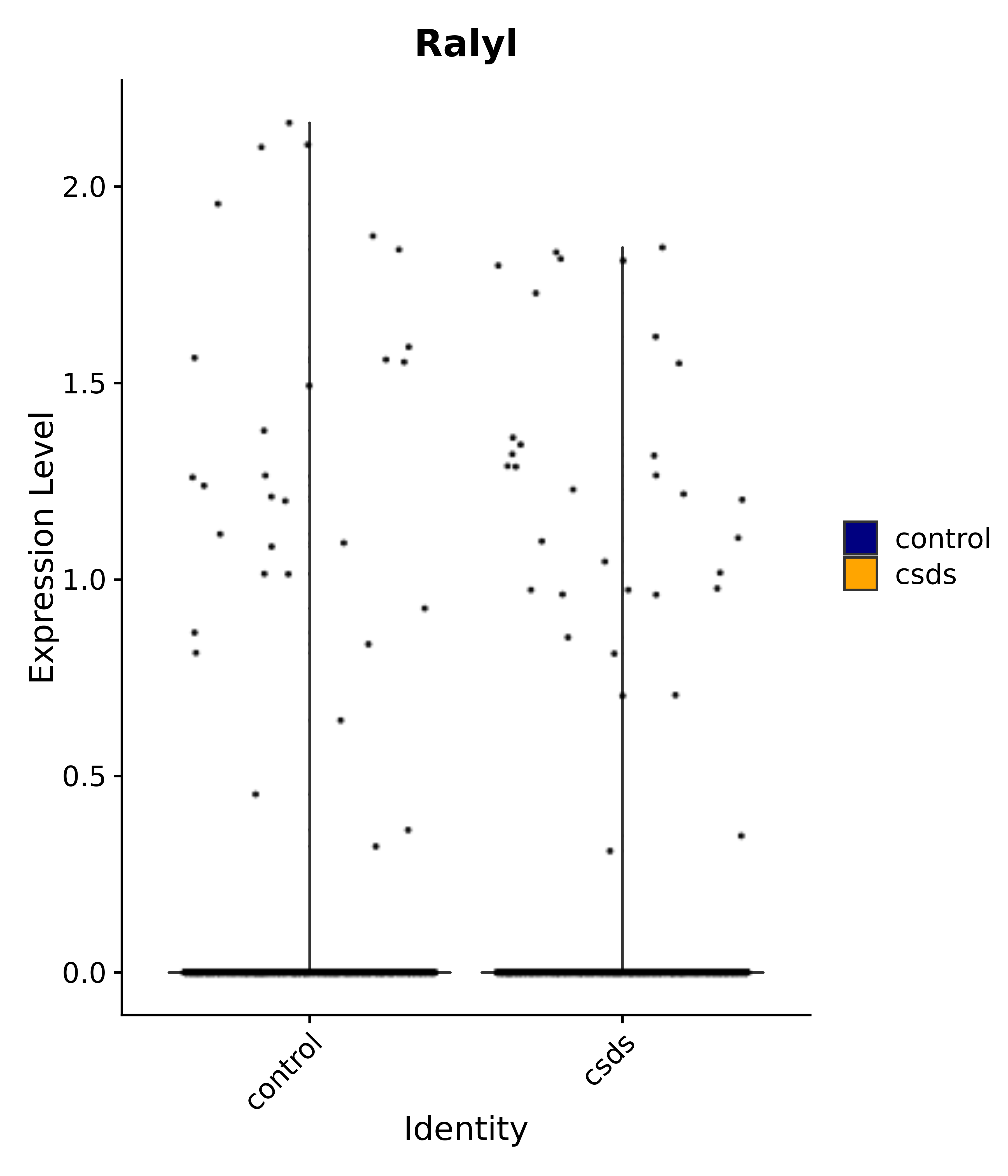
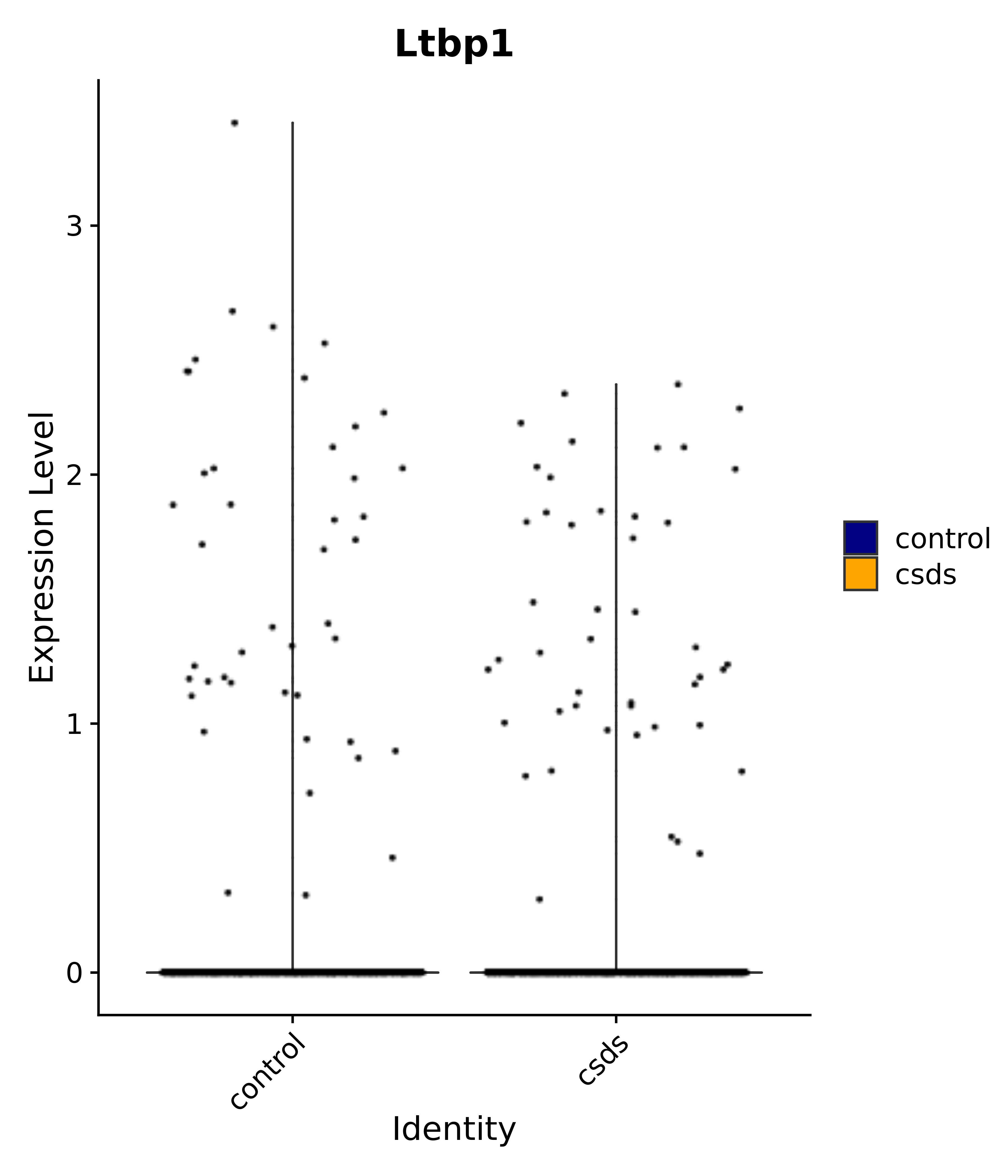
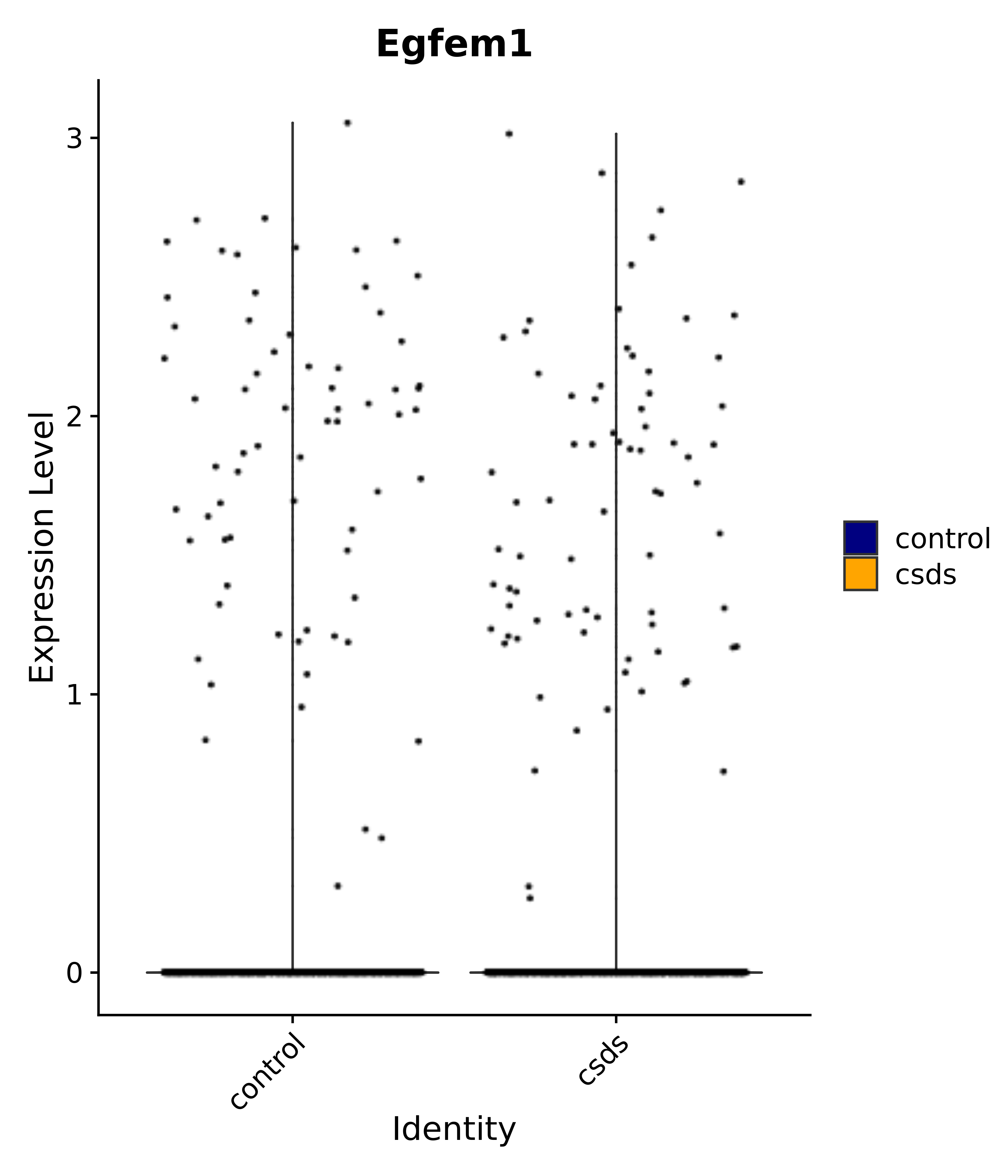
"Emx2", "Sgip1", "Myoc", "Hivep3", "Dcc", "Ralyl", "Ltbp1", "Egfem1", # 7 VPH
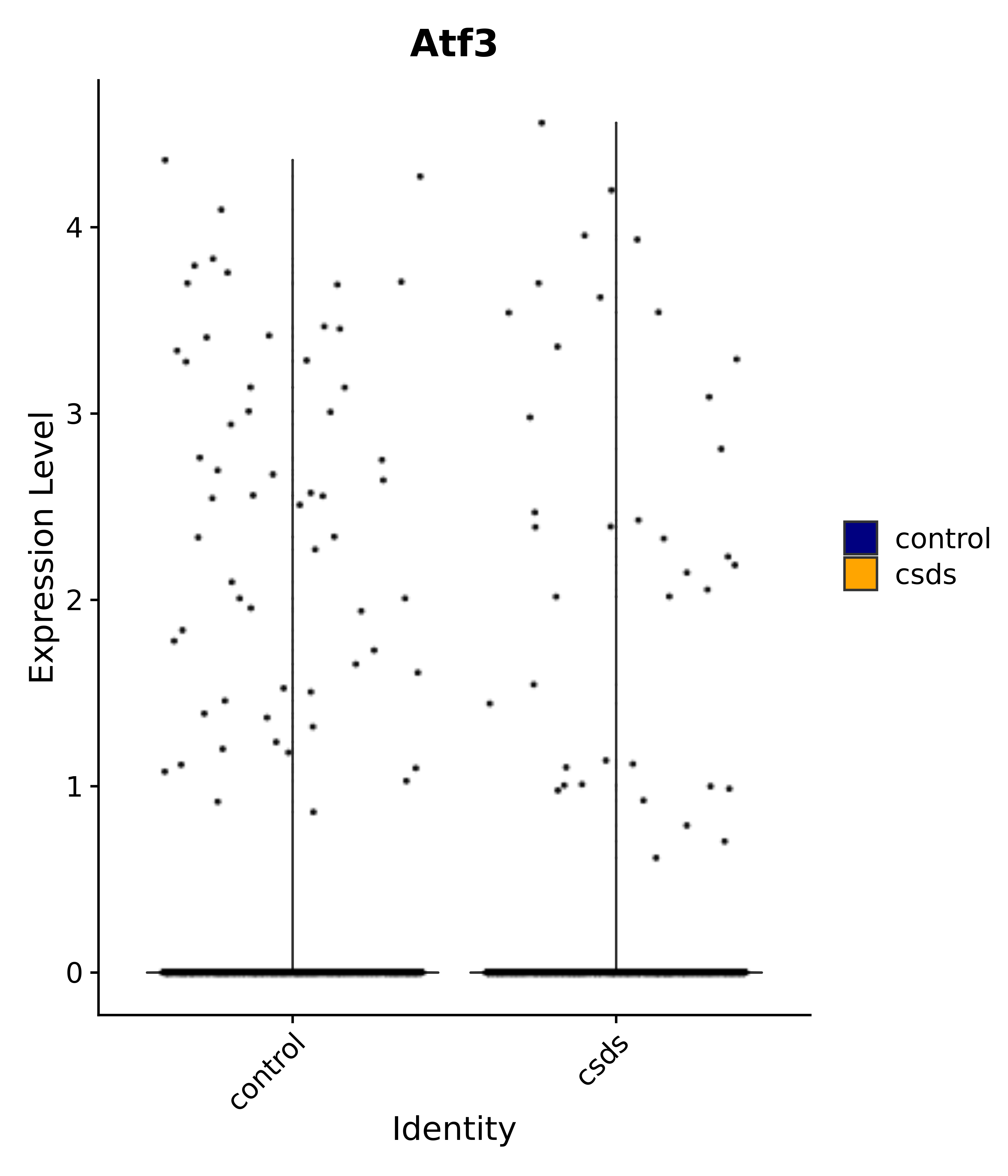
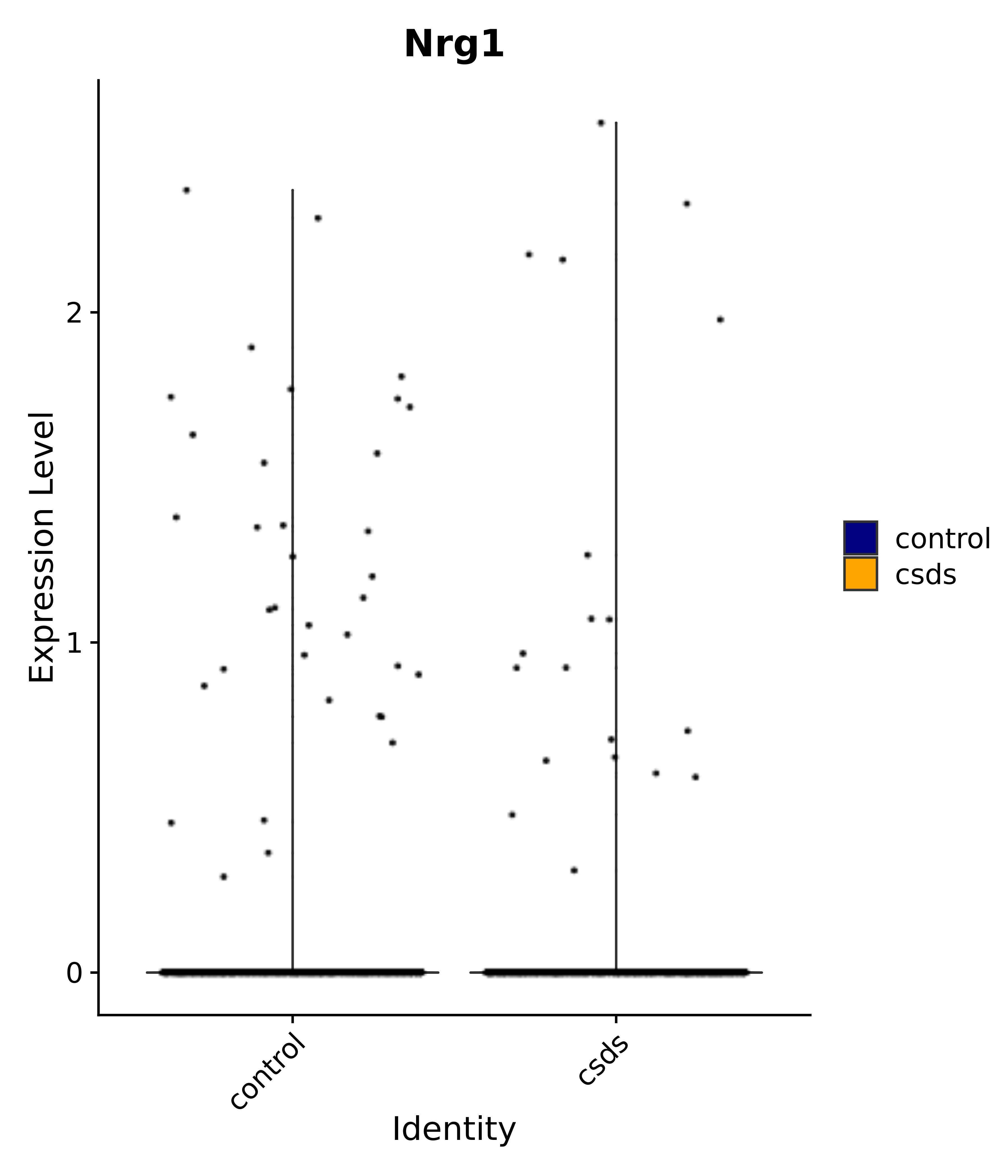
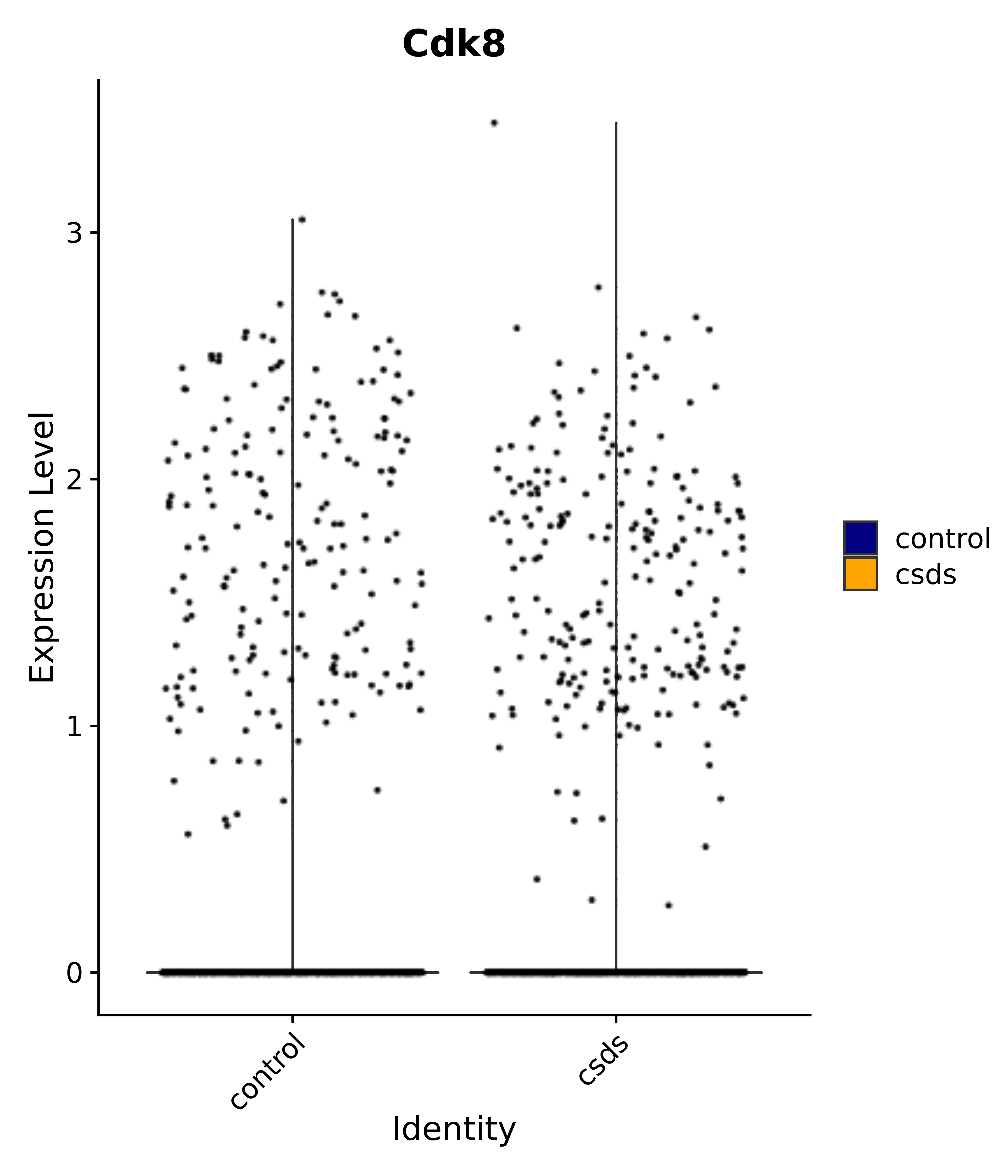
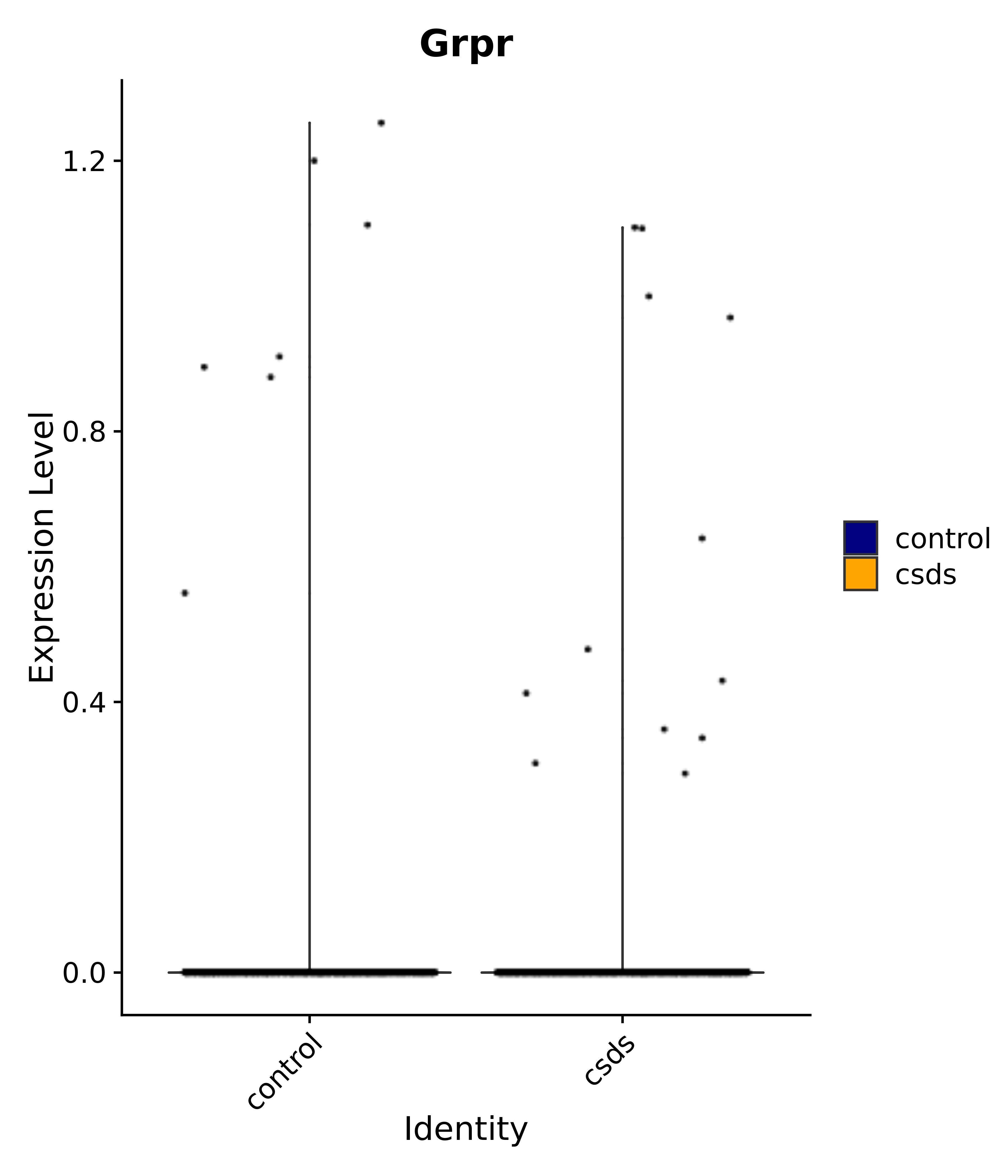
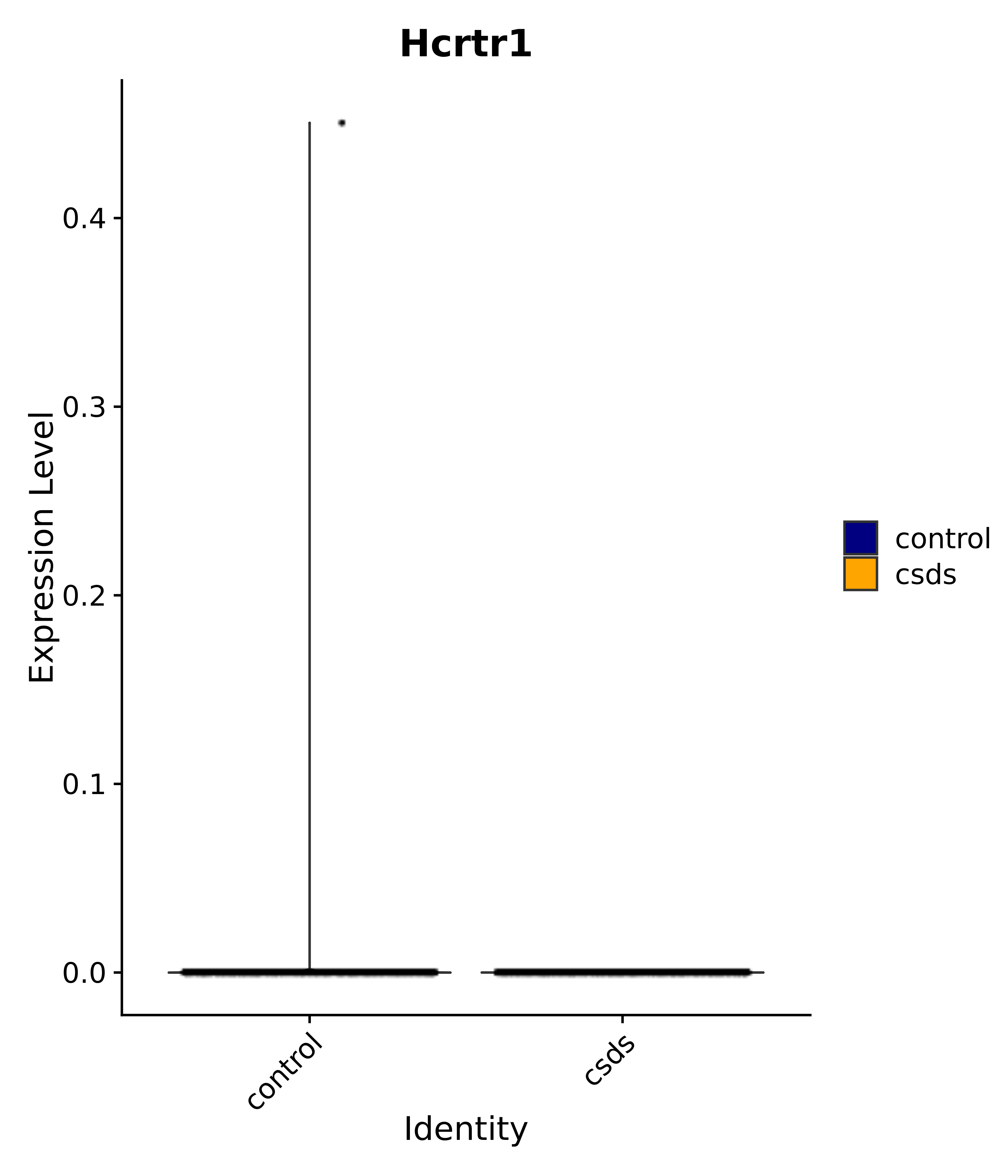
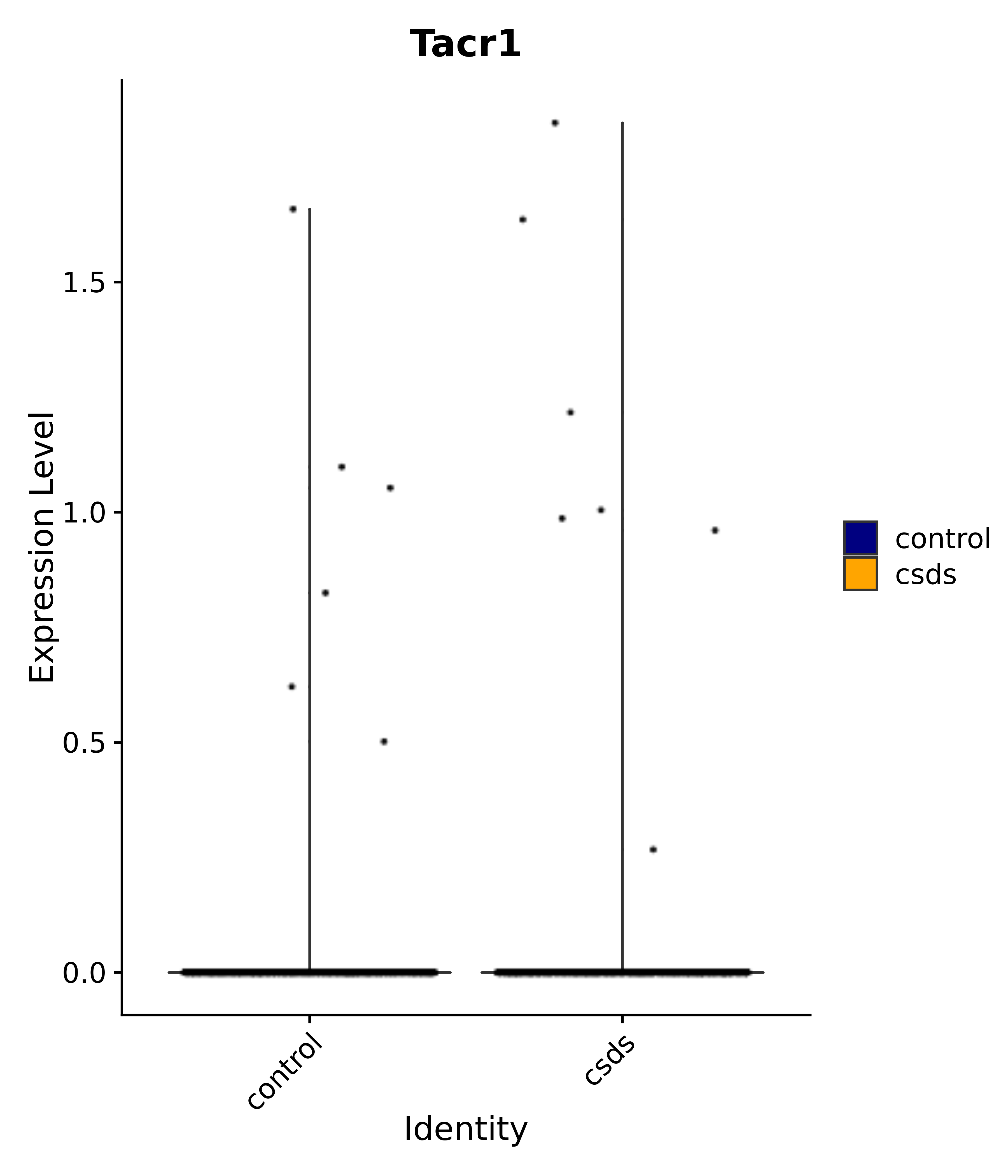
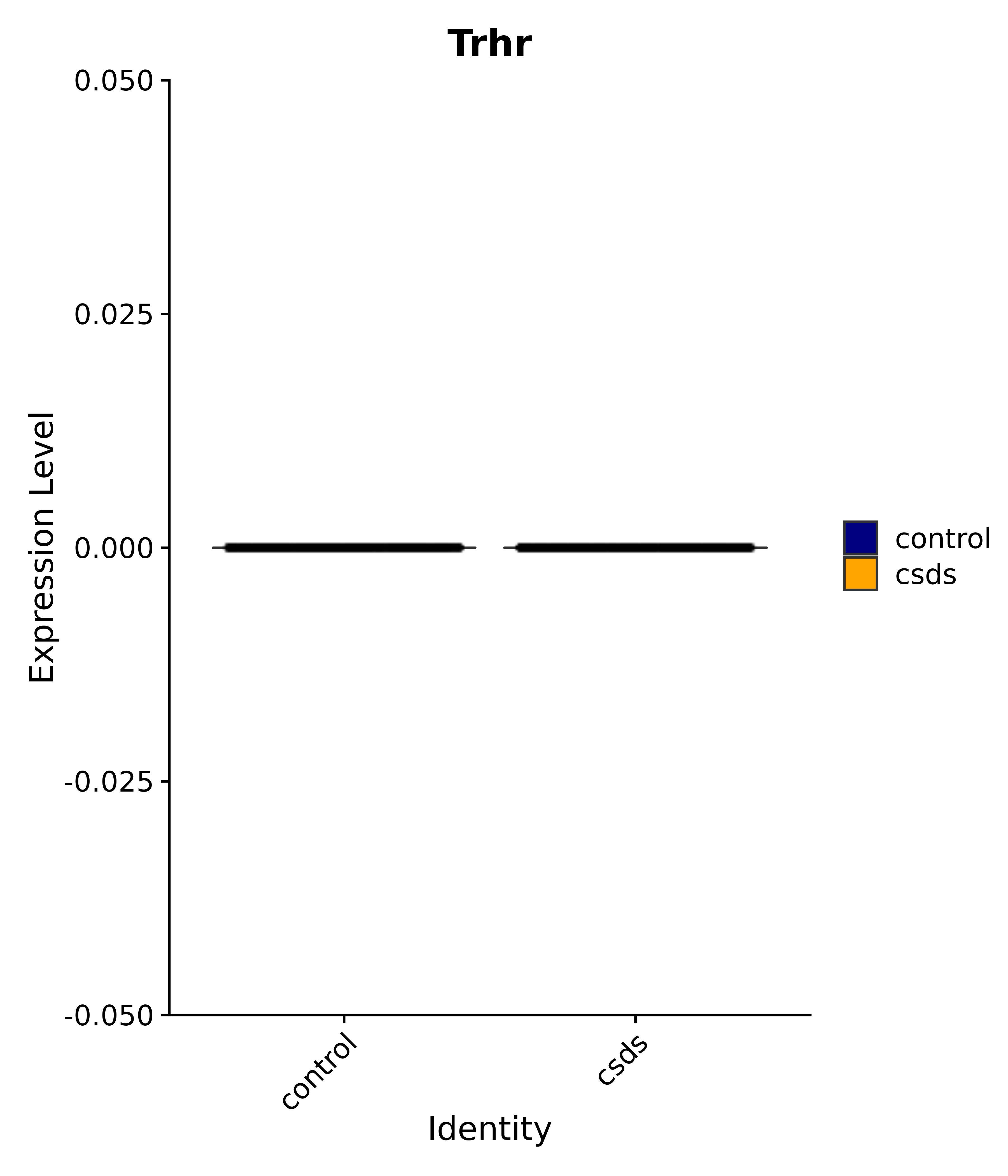
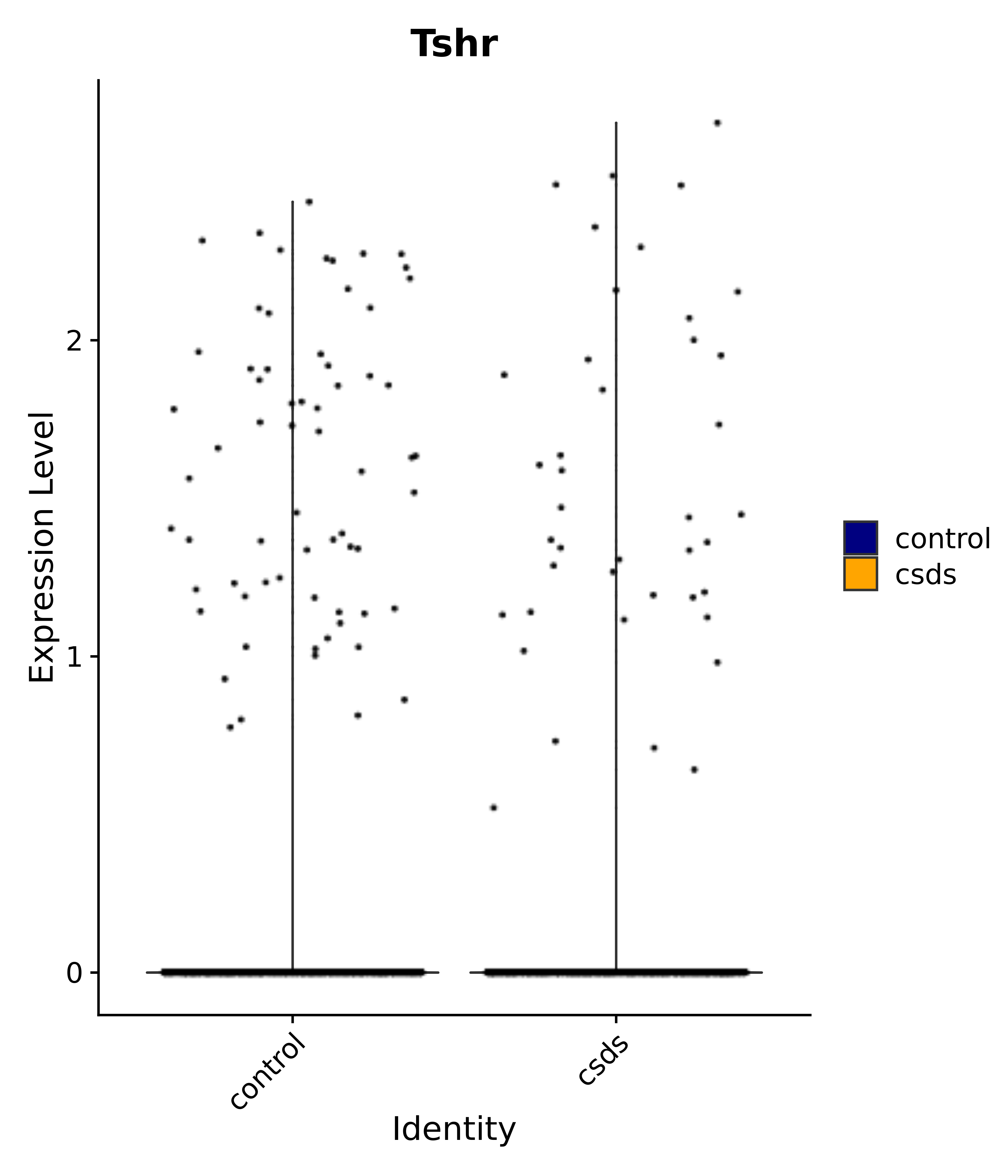
"Klf4", "Atf3", "Nrg1", "Cdk8", "Grpr", "Qrfpr", "Hcrtr1", "Hcrtr2", "Tacr1", "Trhr", "Tshr",
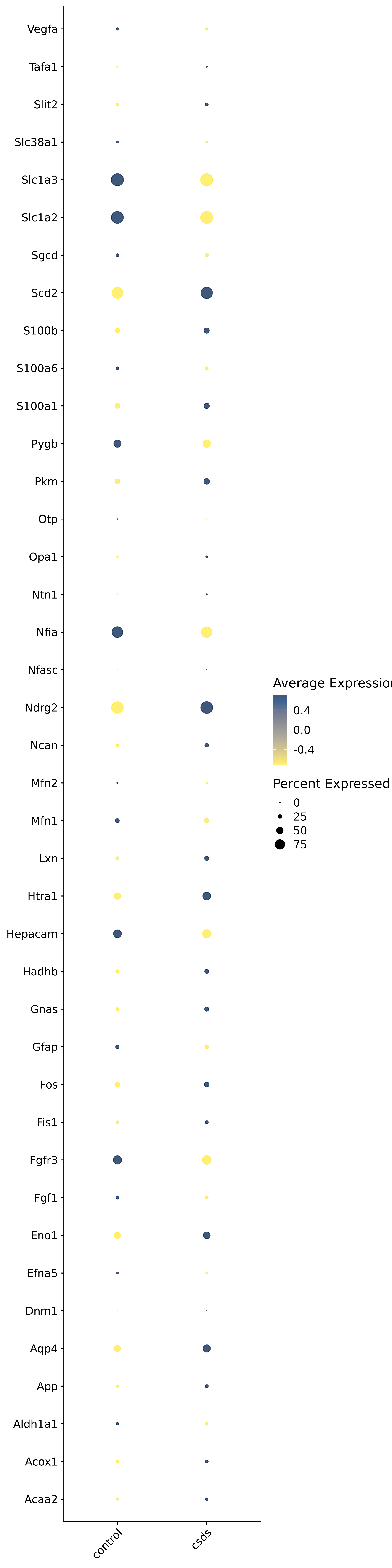
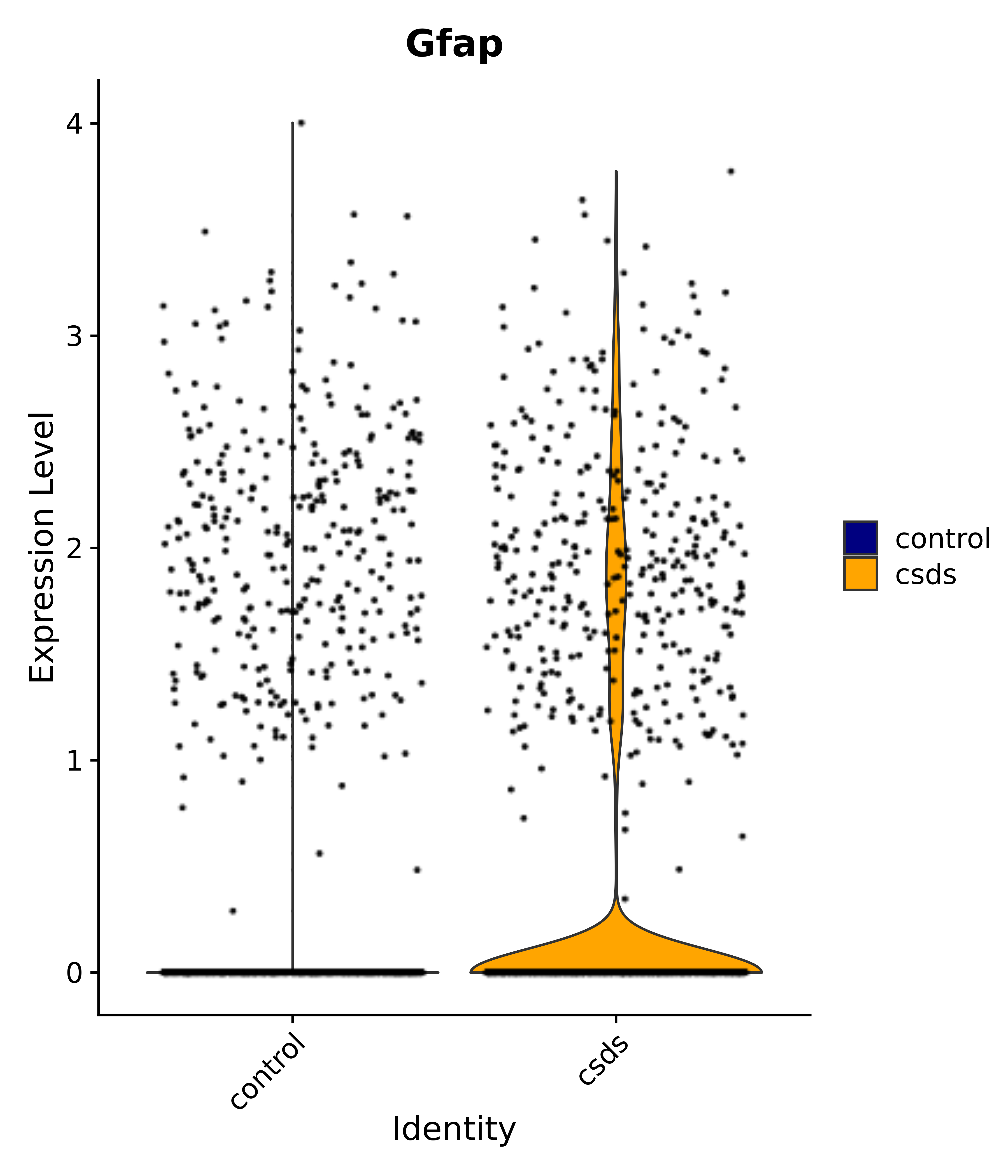
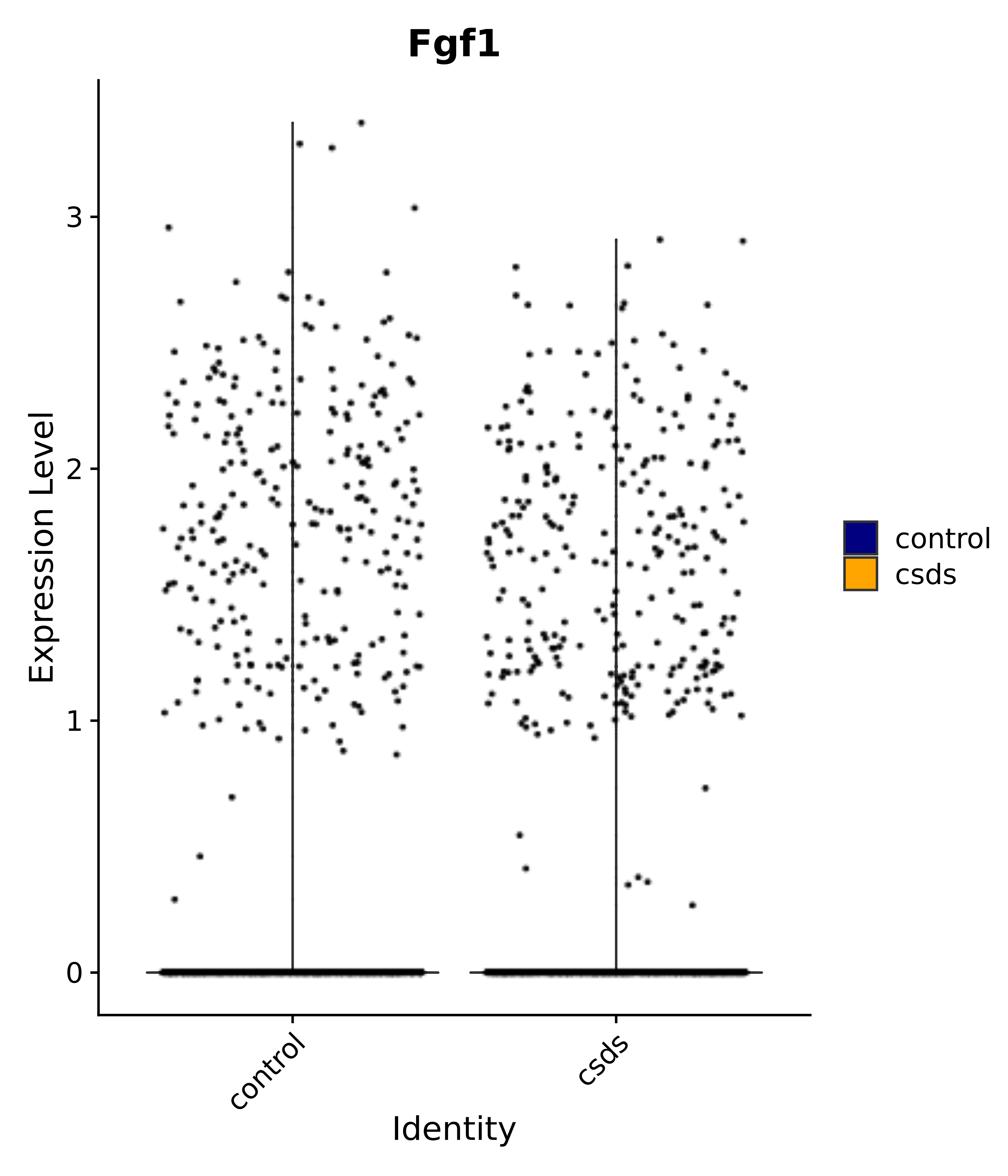
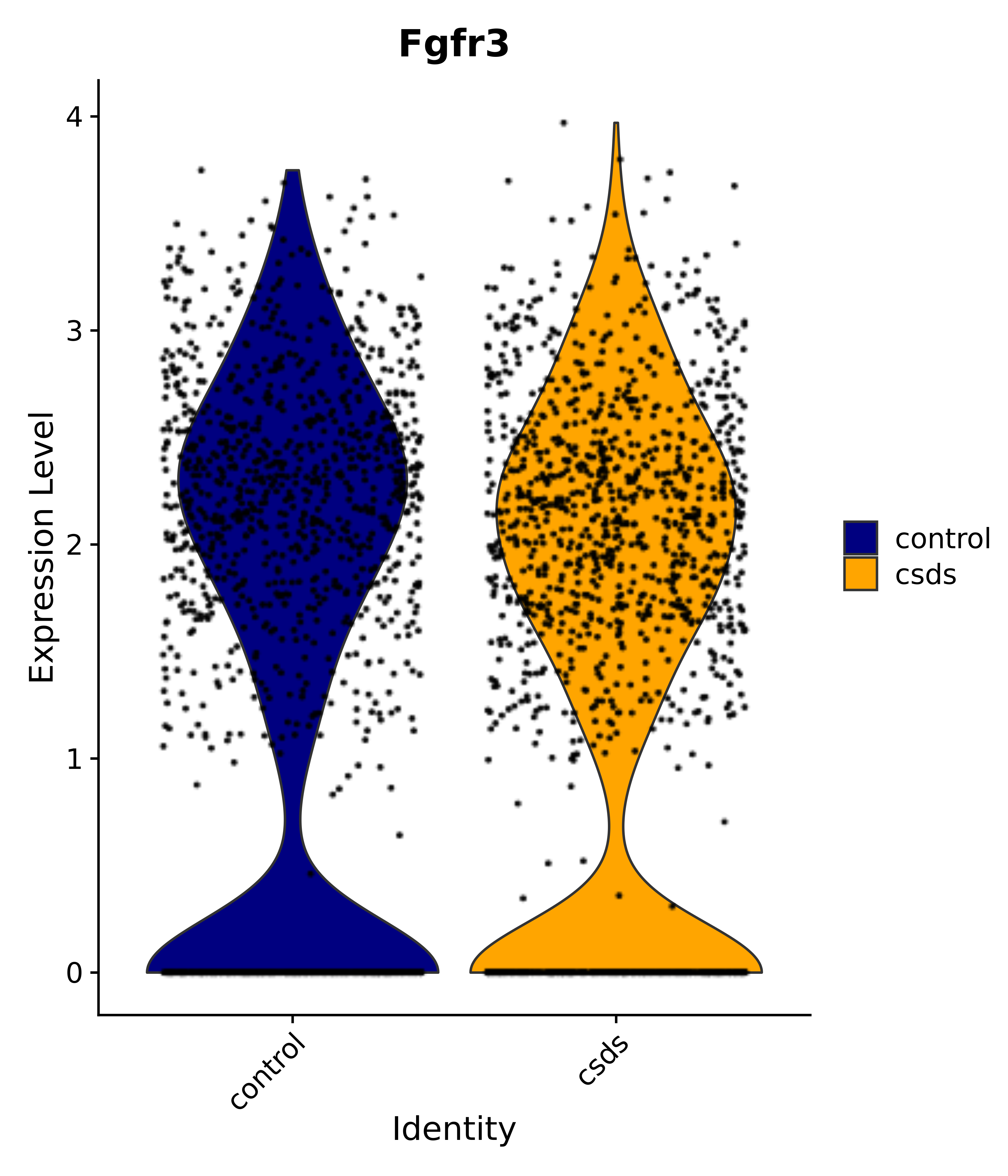
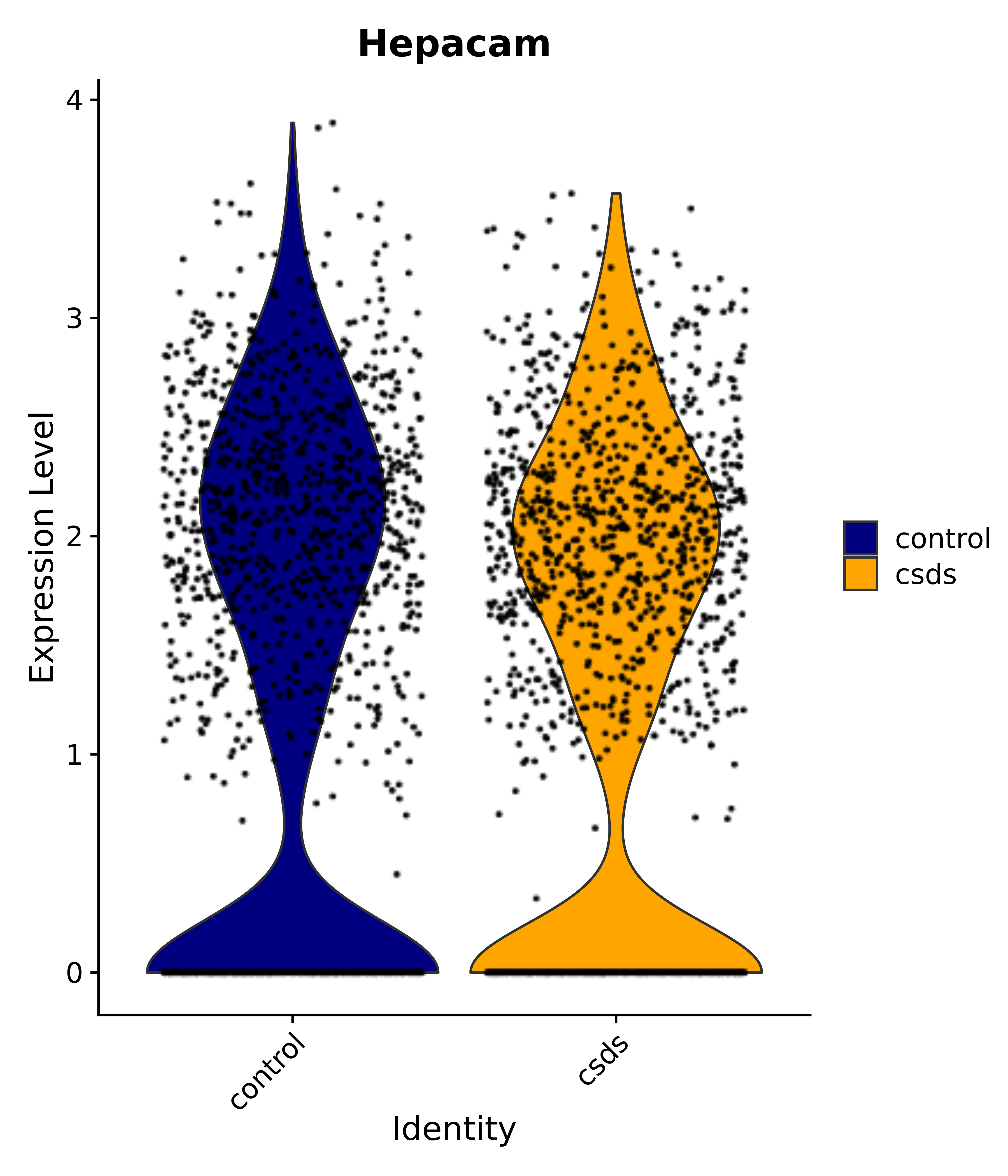
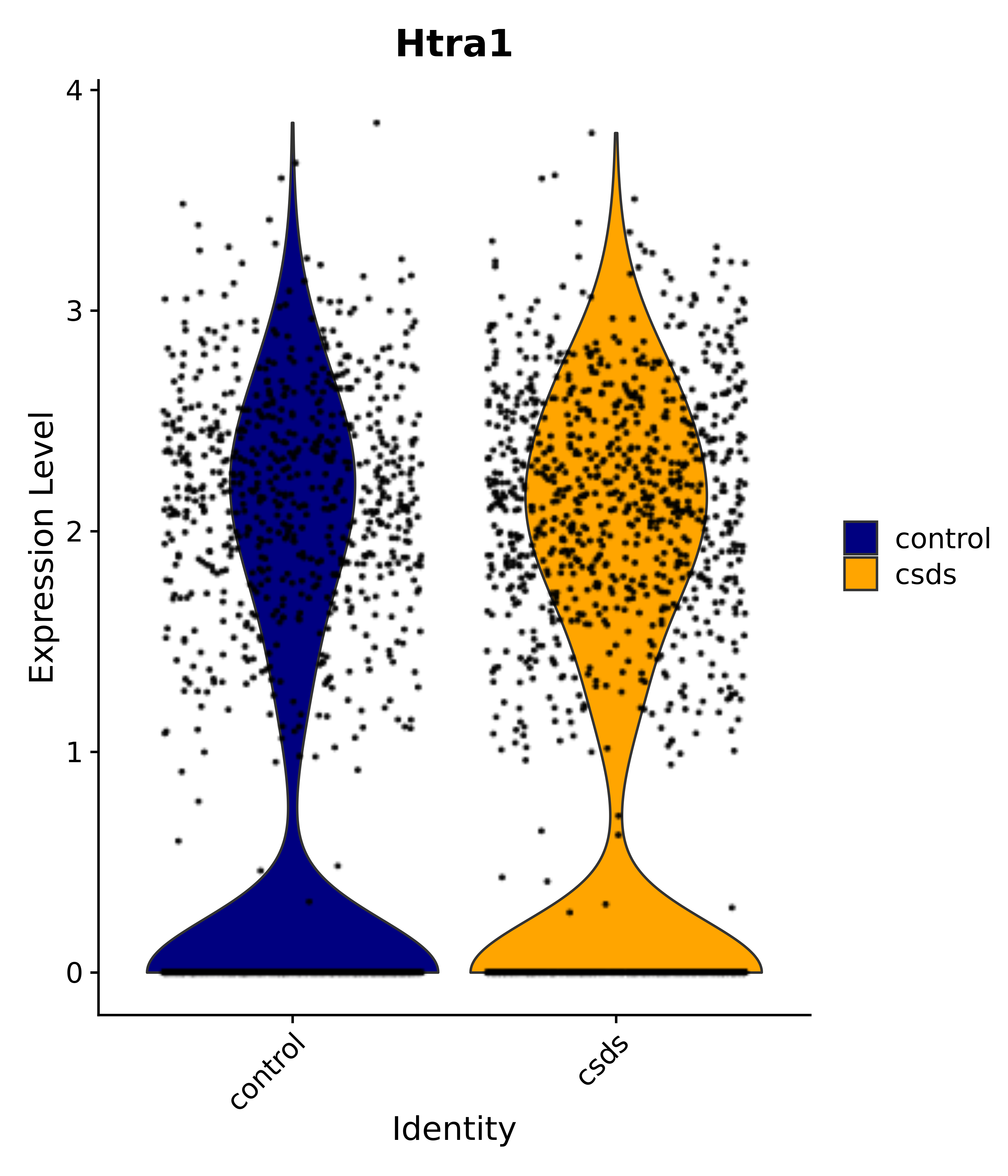
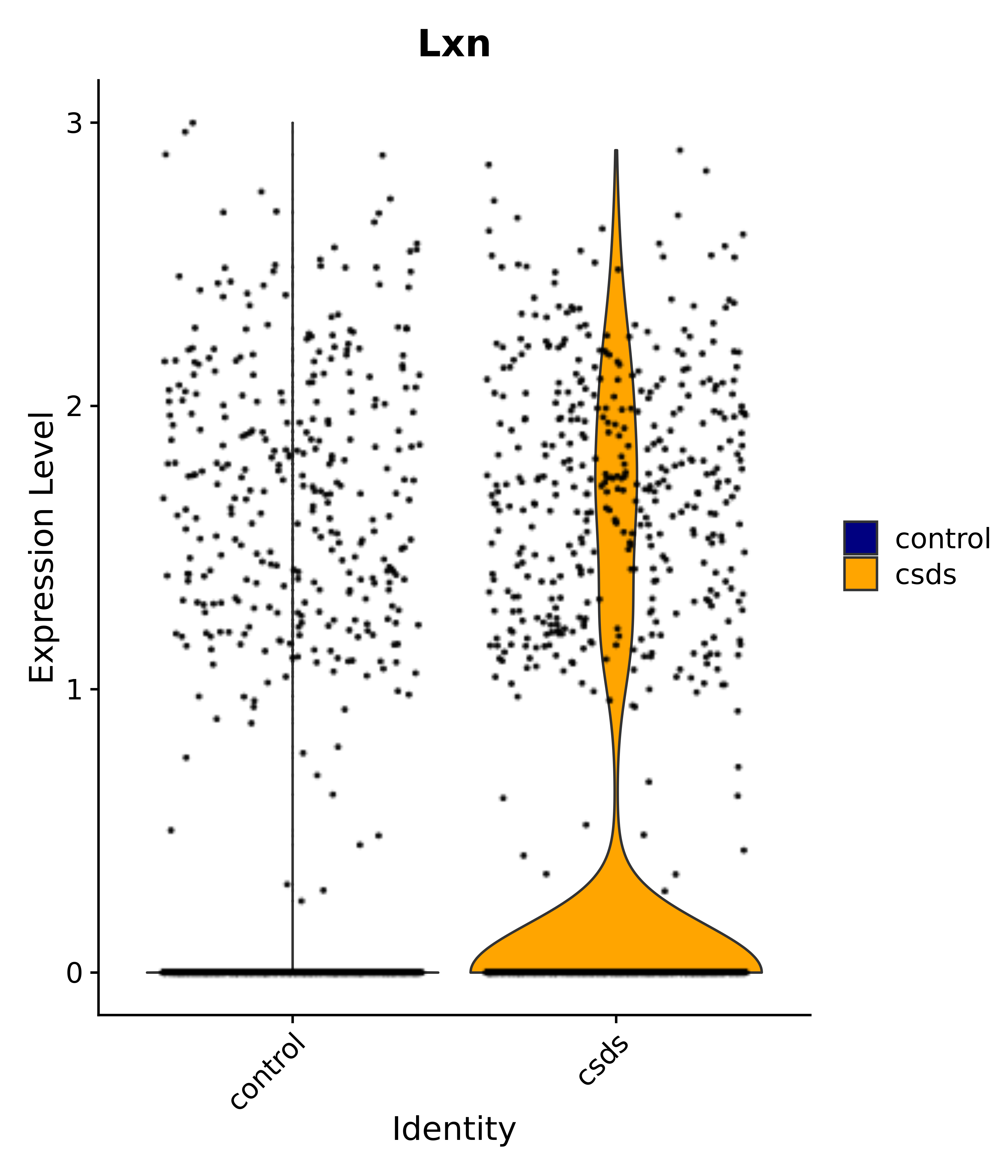
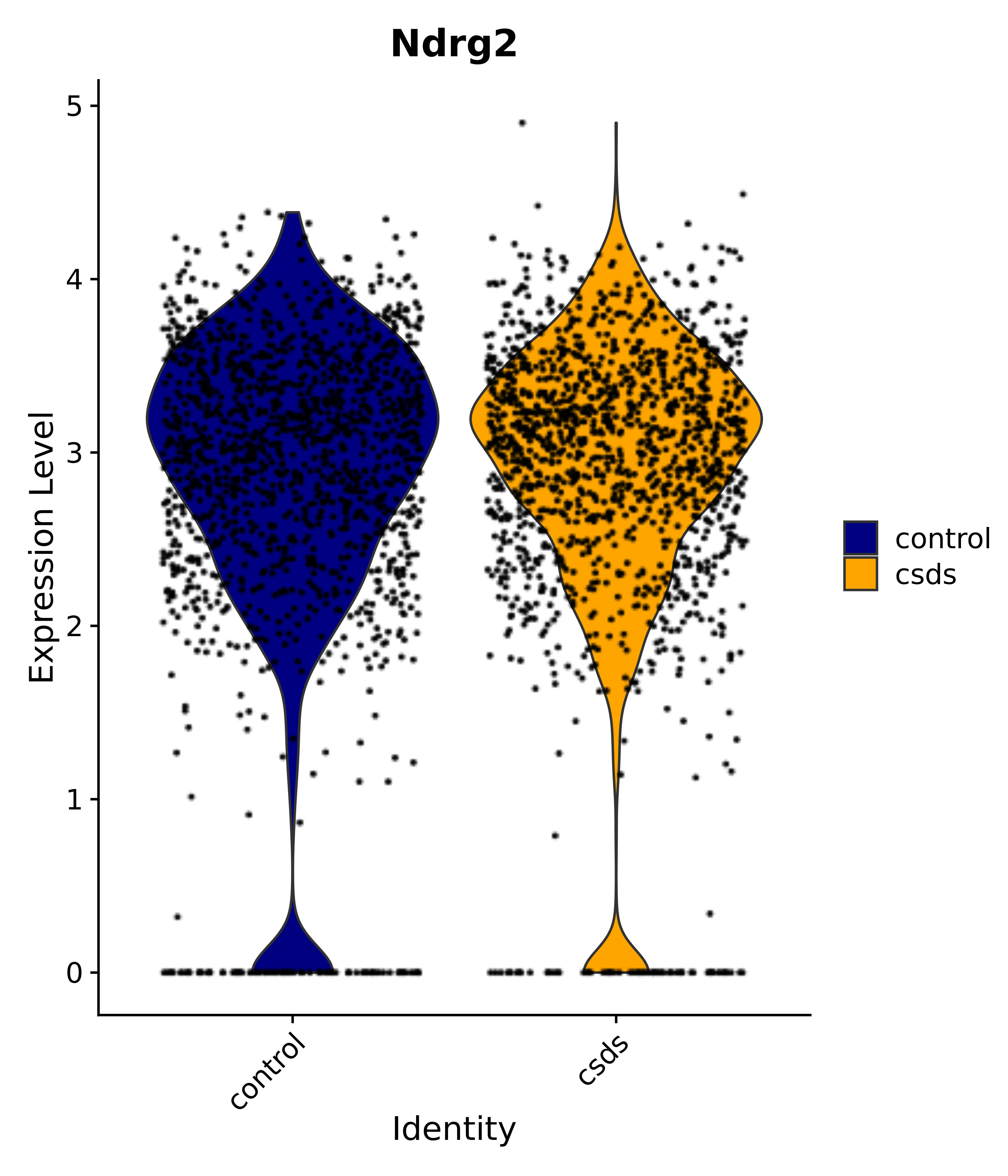
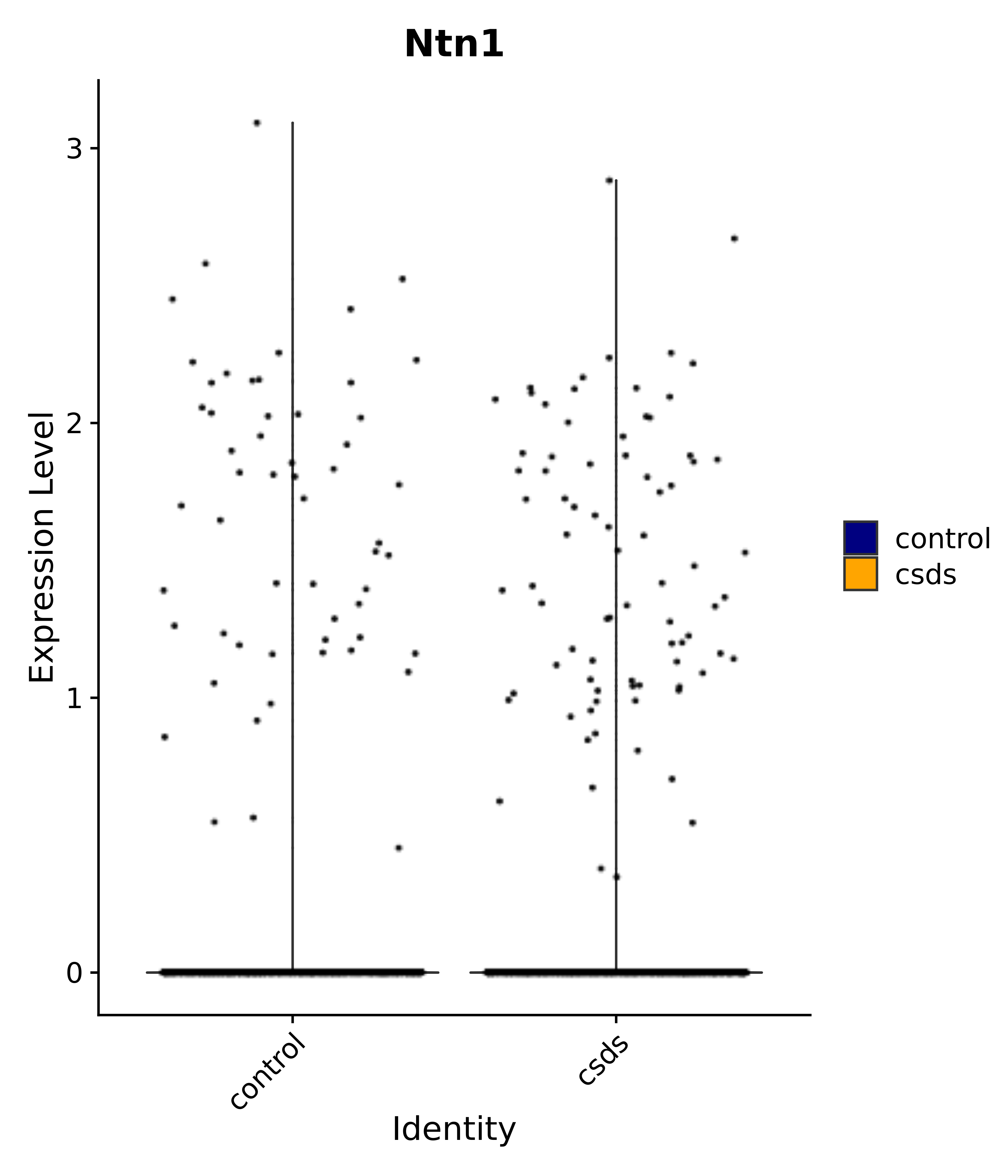
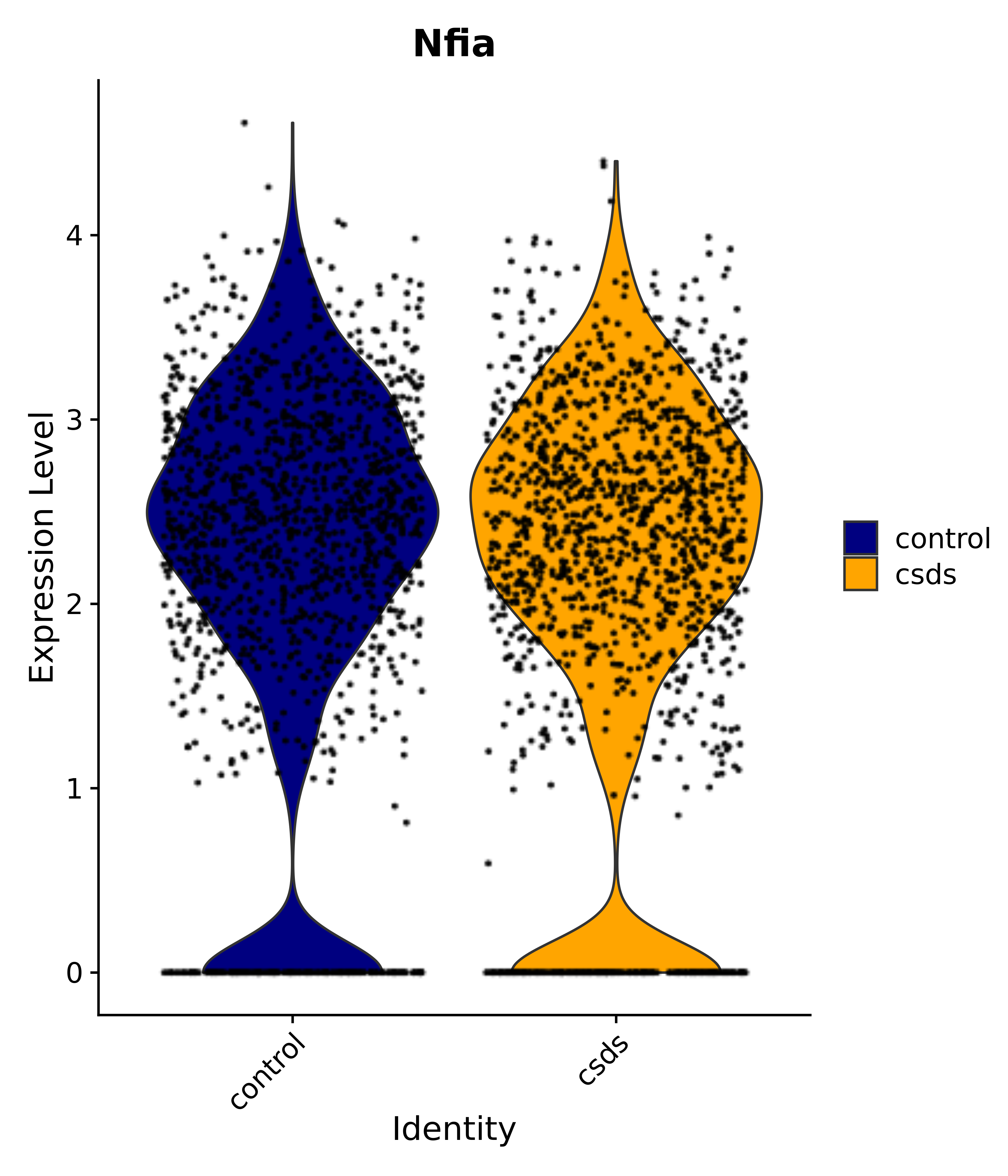
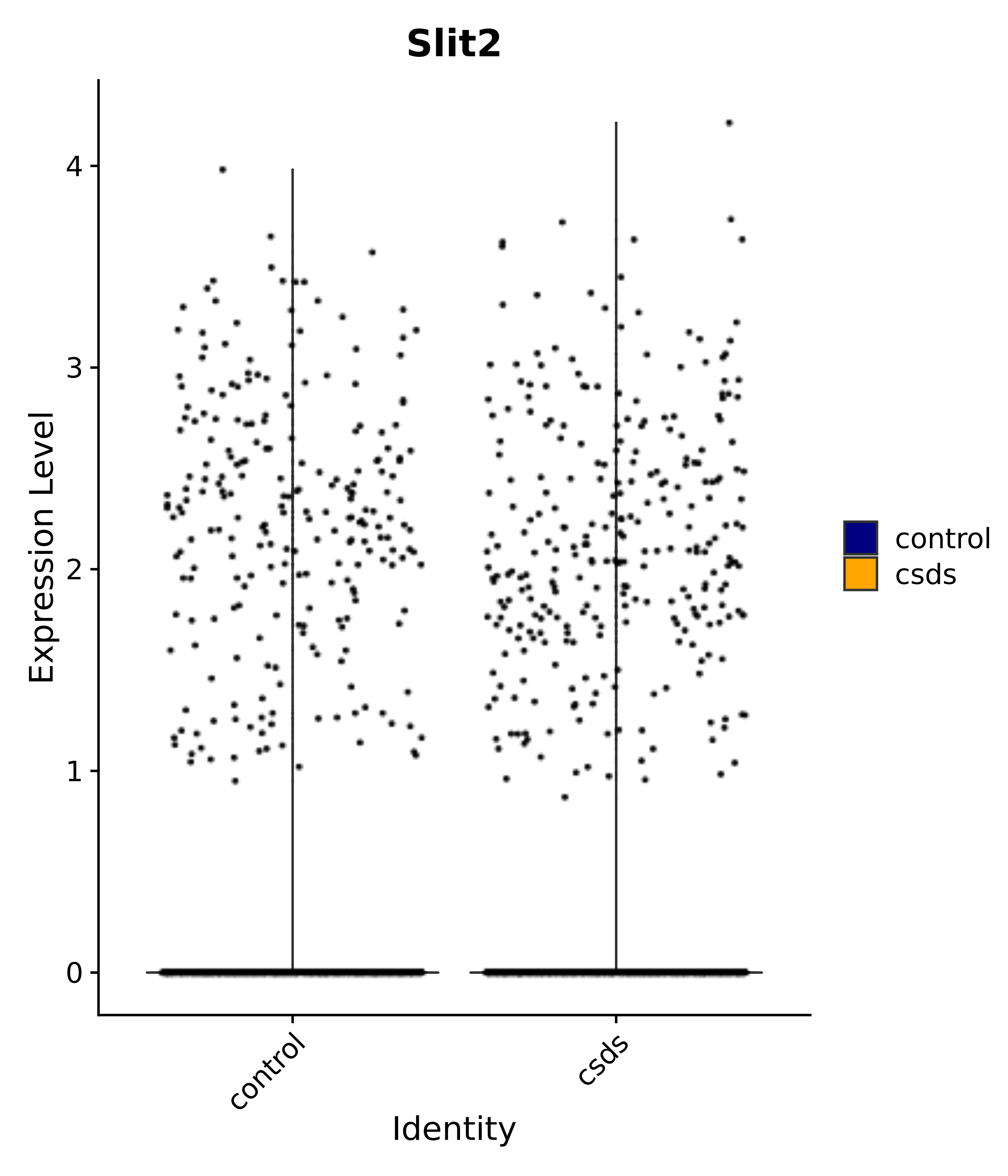
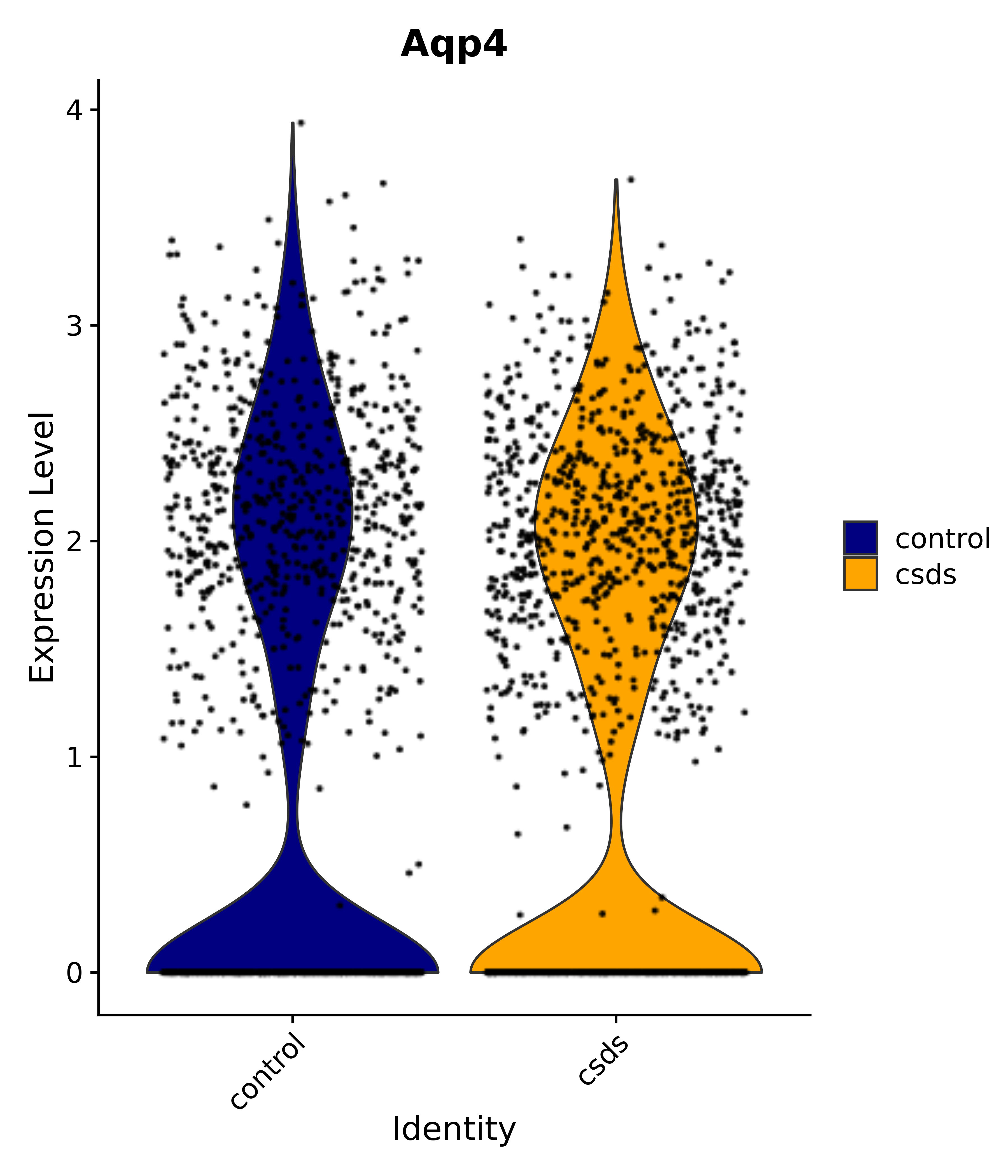
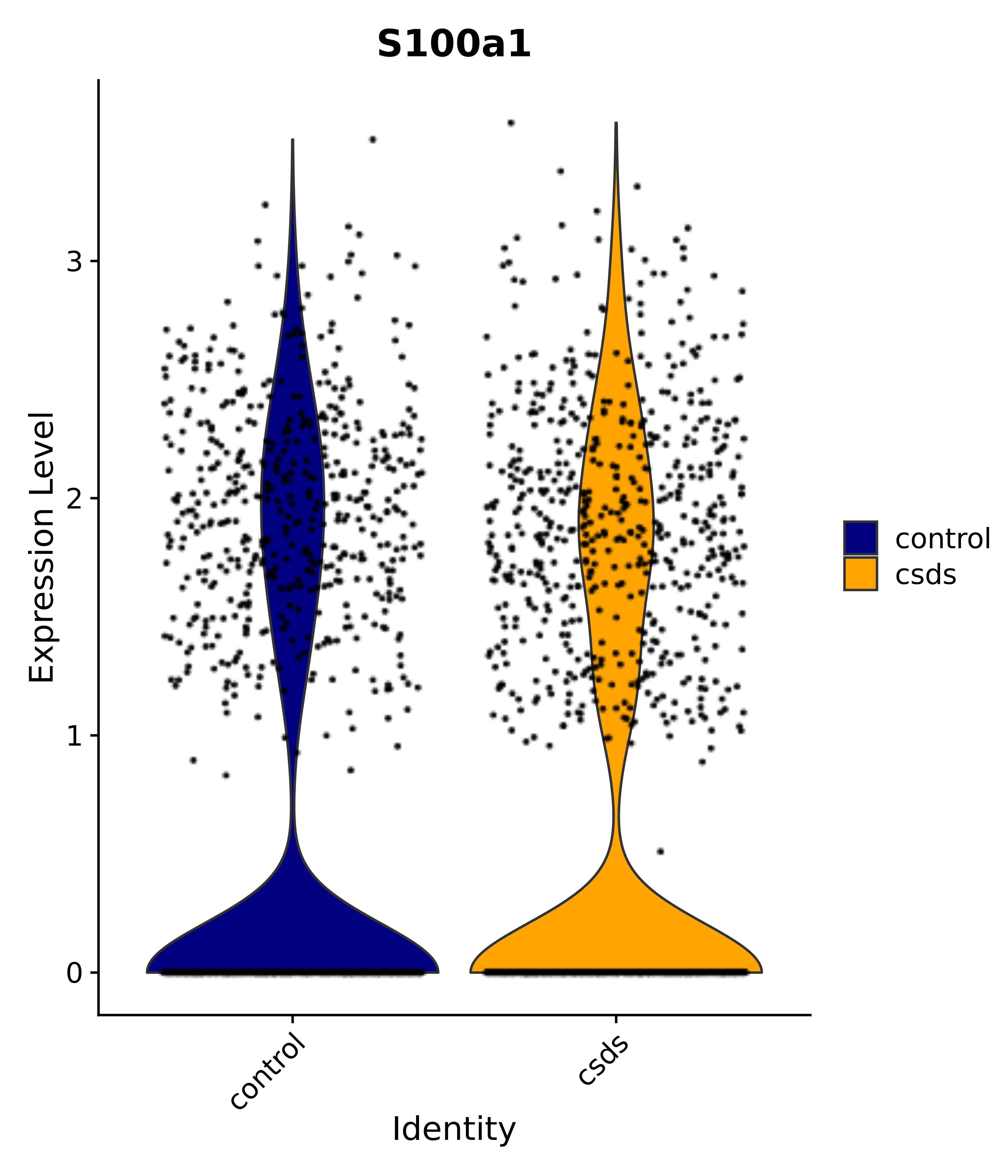
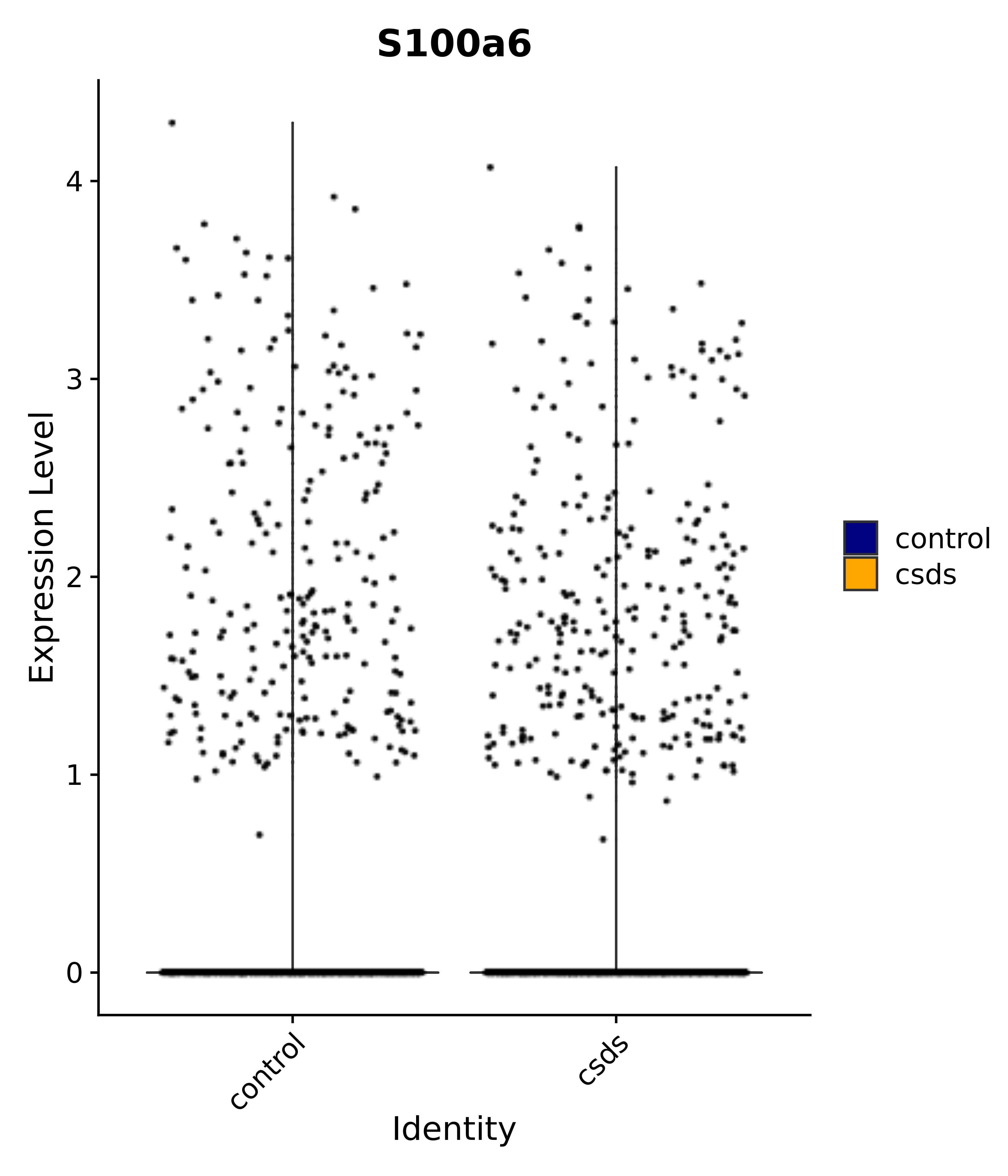
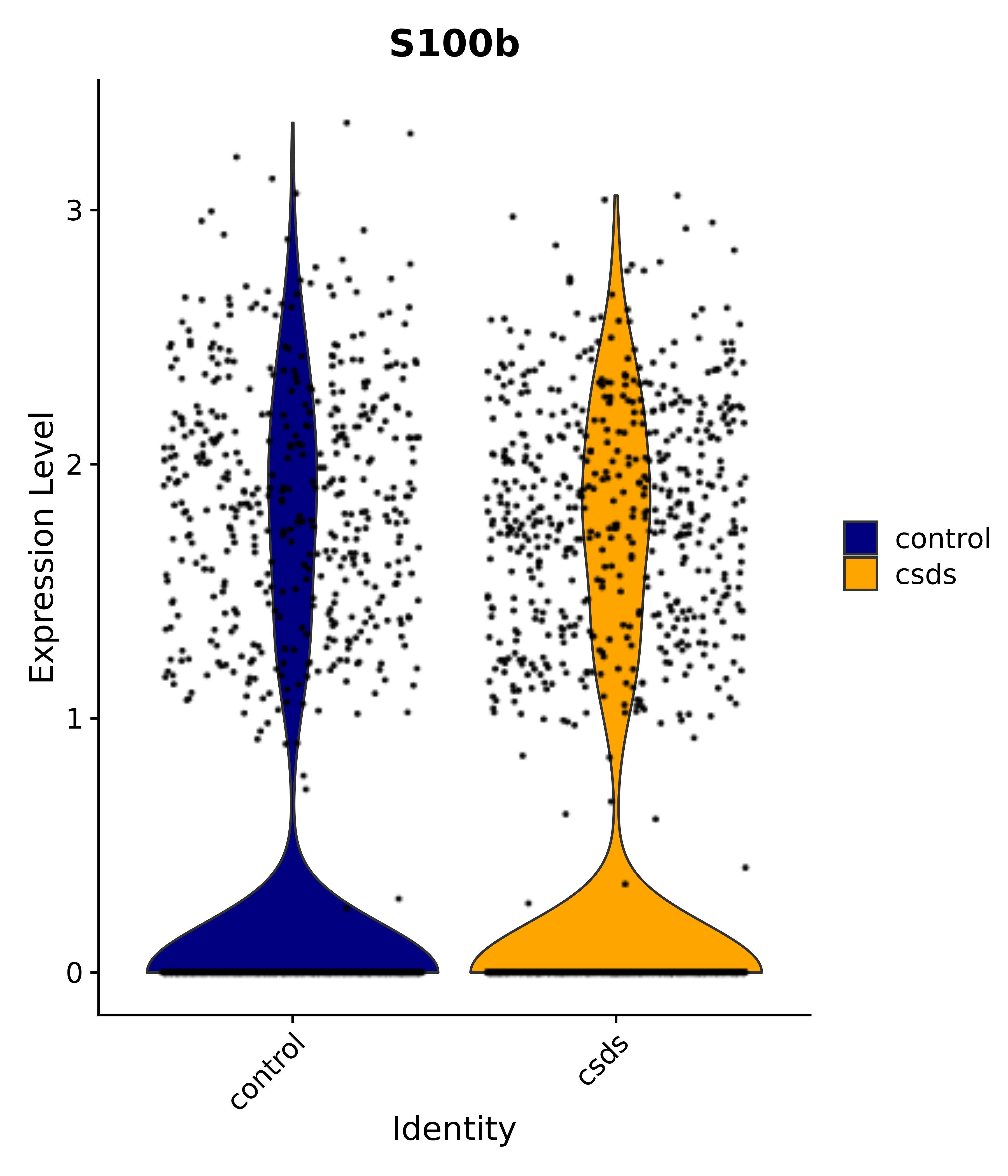
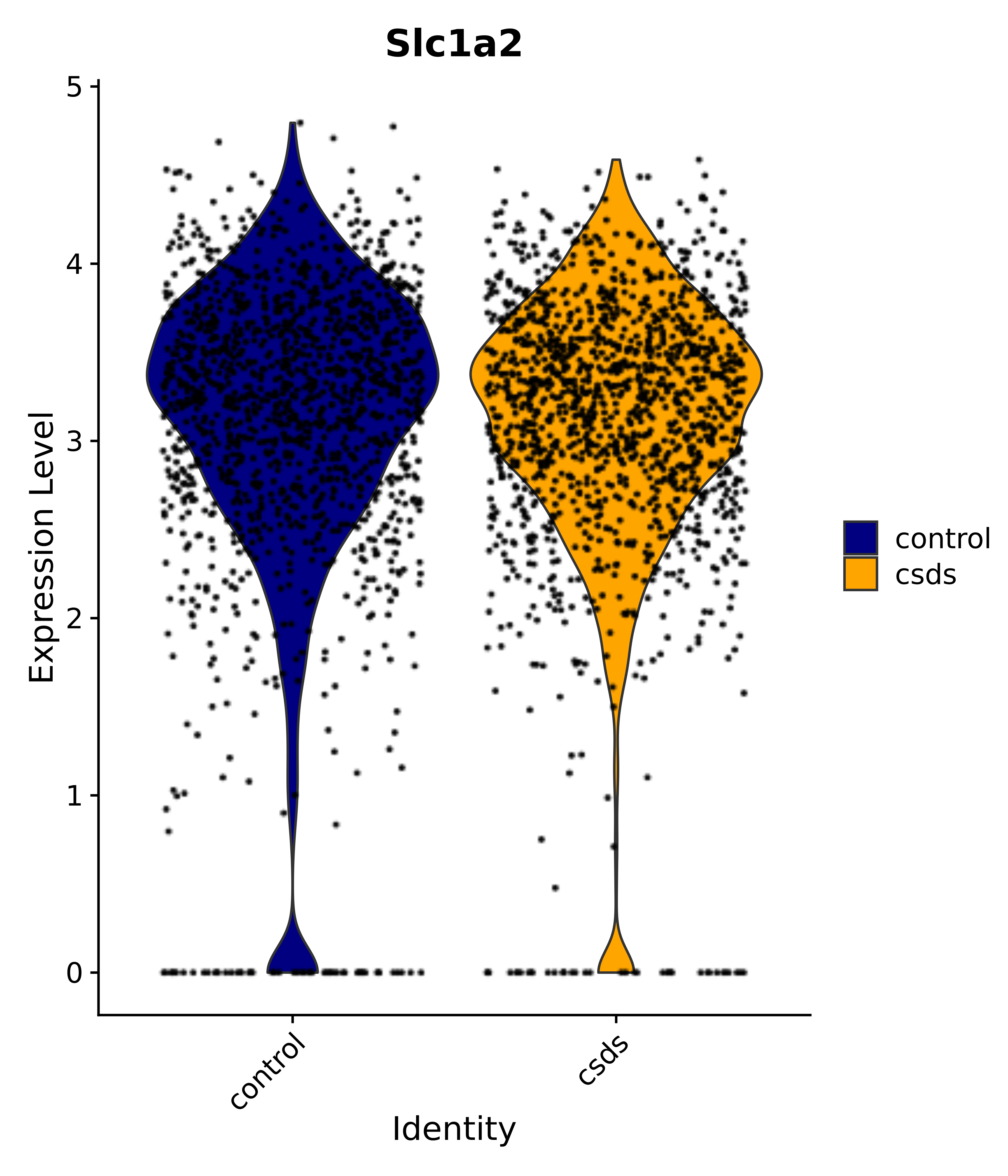
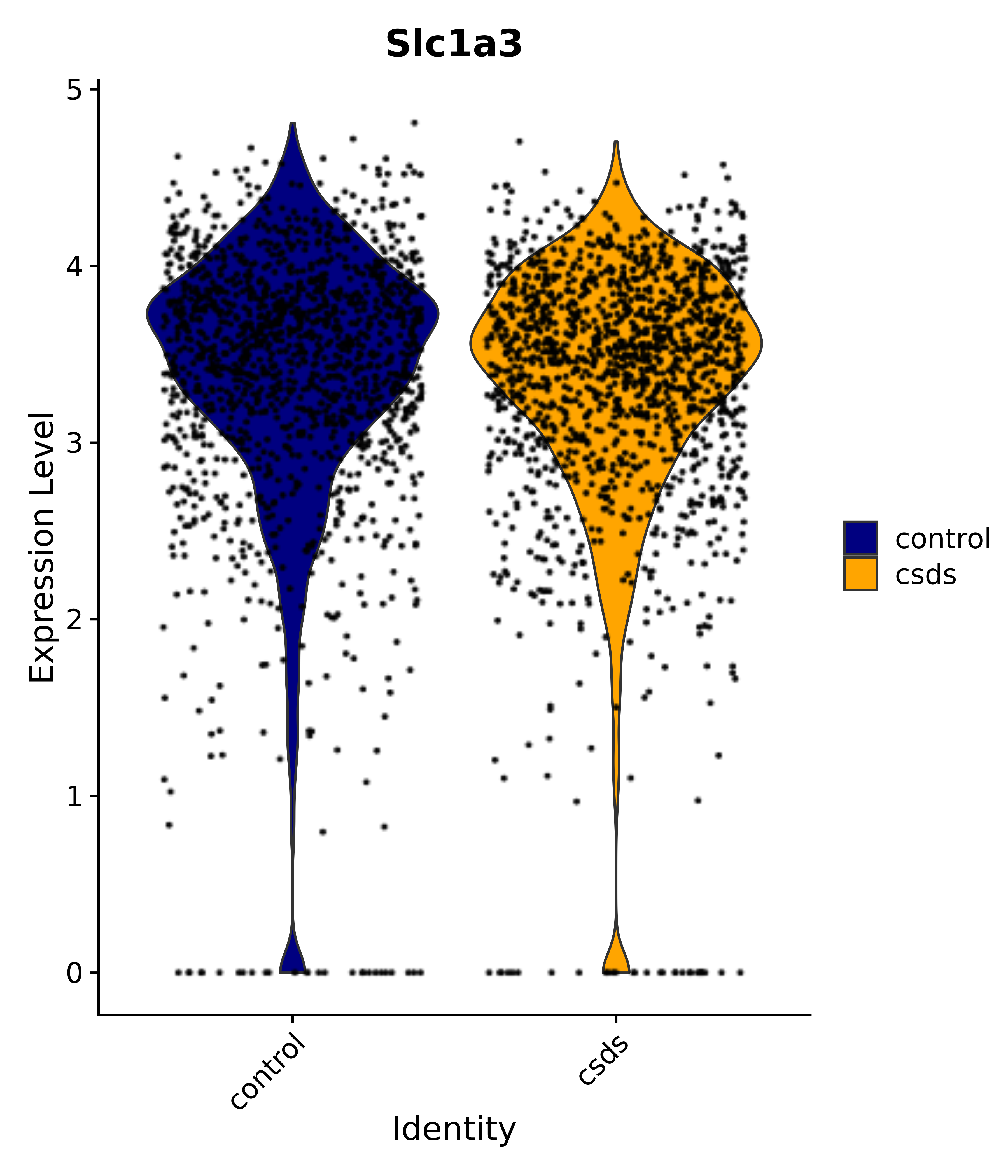
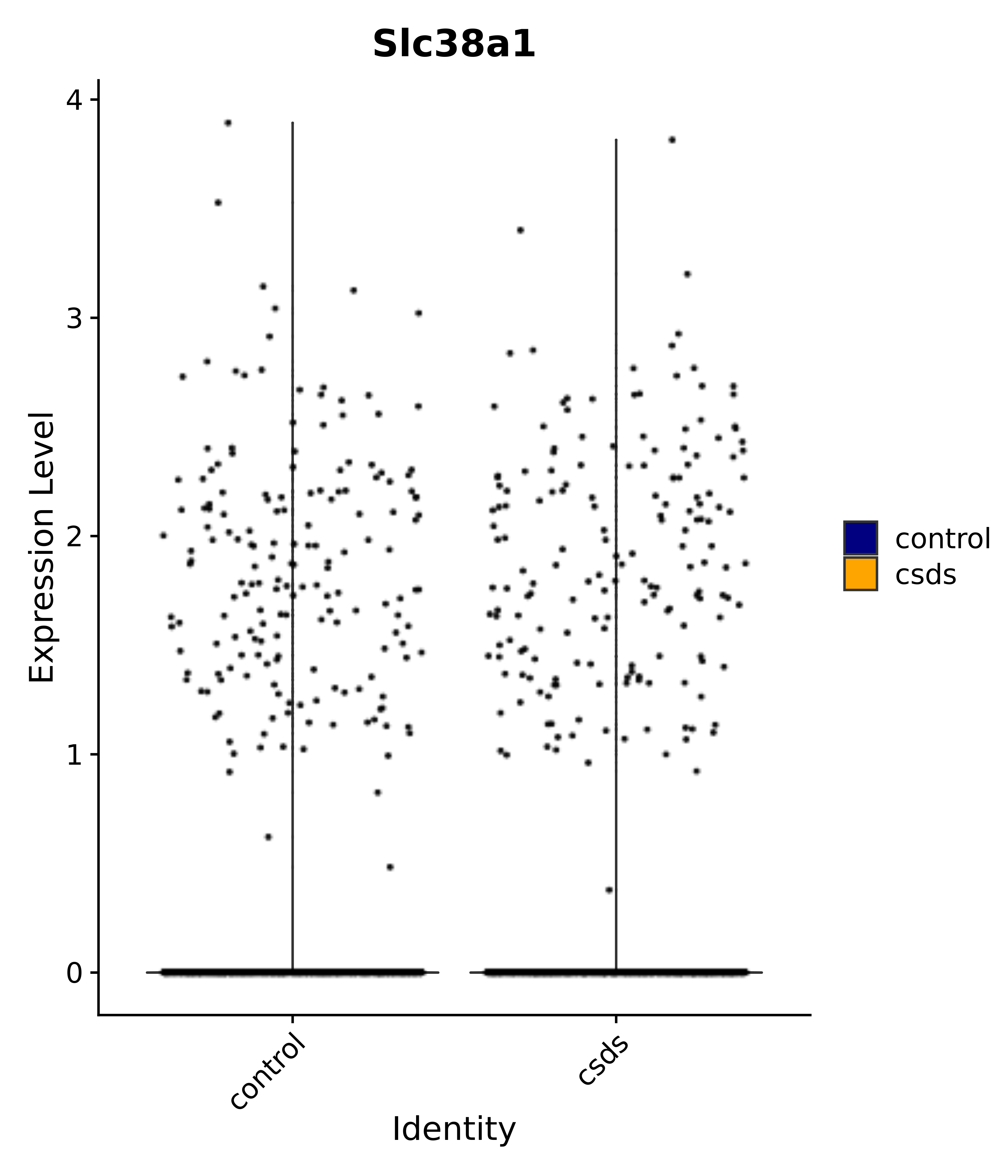
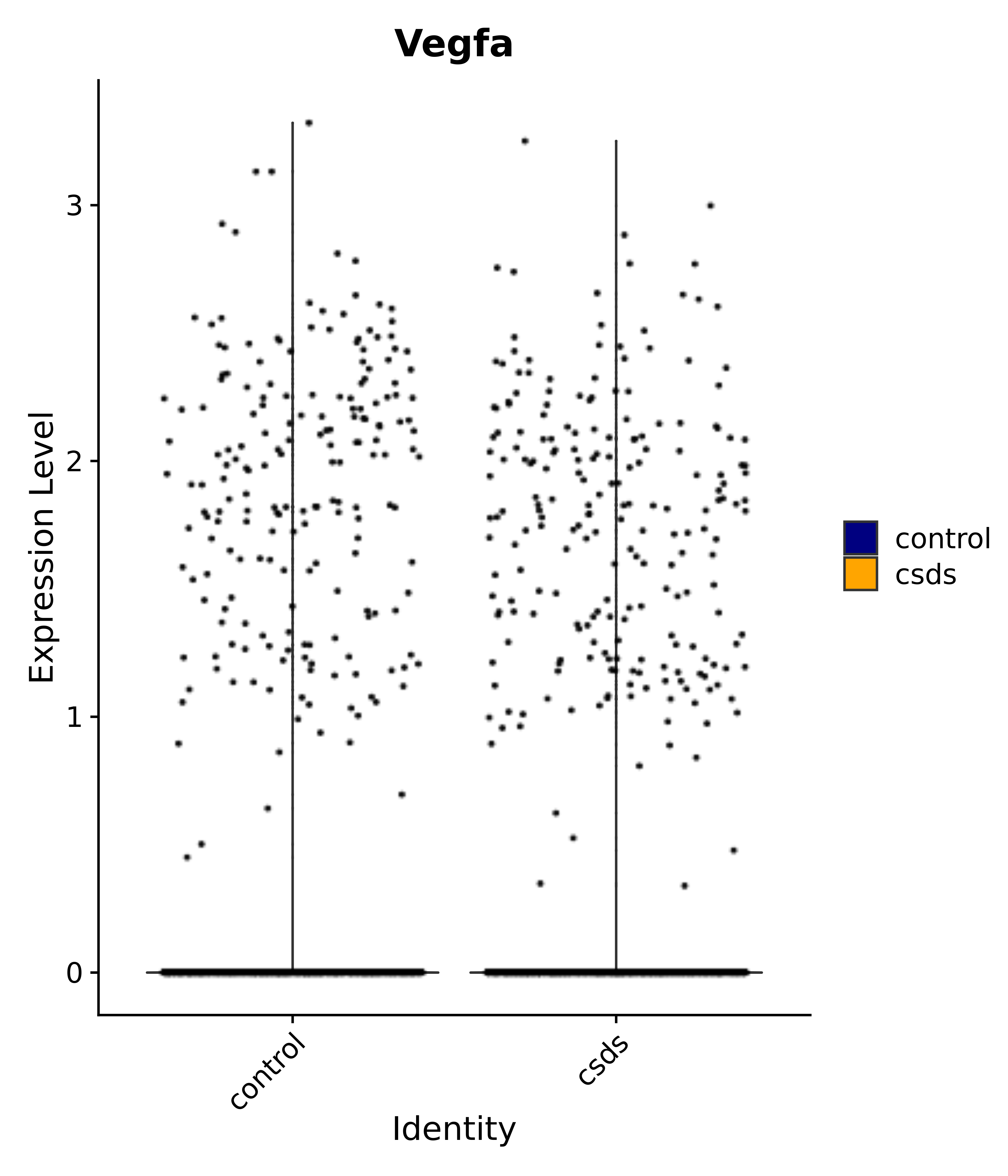
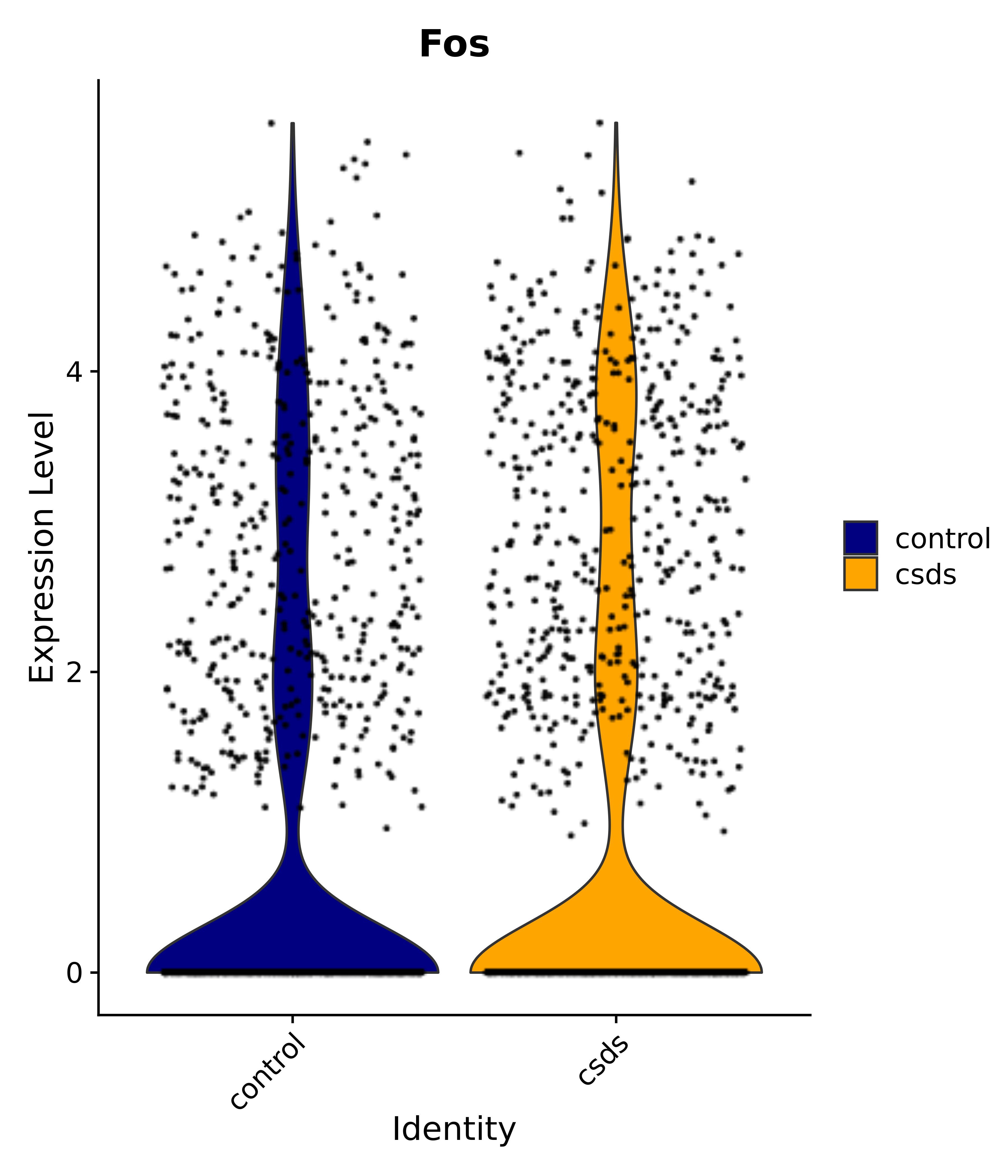
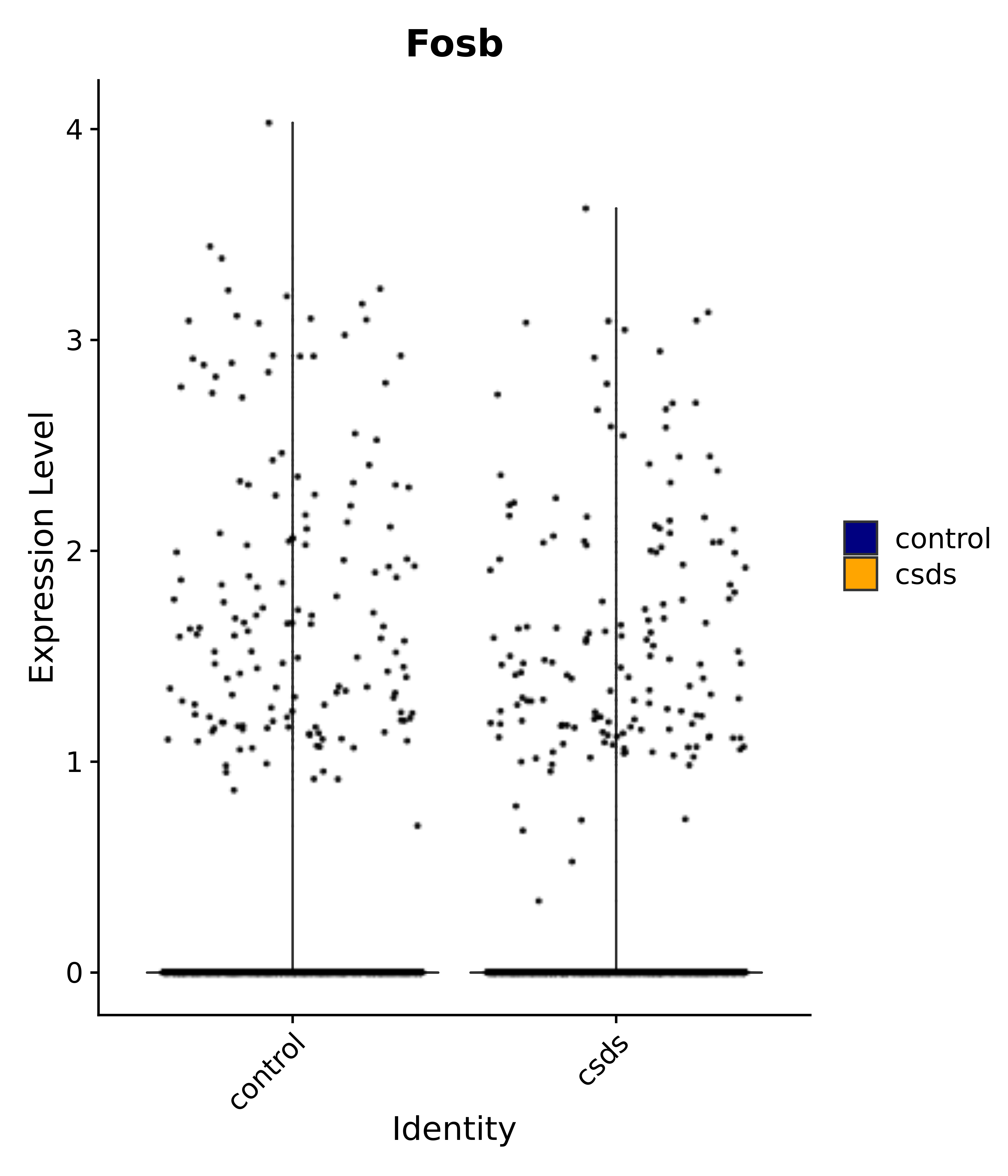
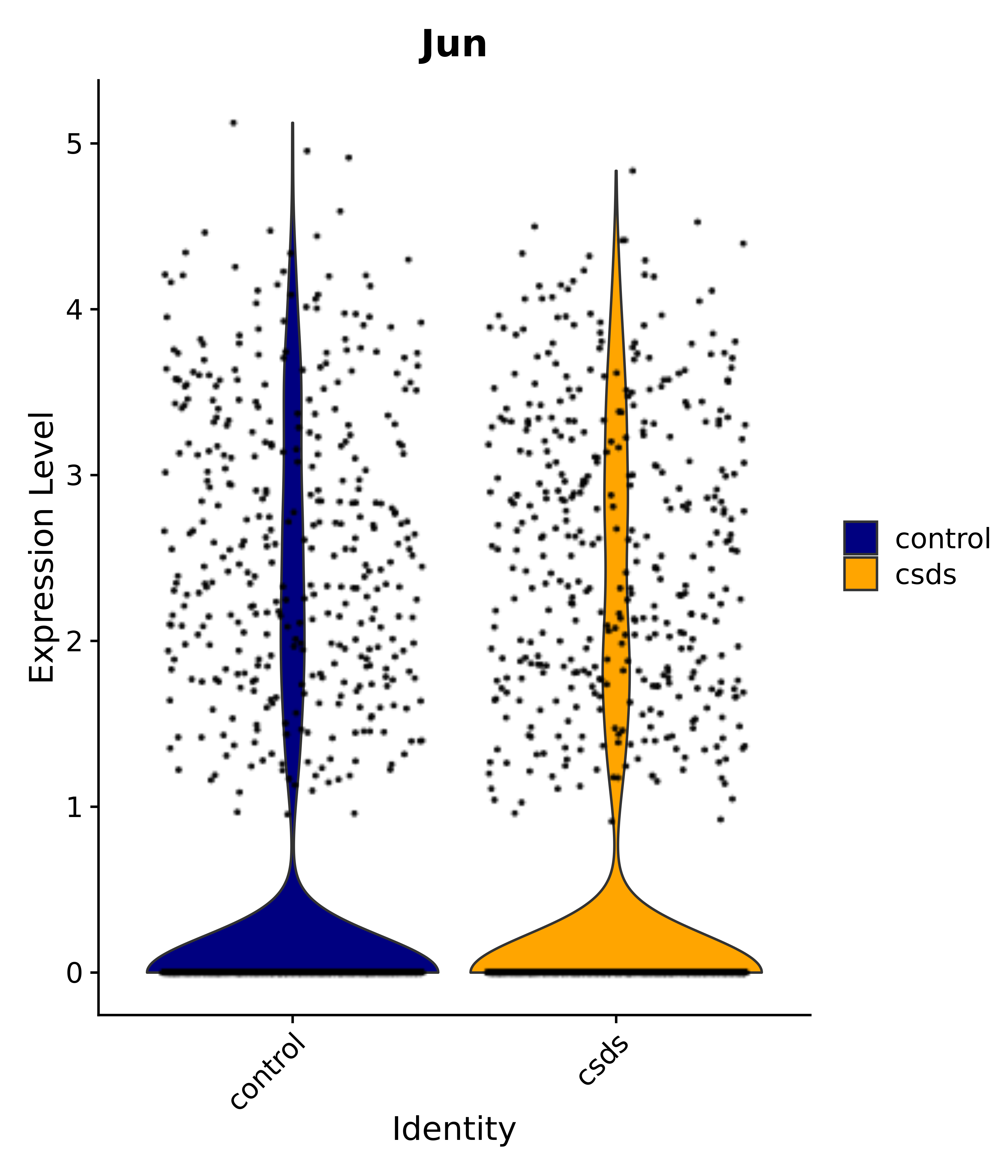
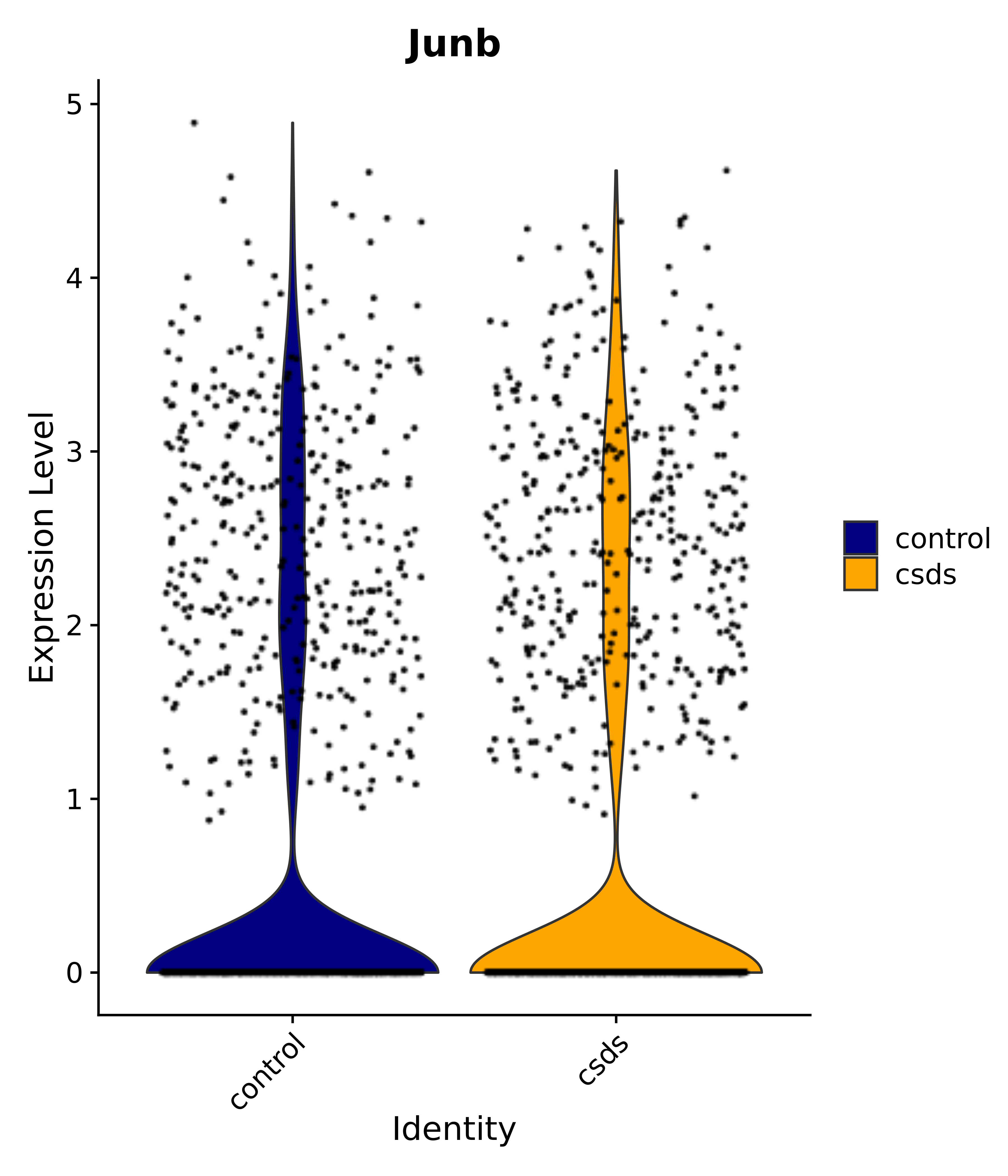
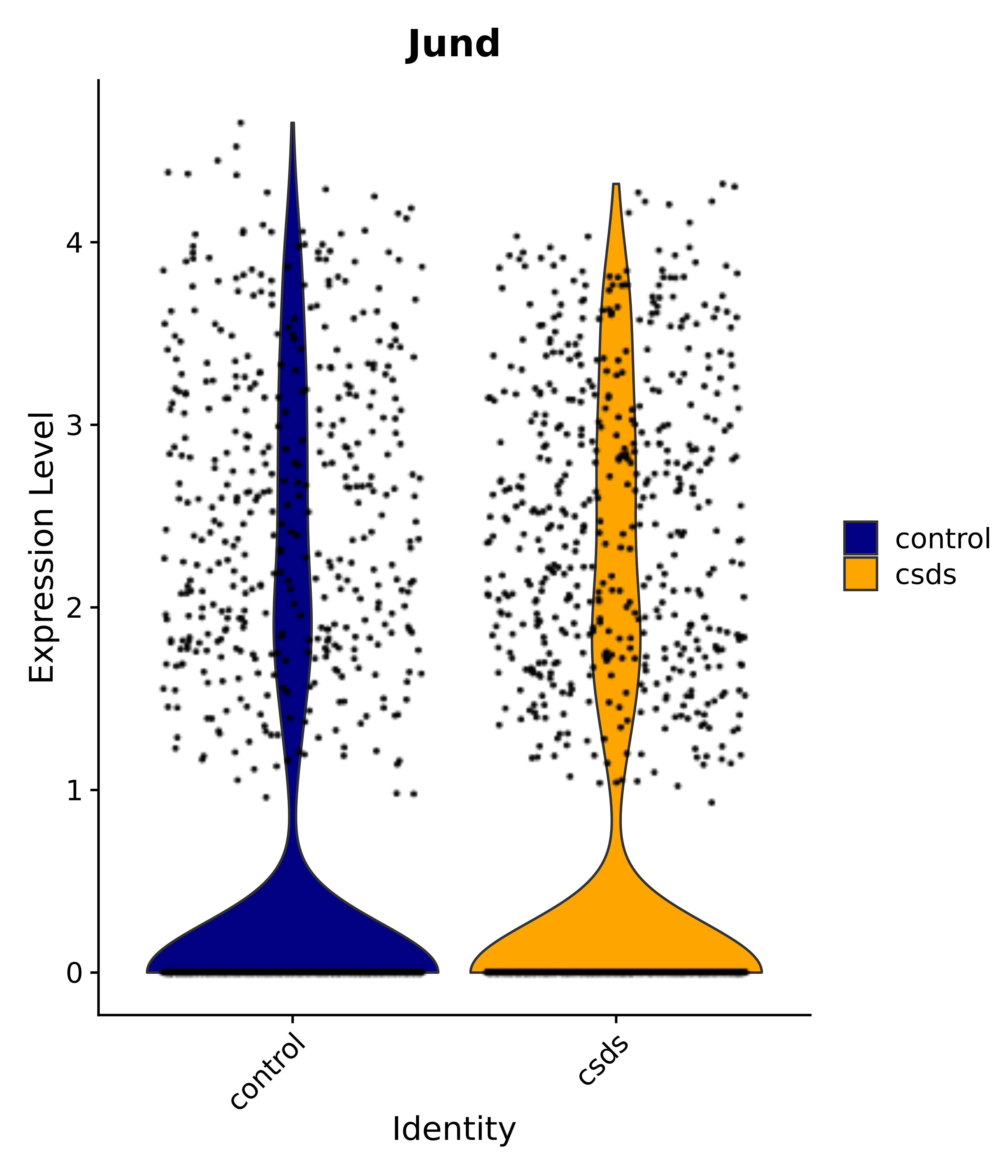
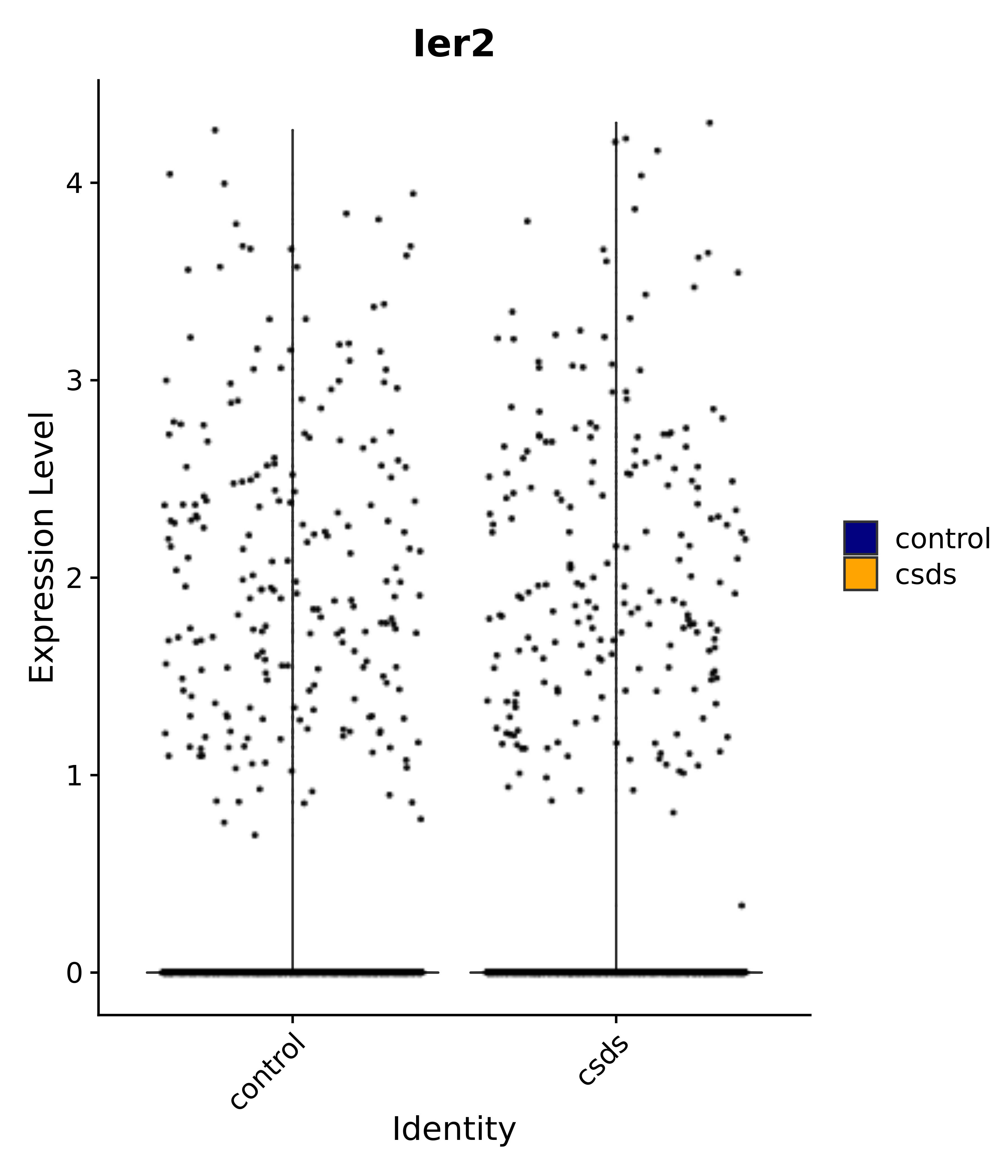
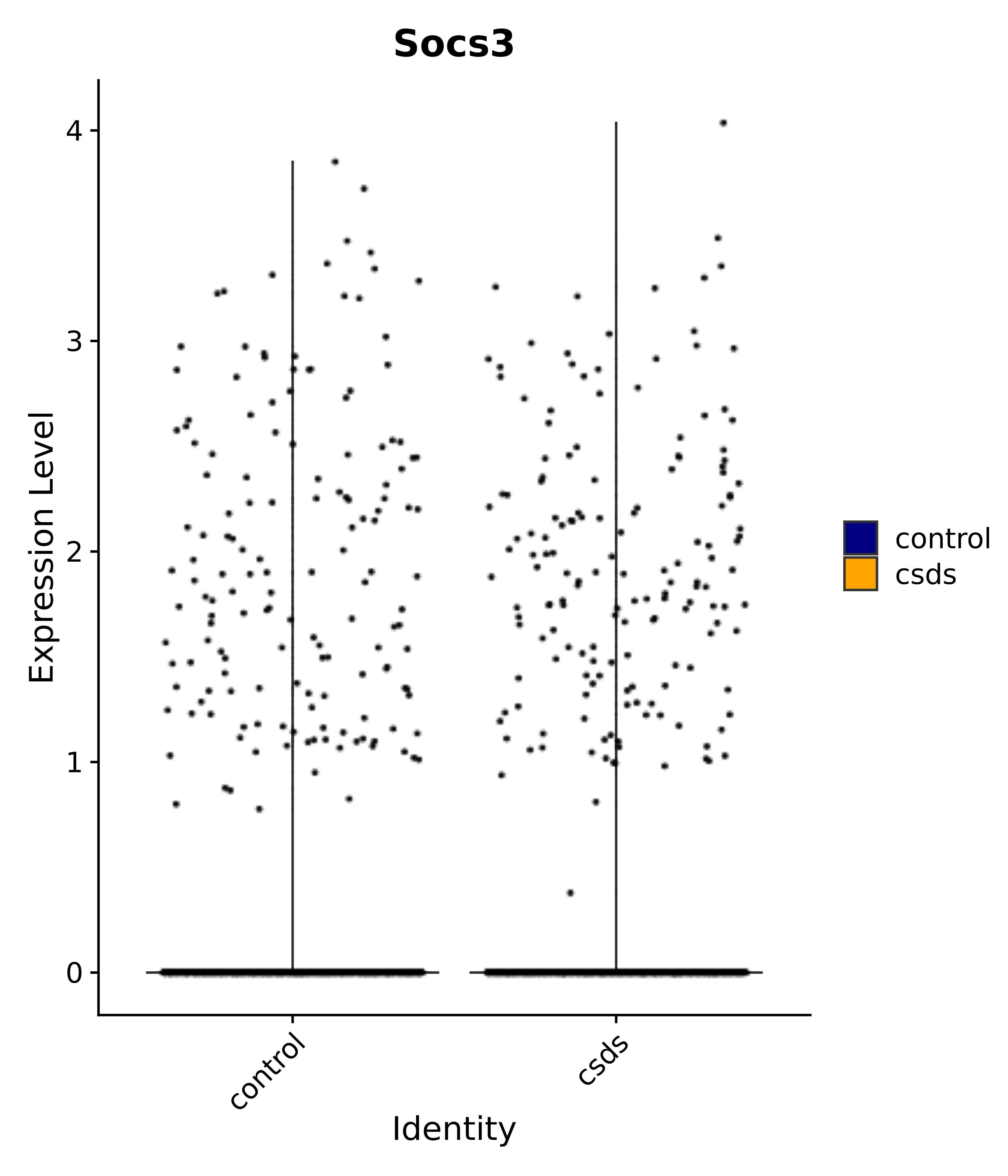
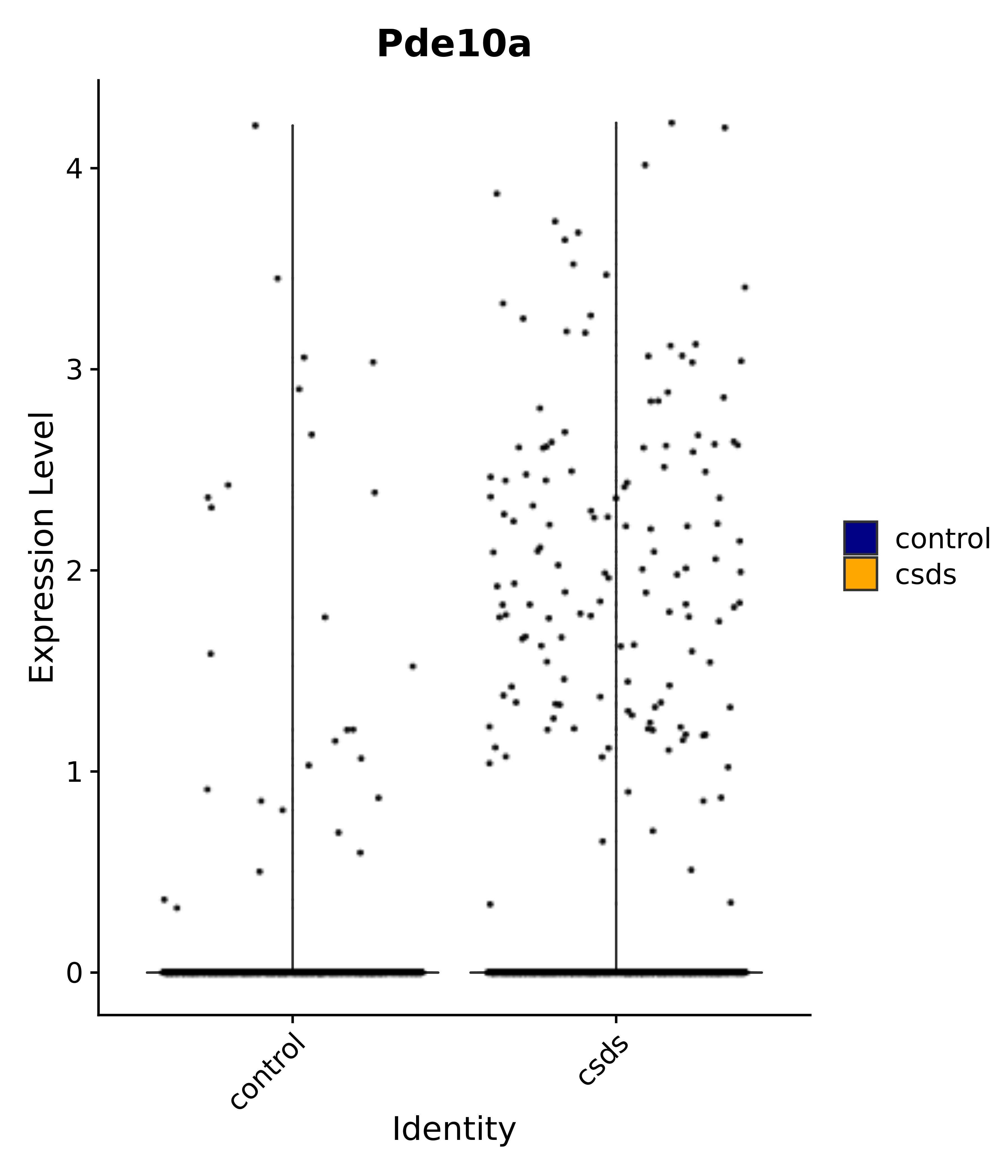
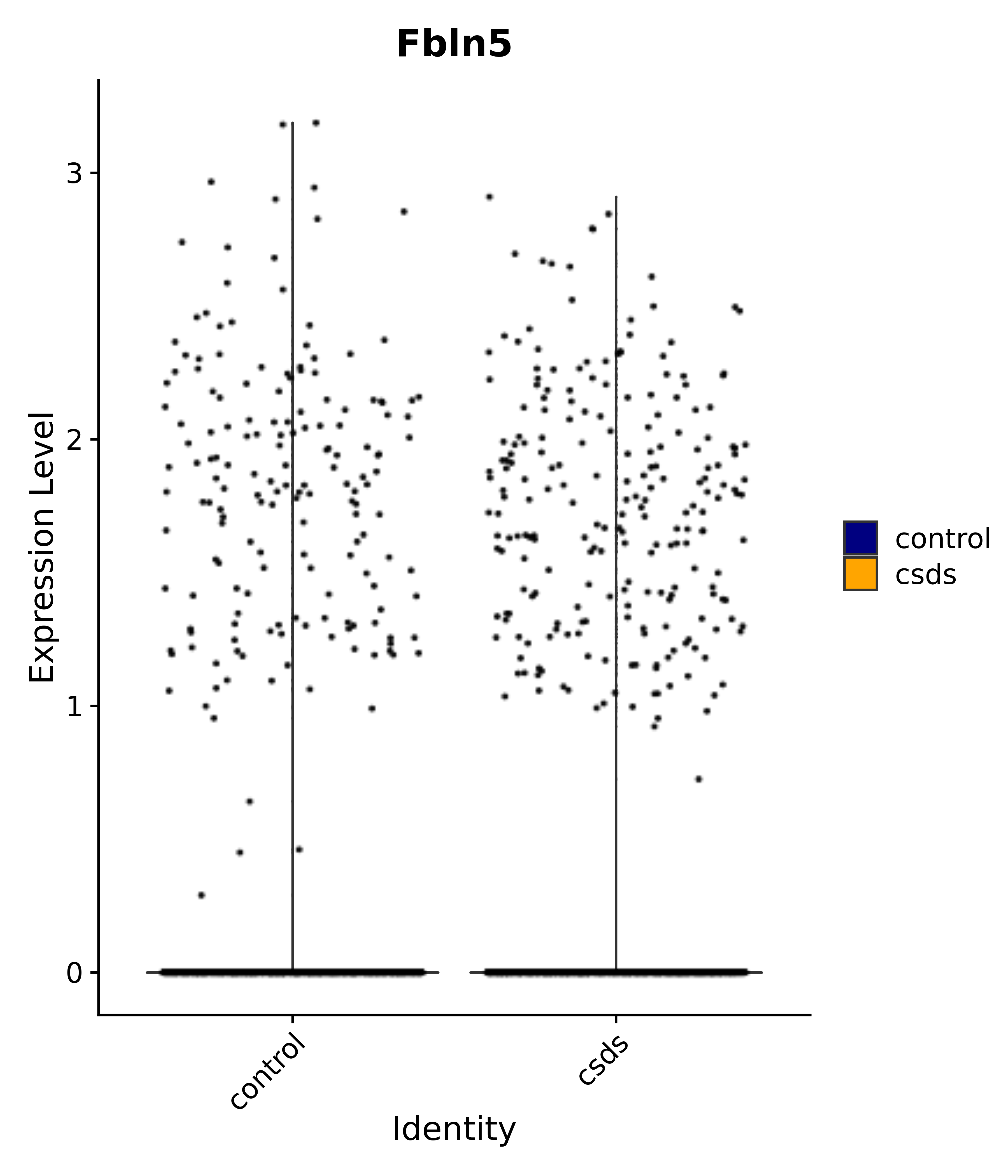
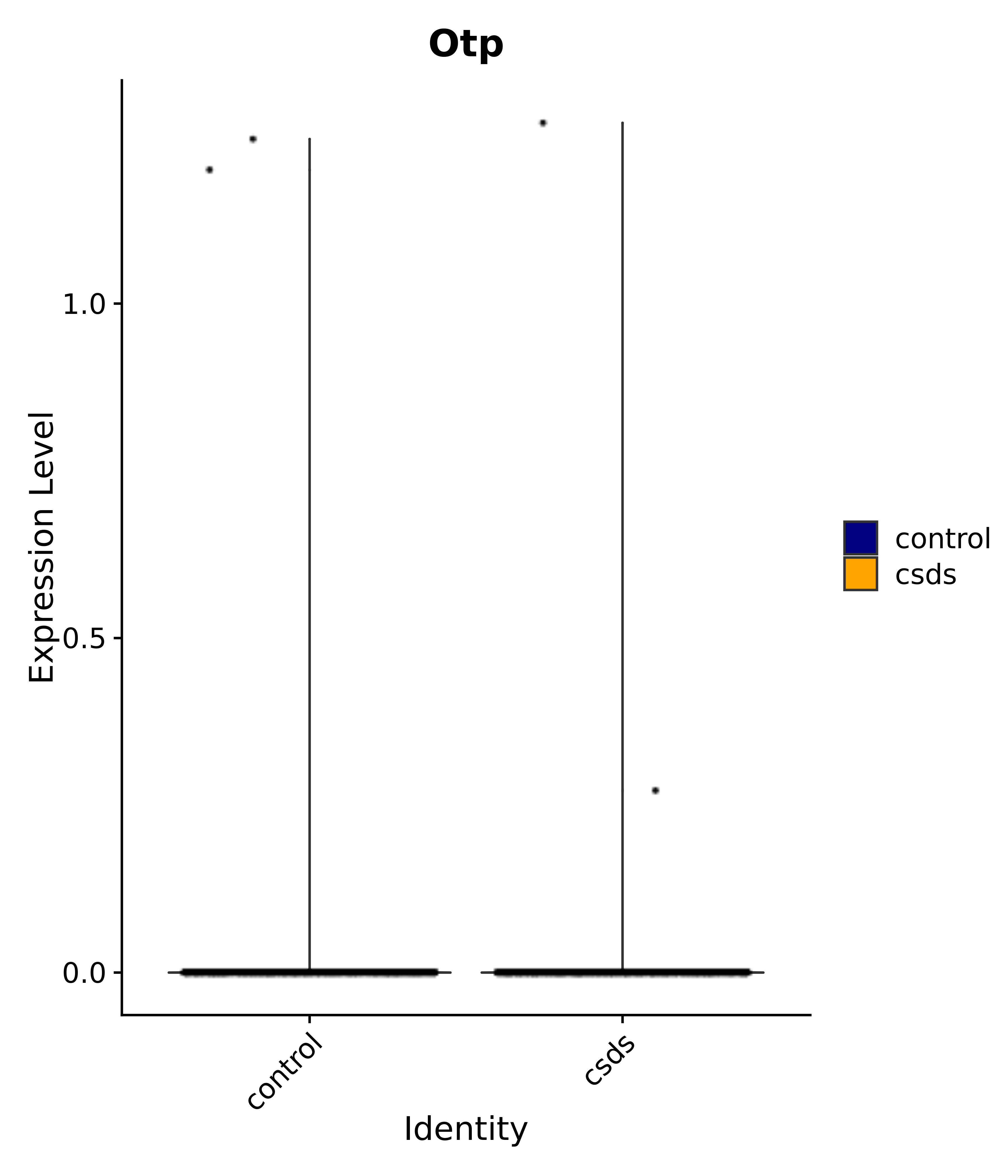
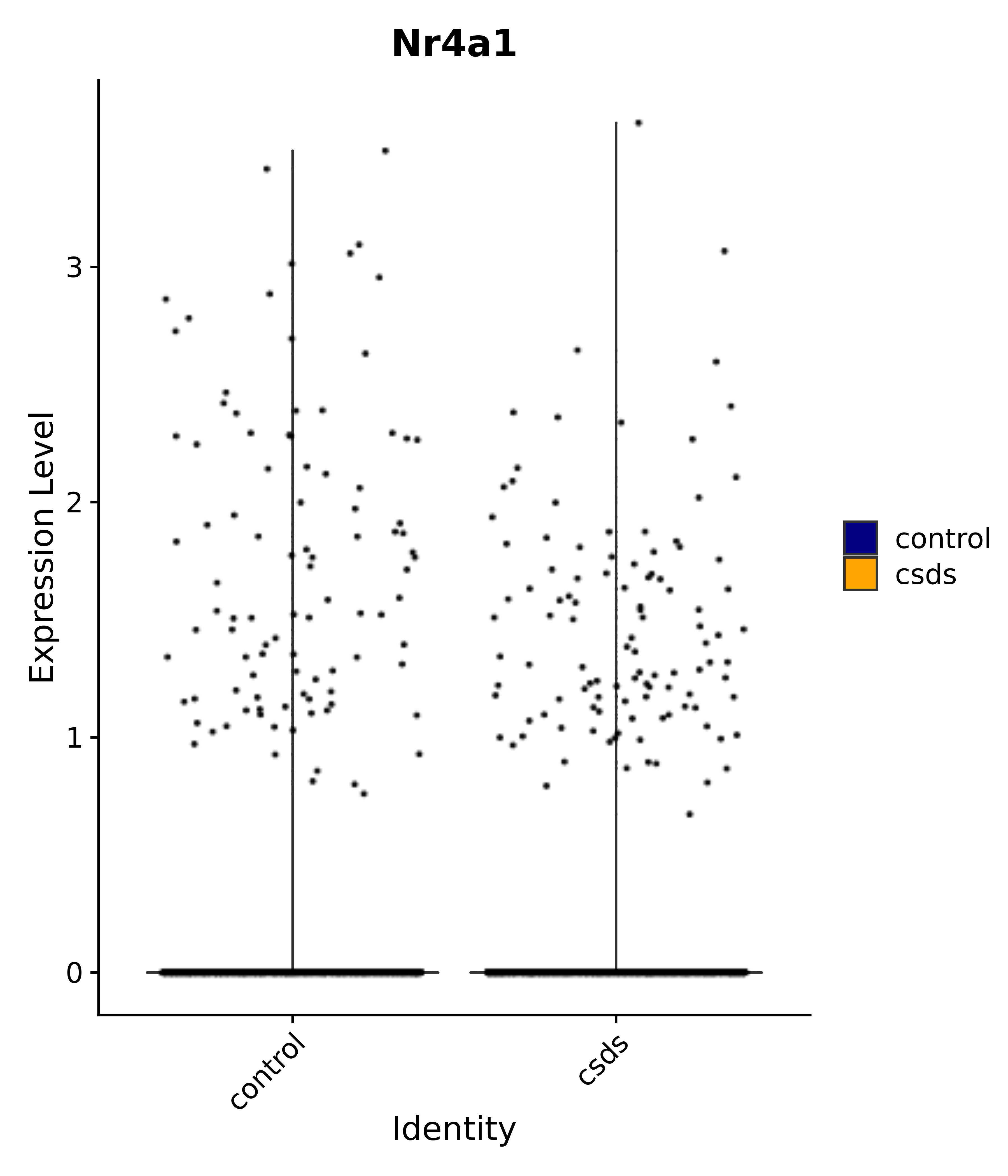
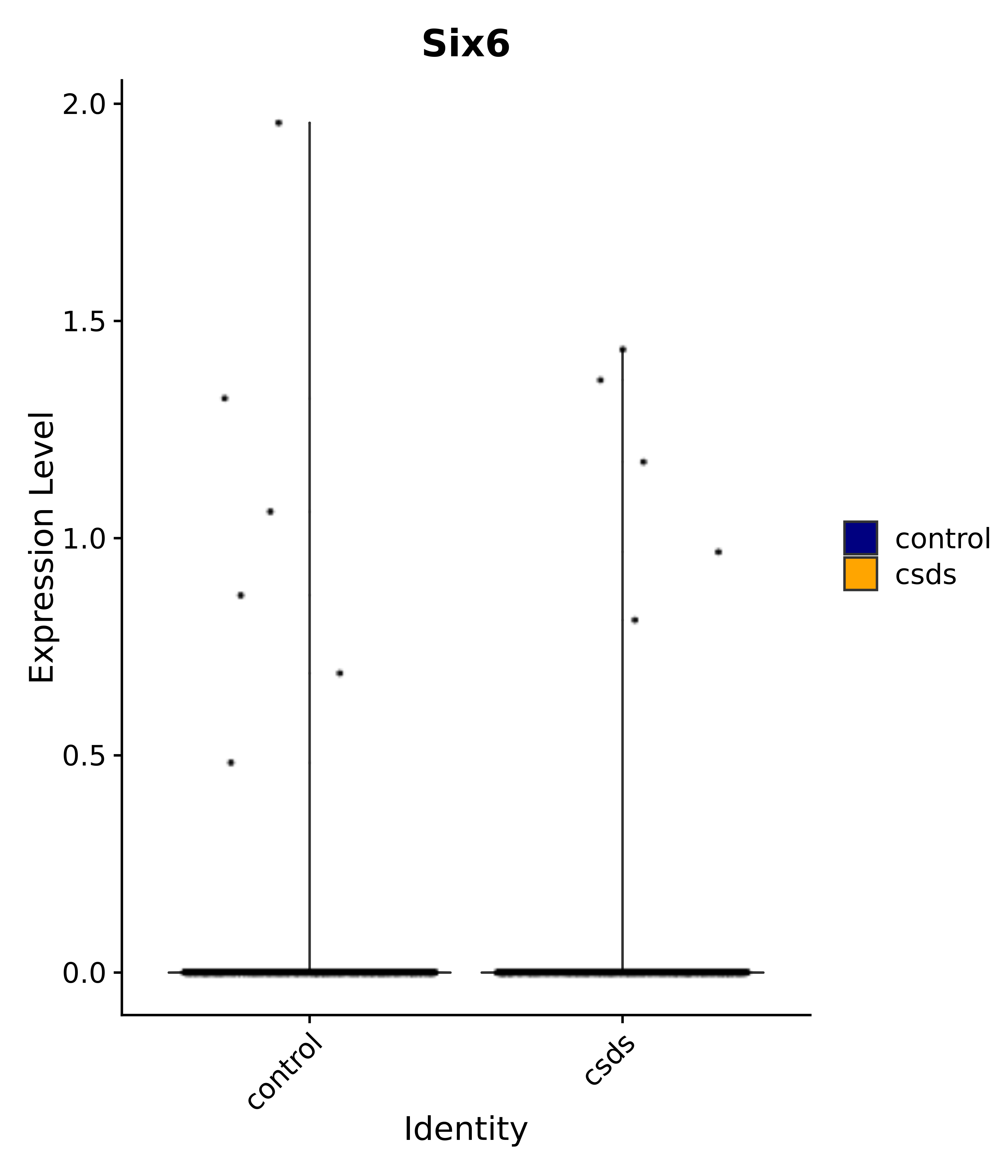
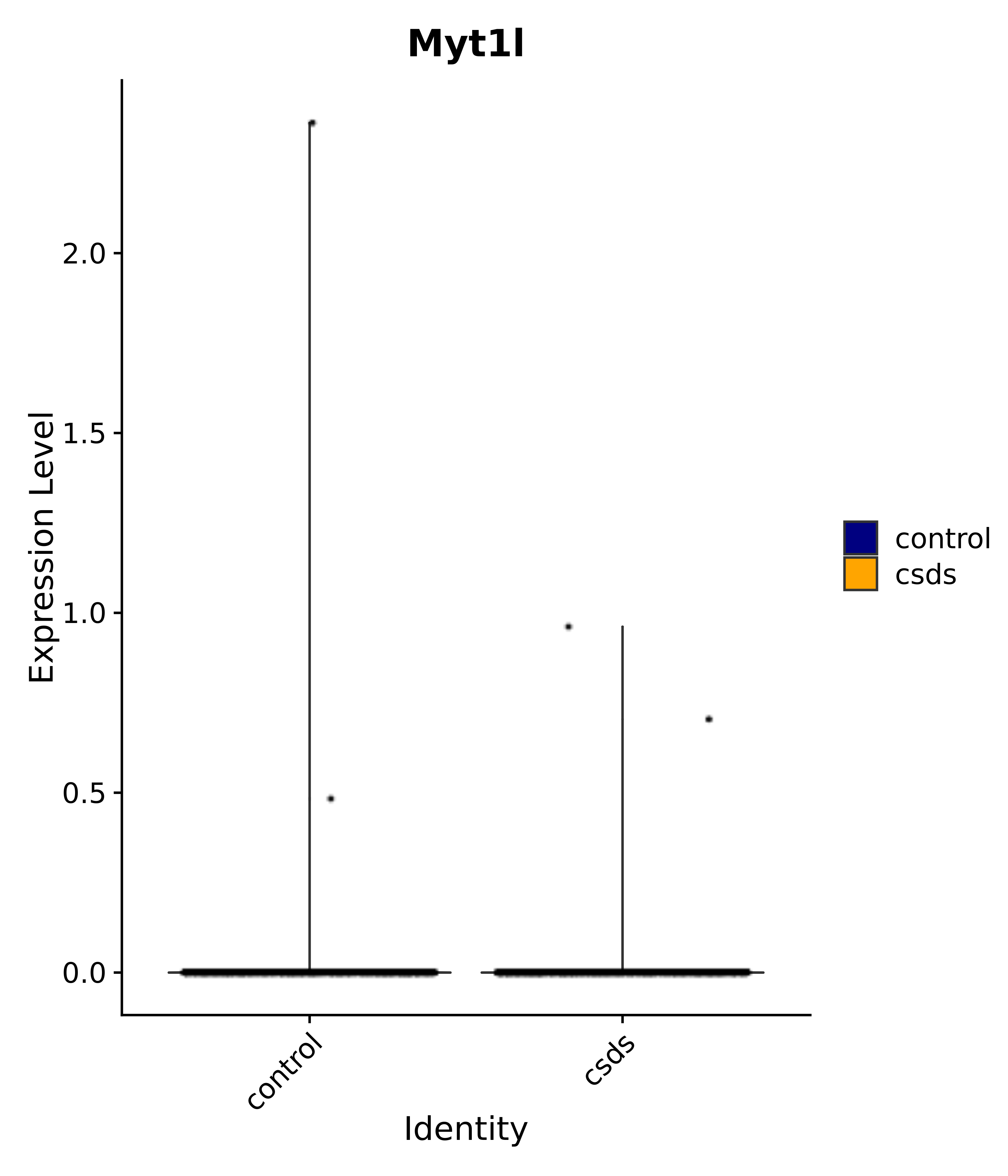
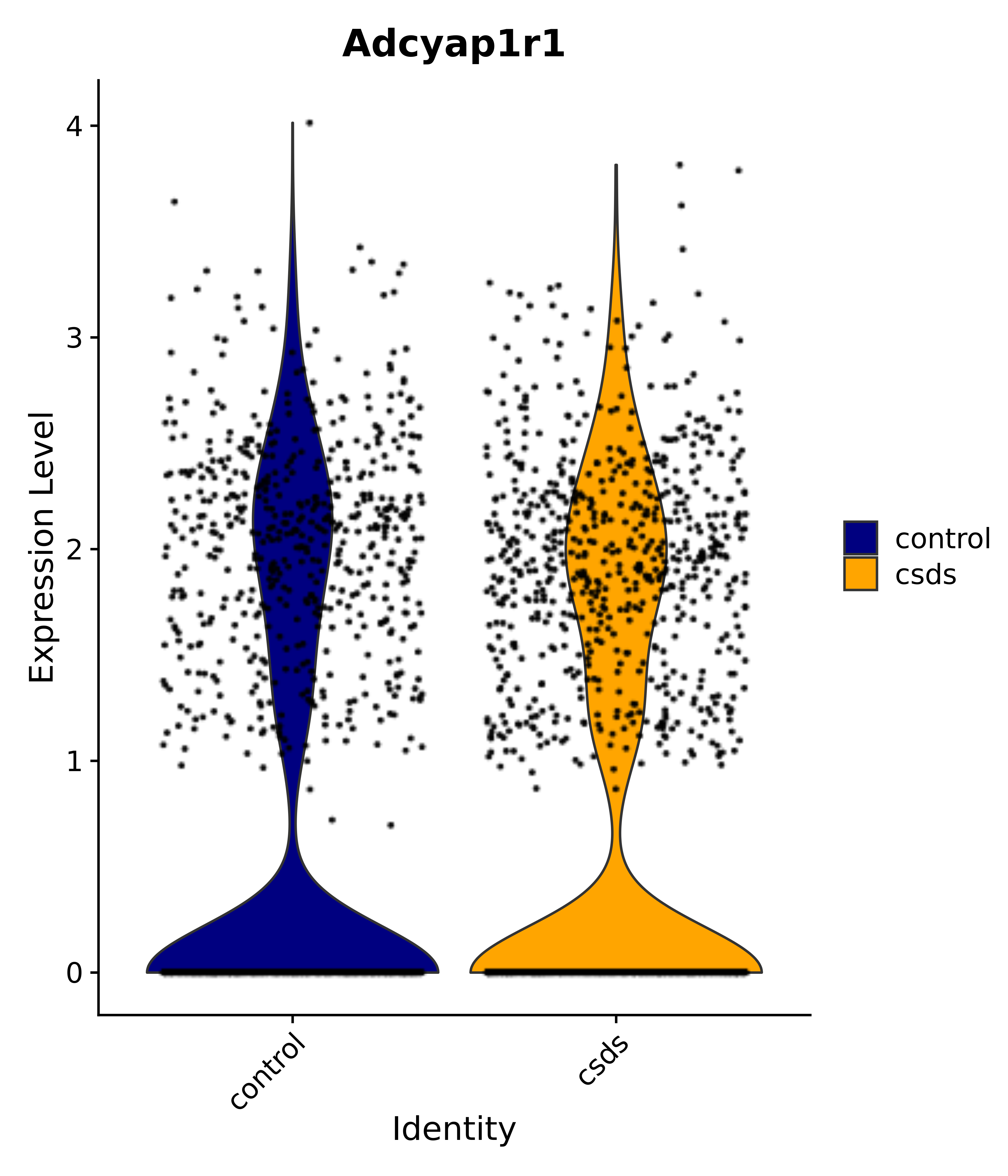
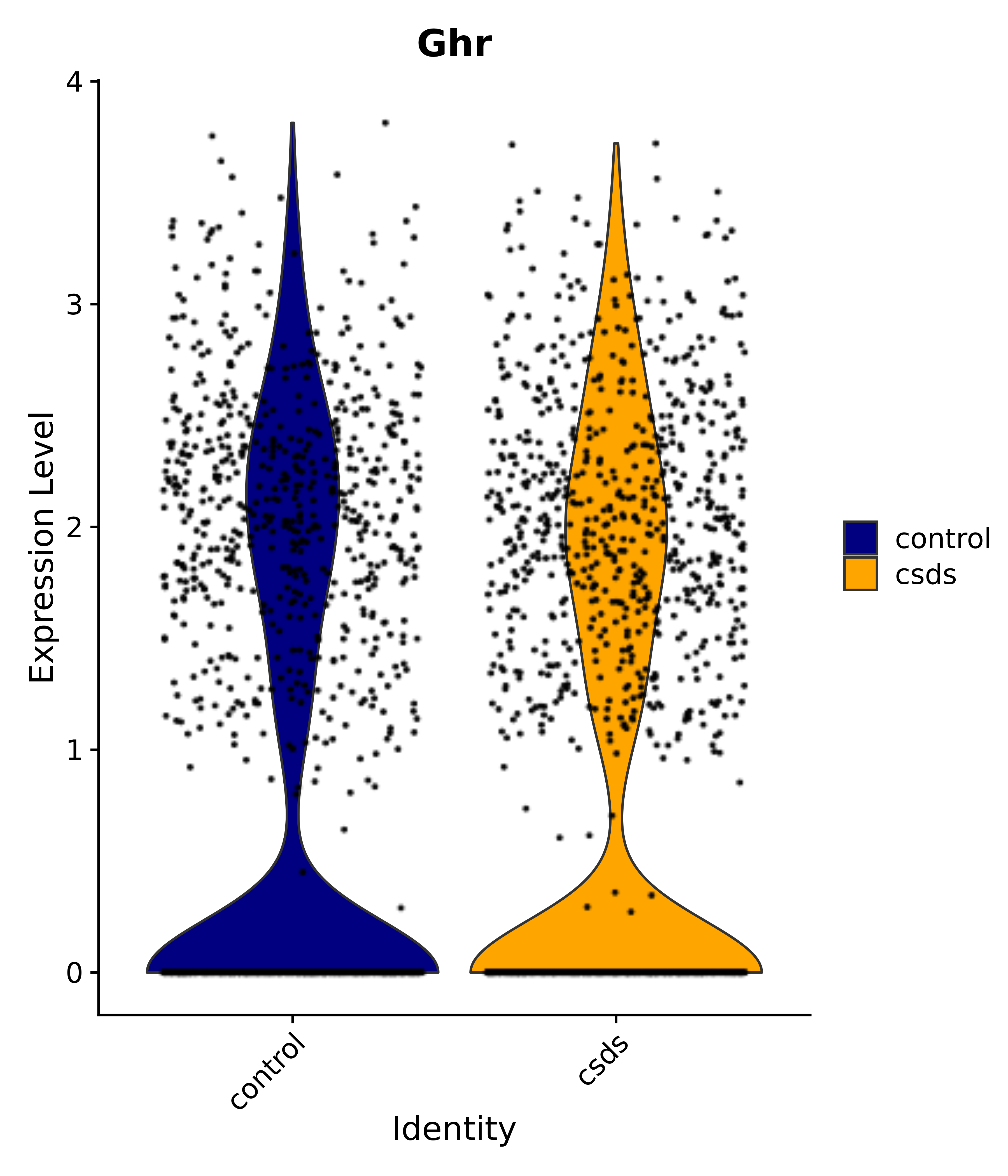
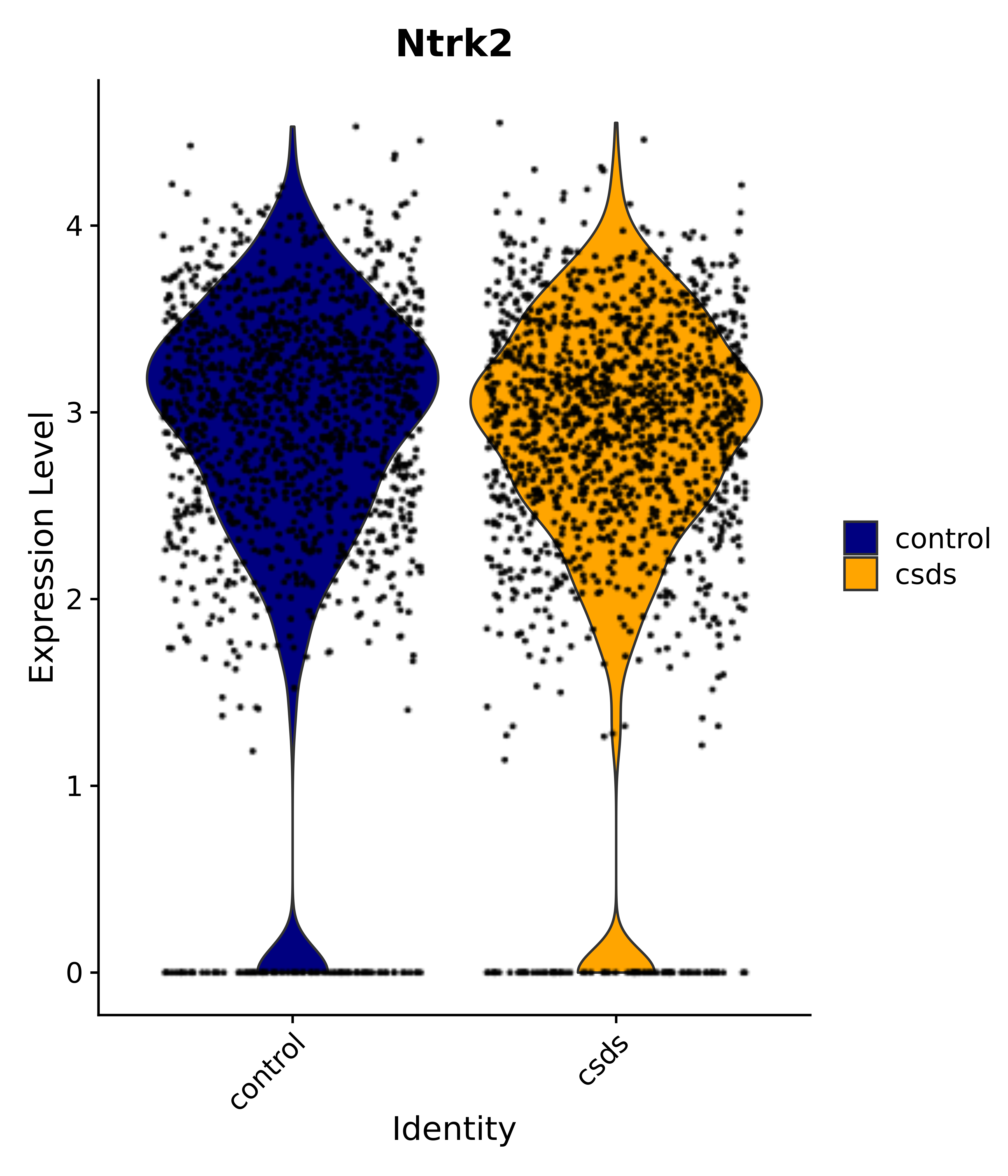
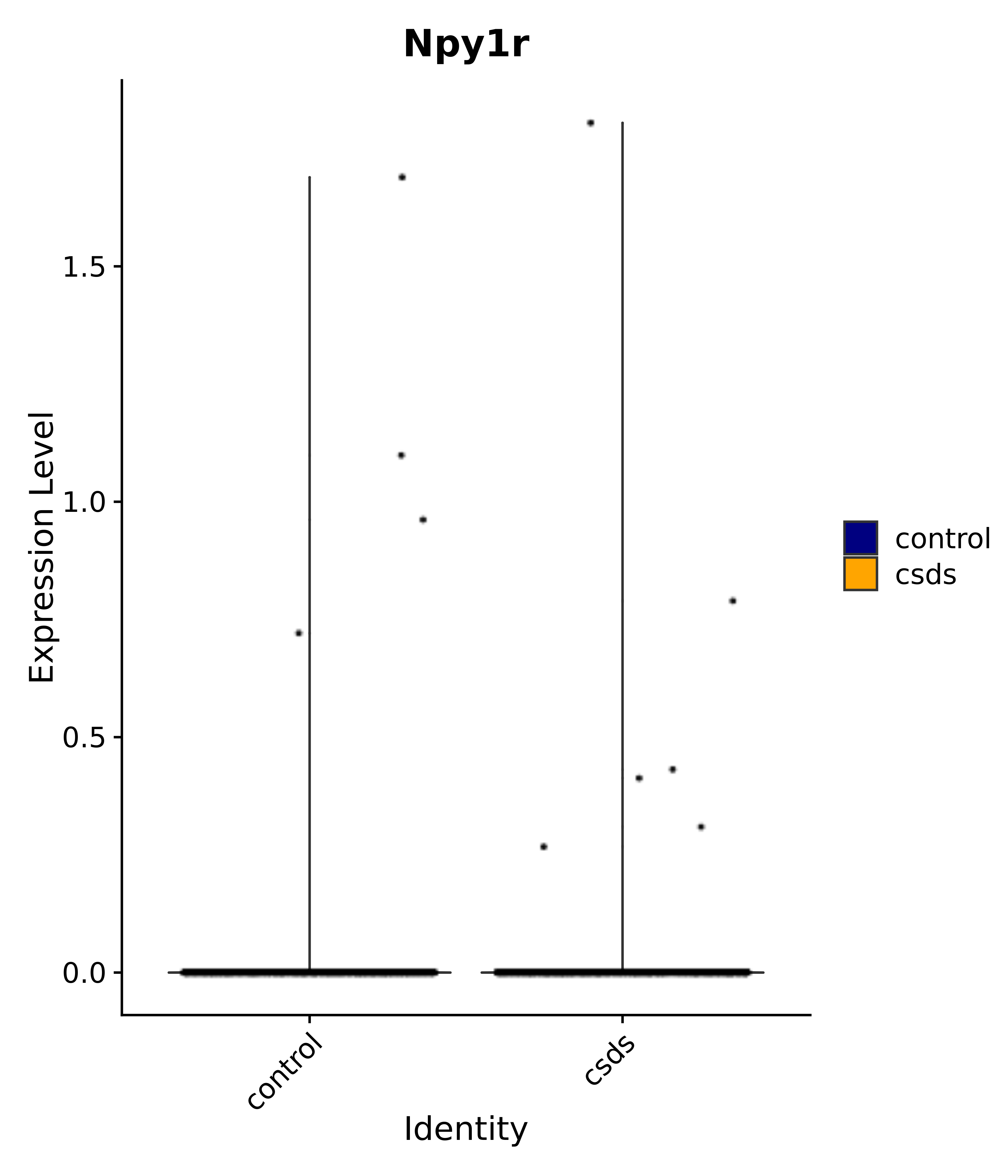
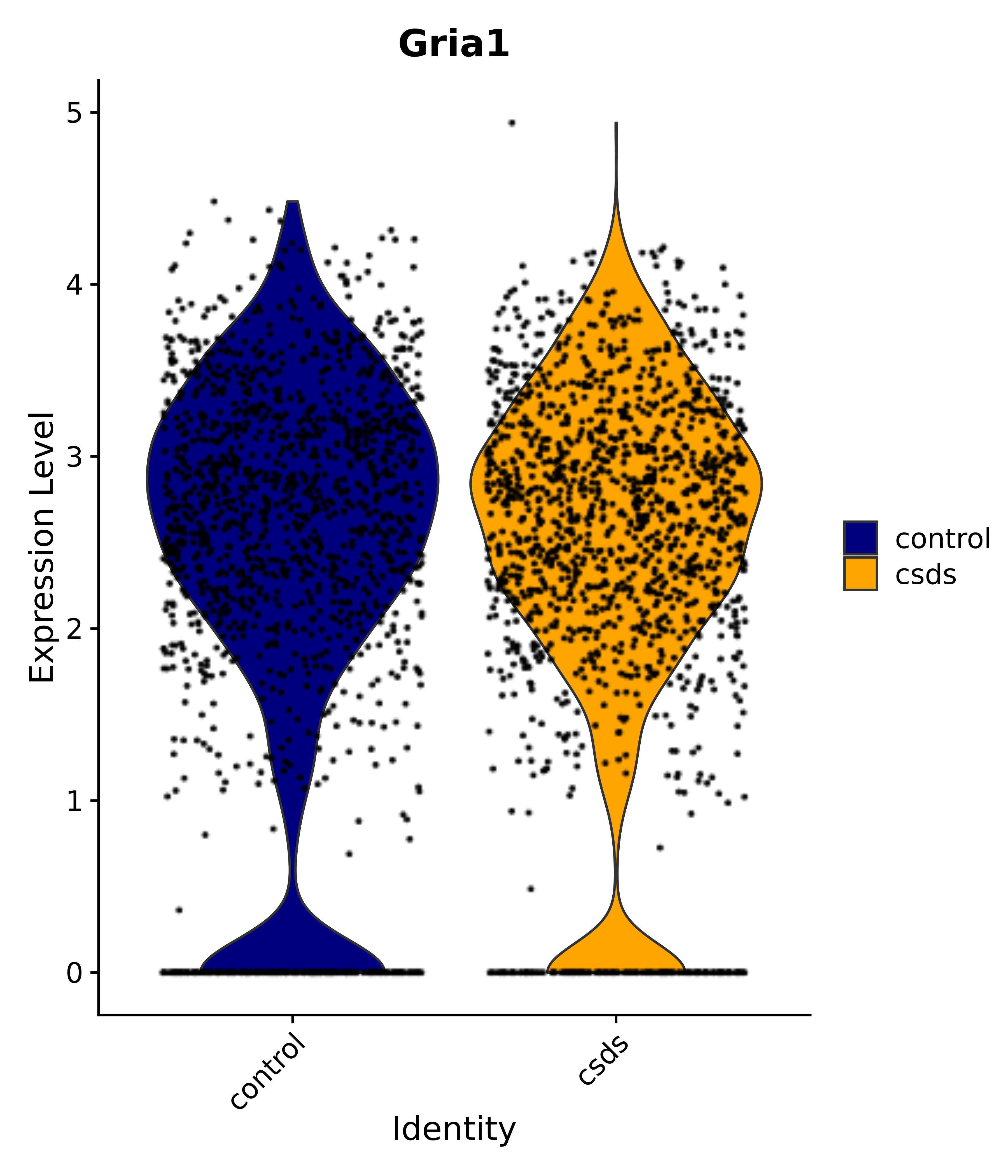
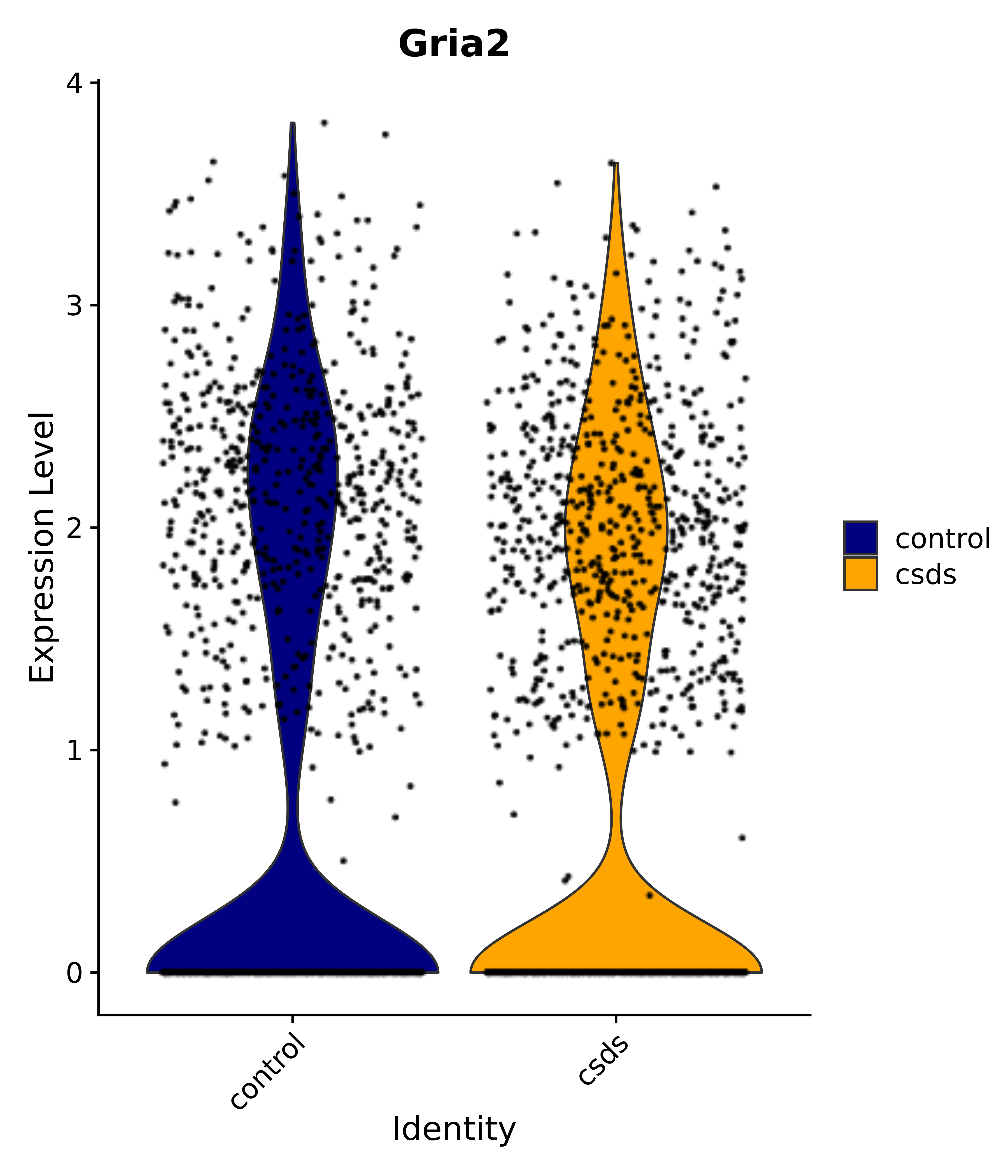
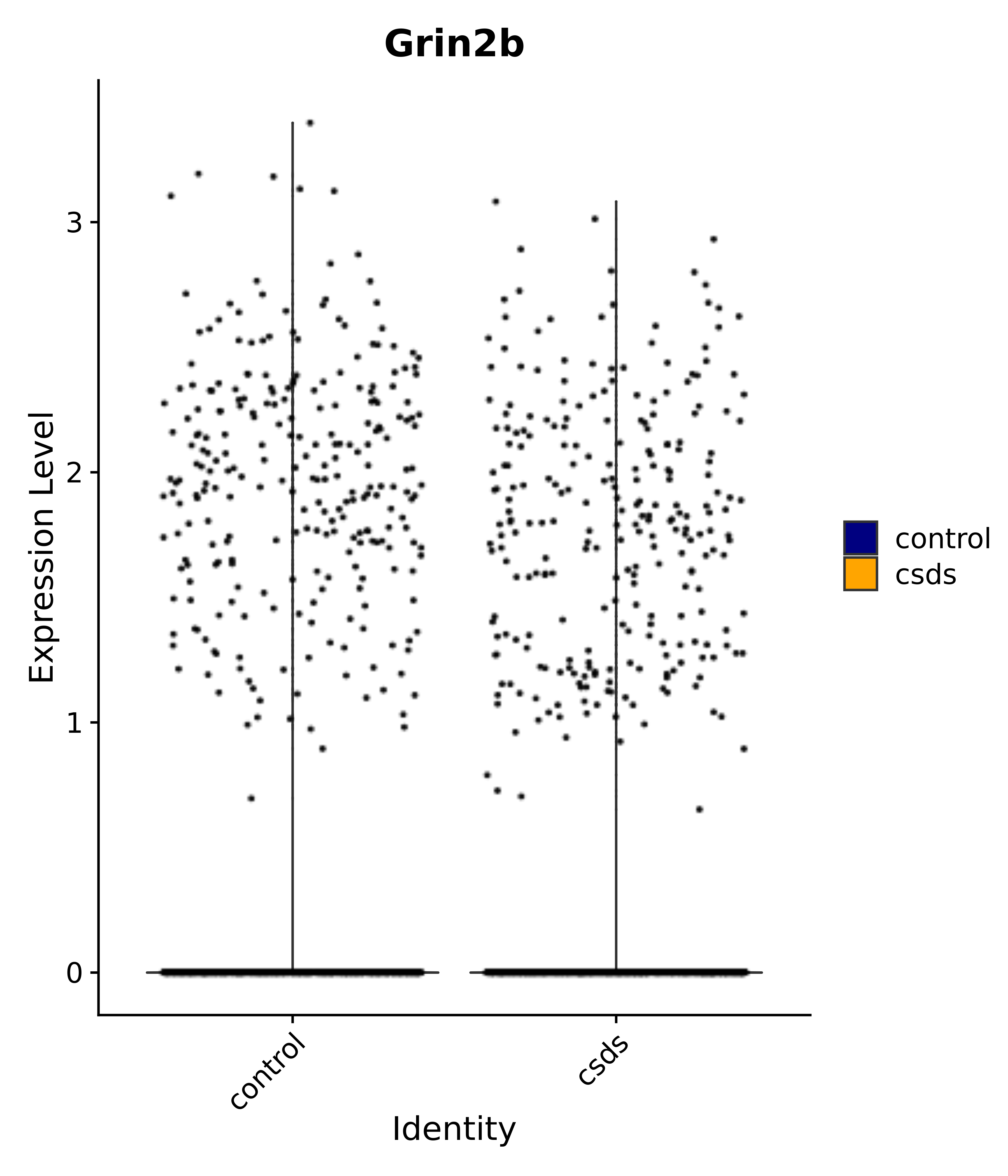
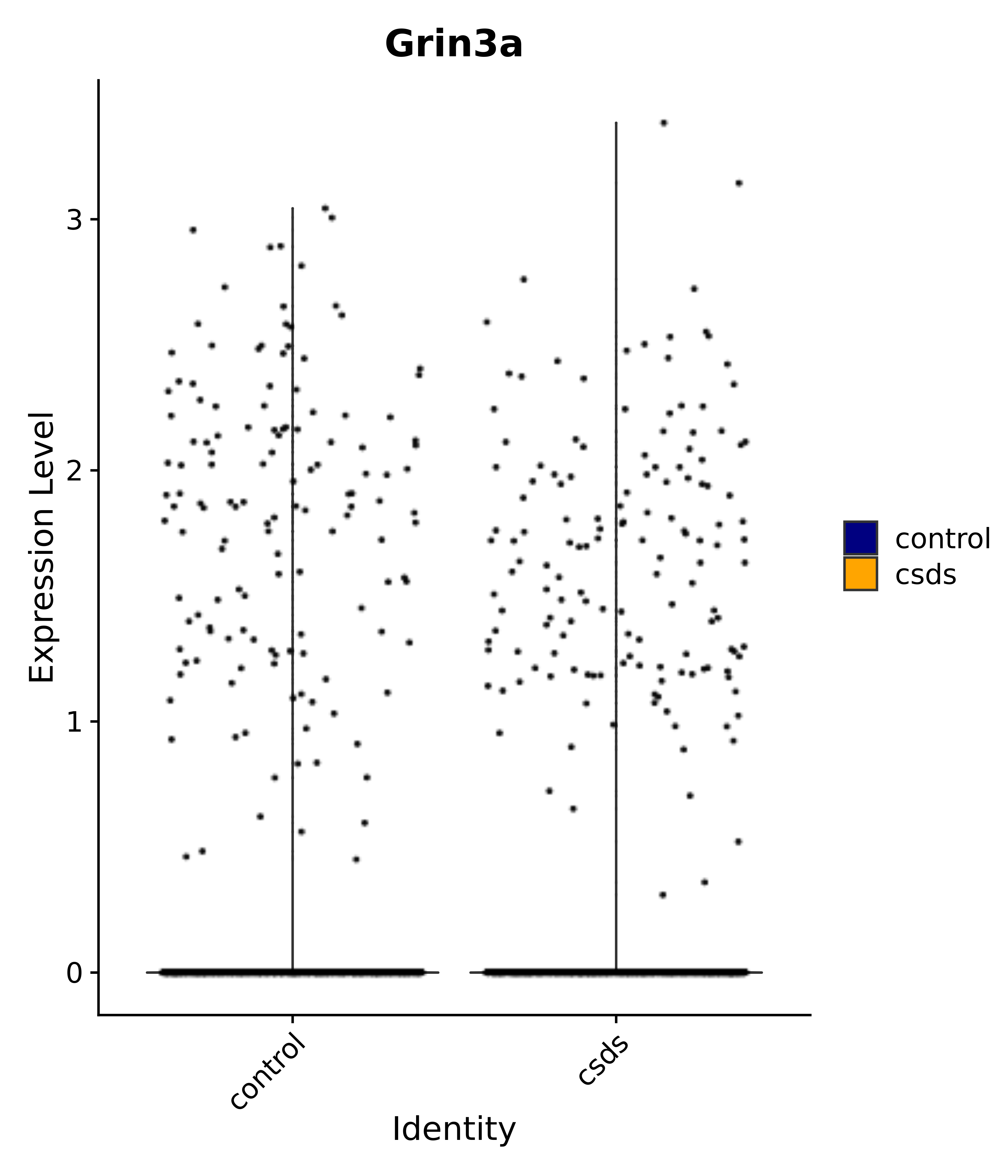
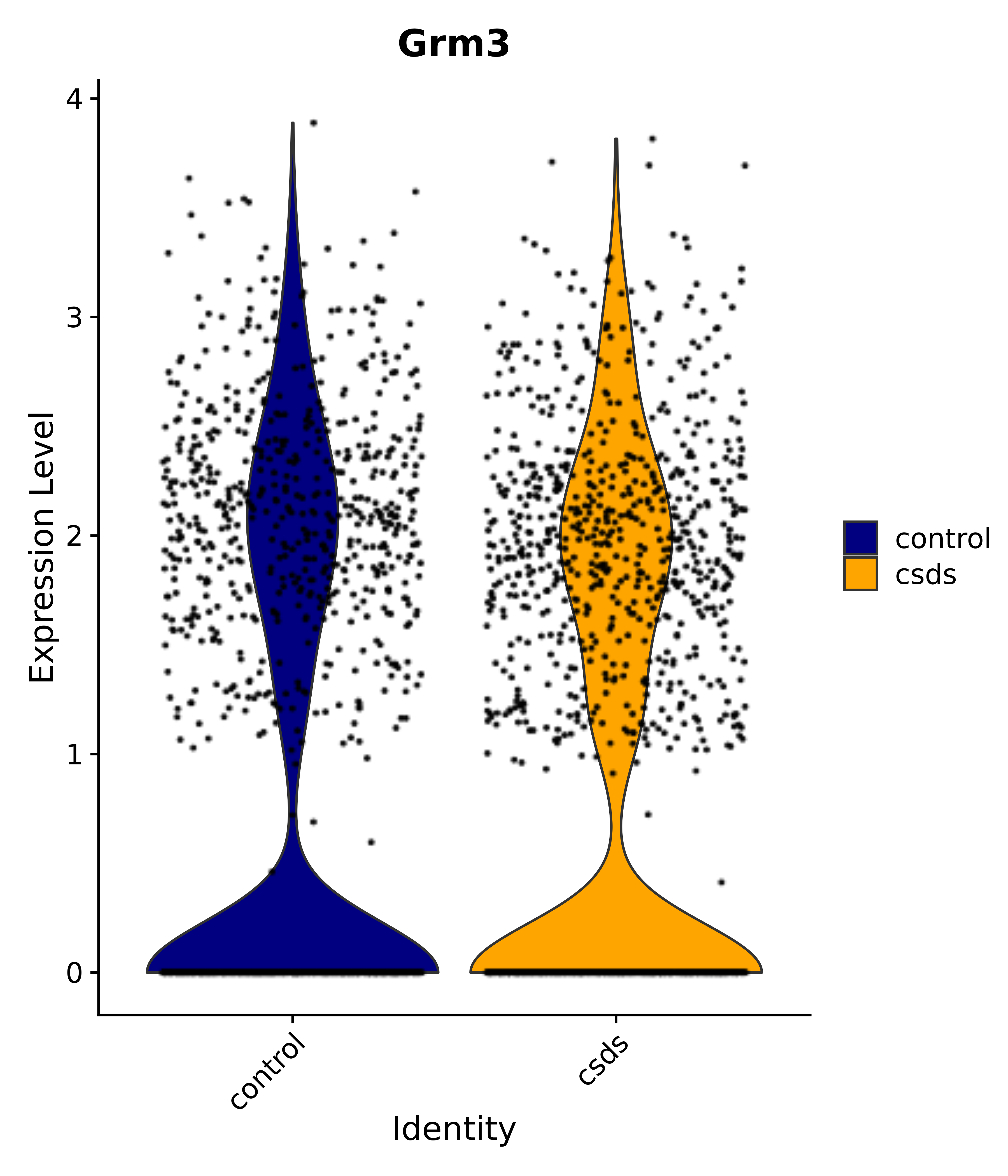
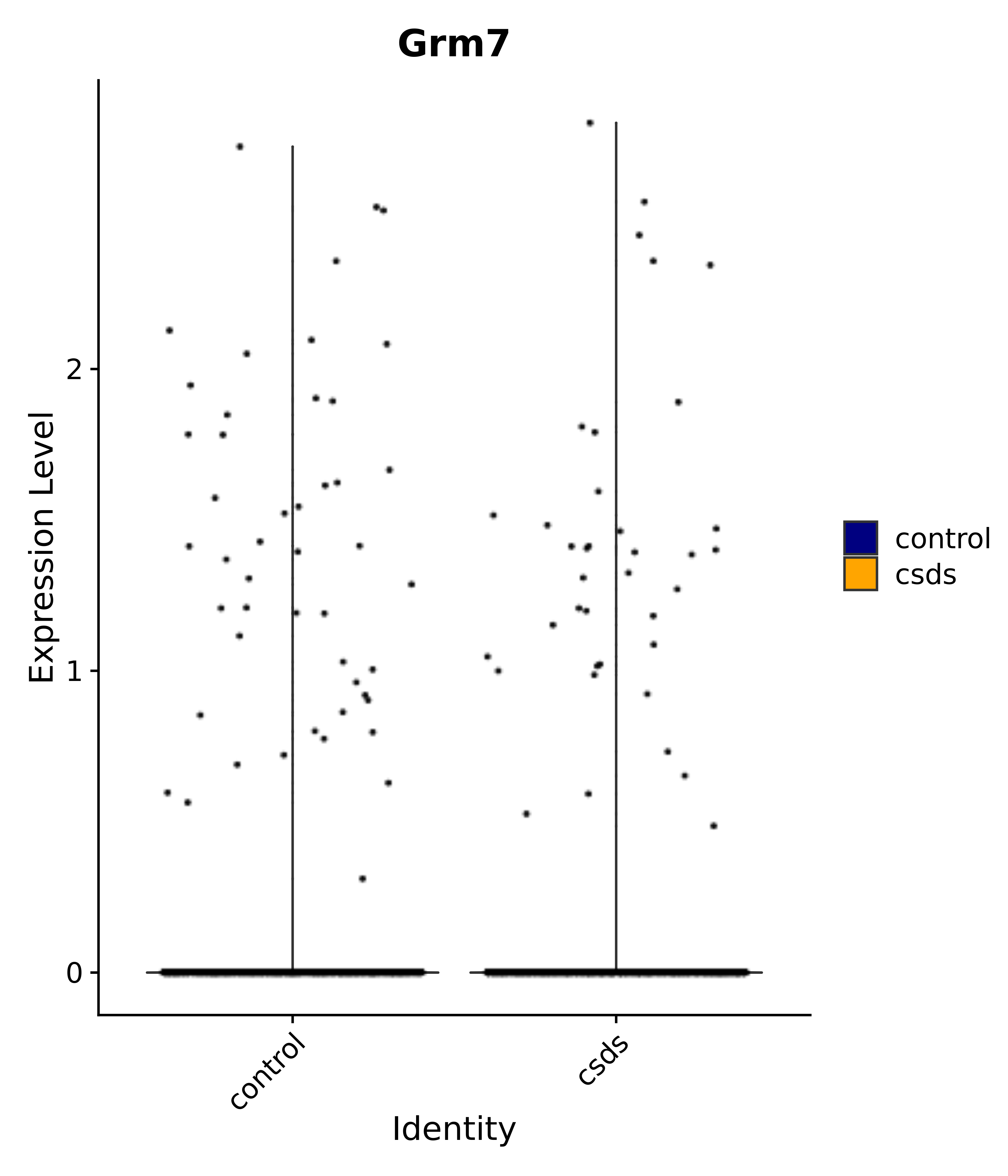
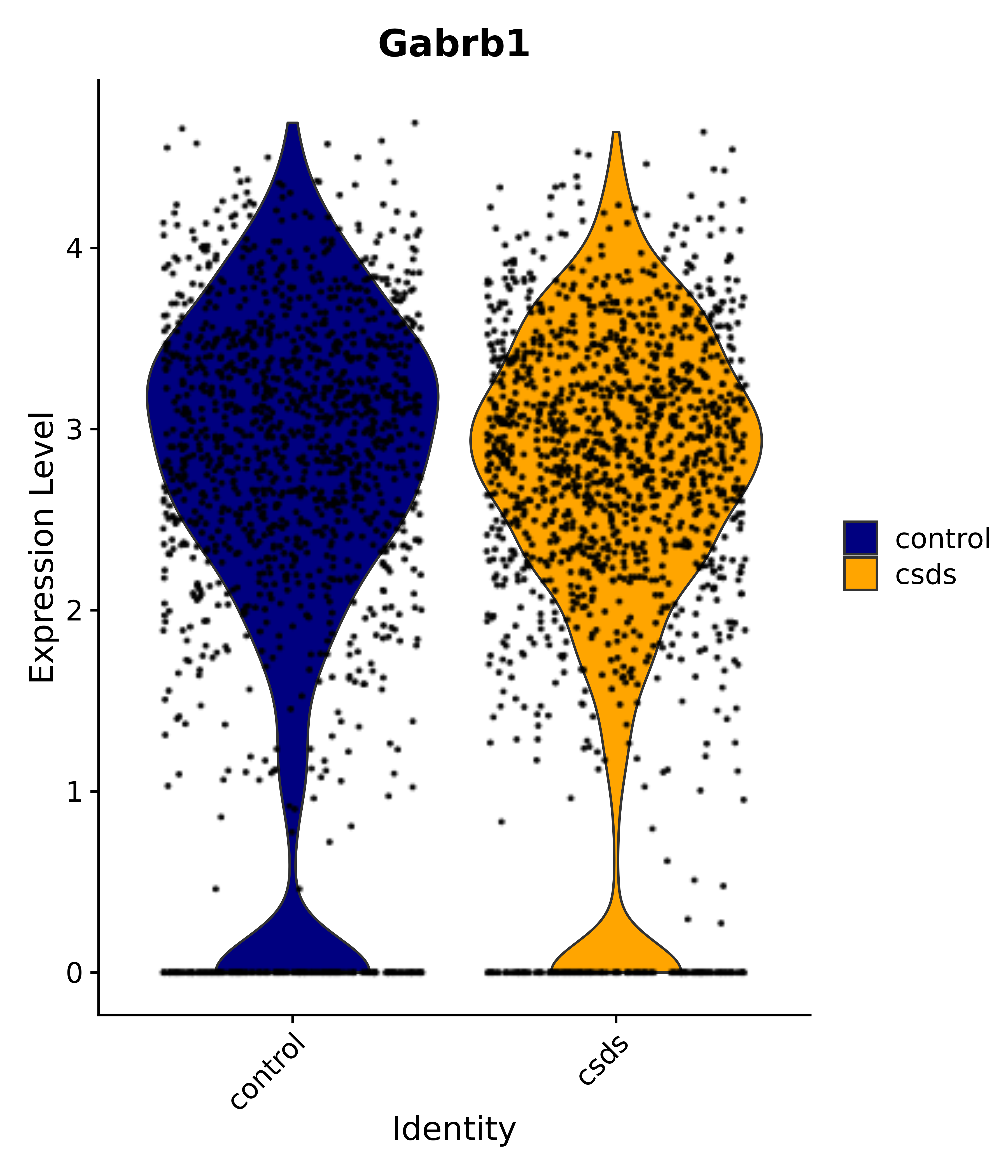
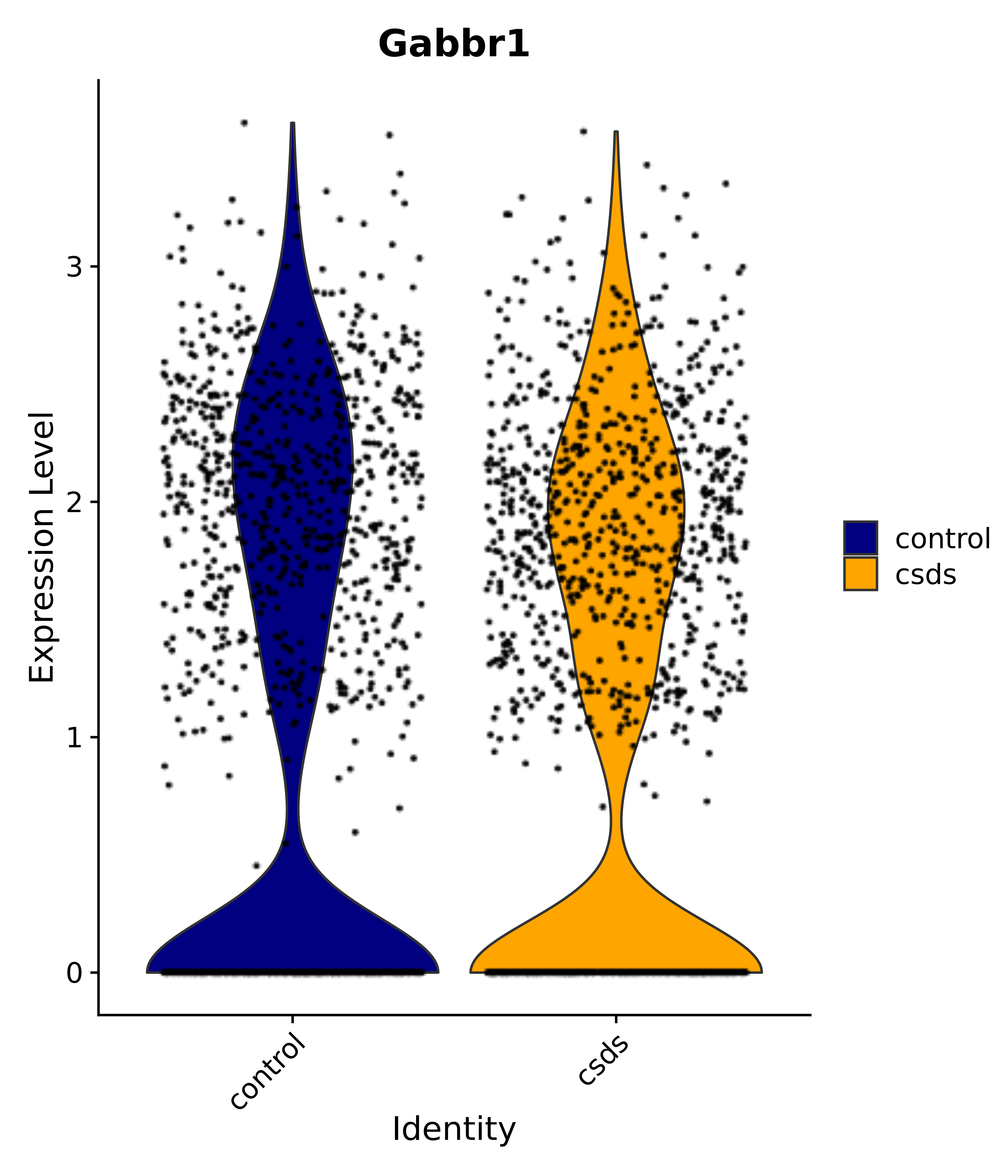
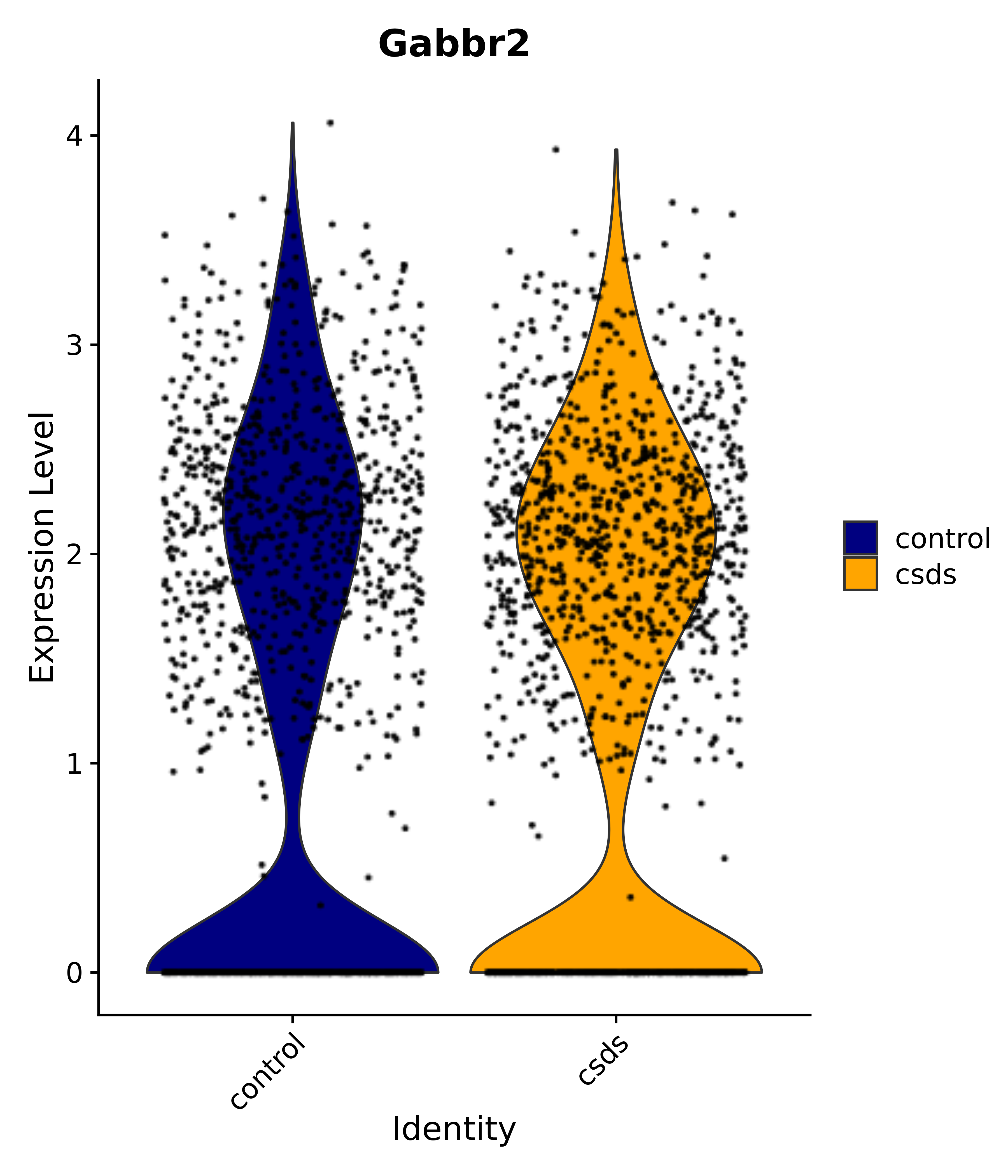
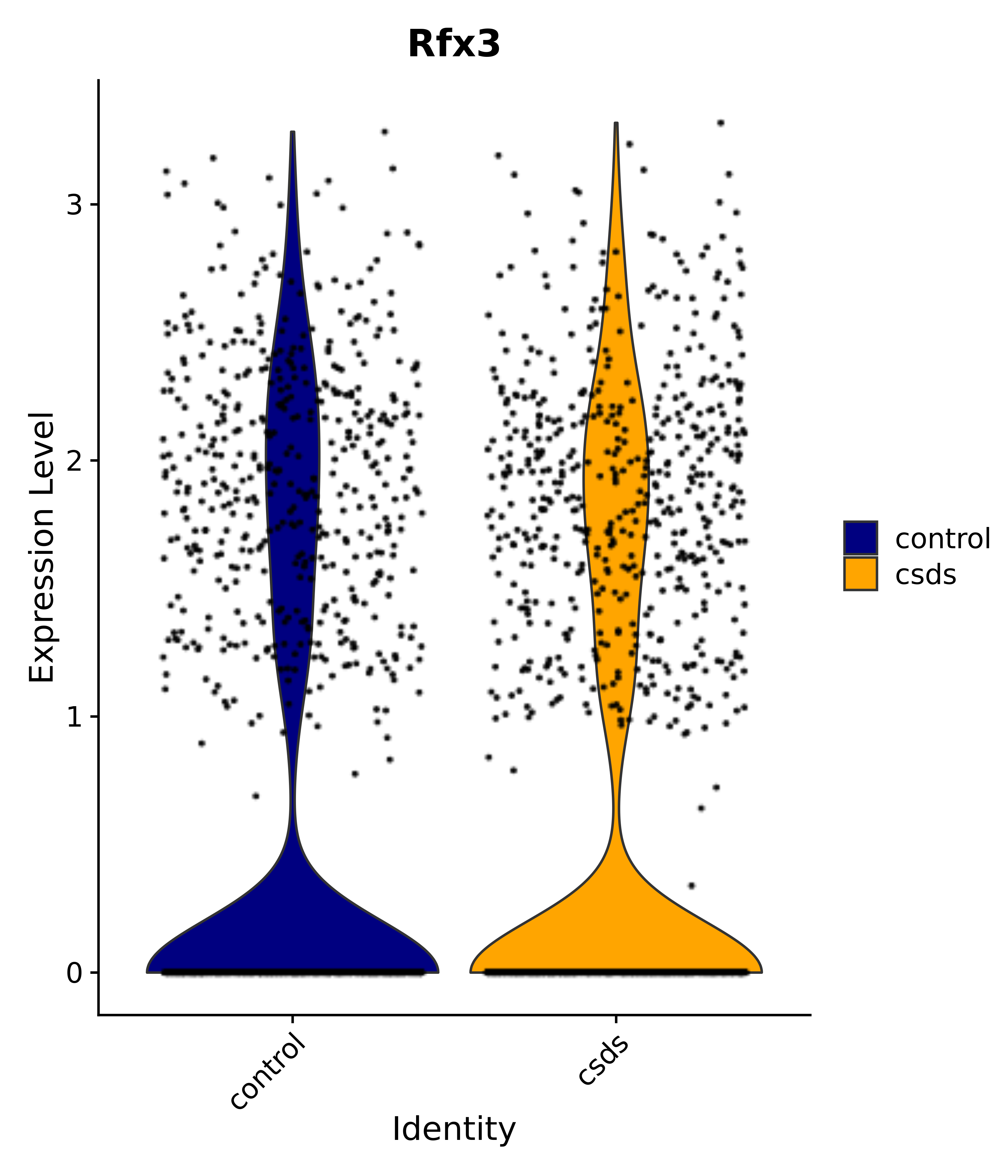
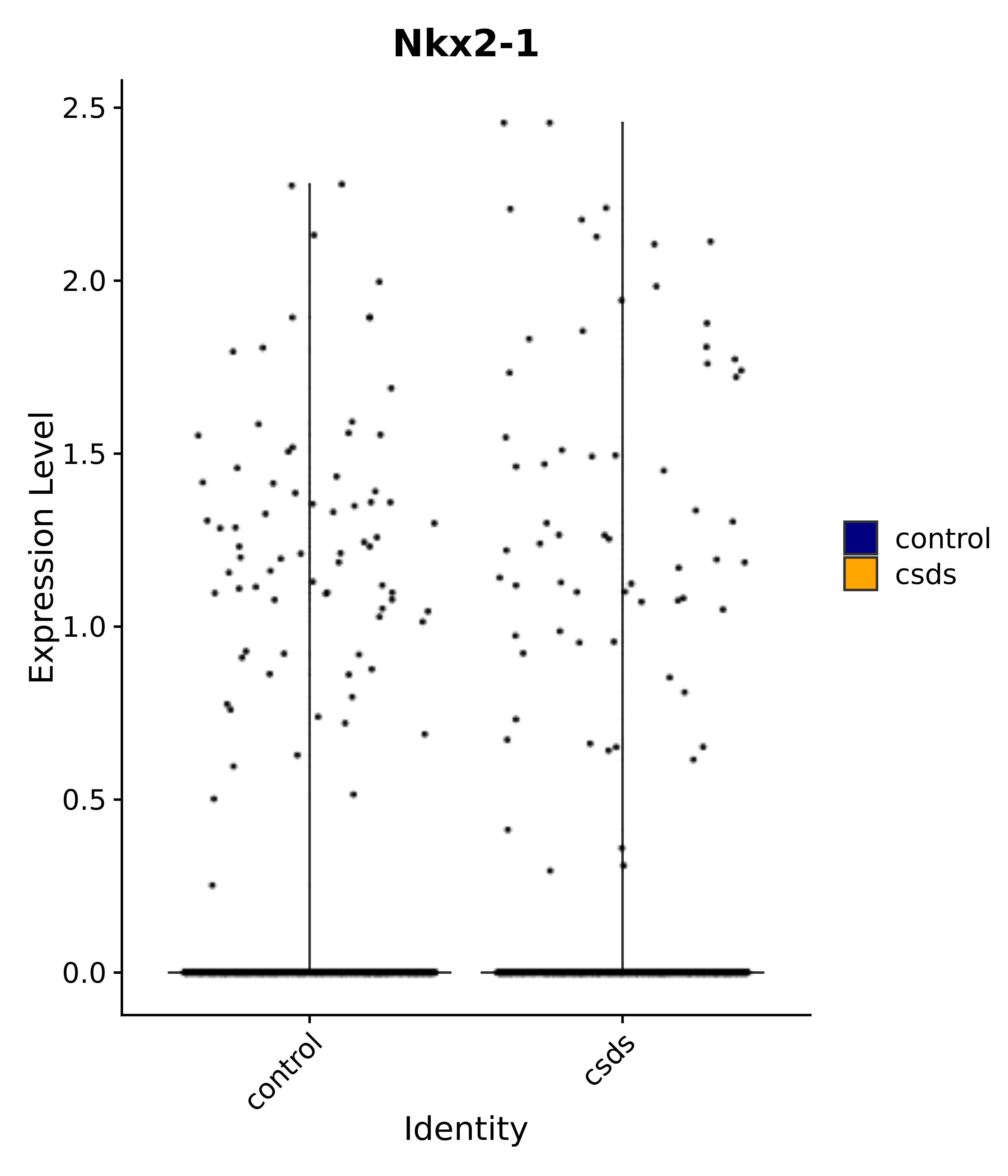
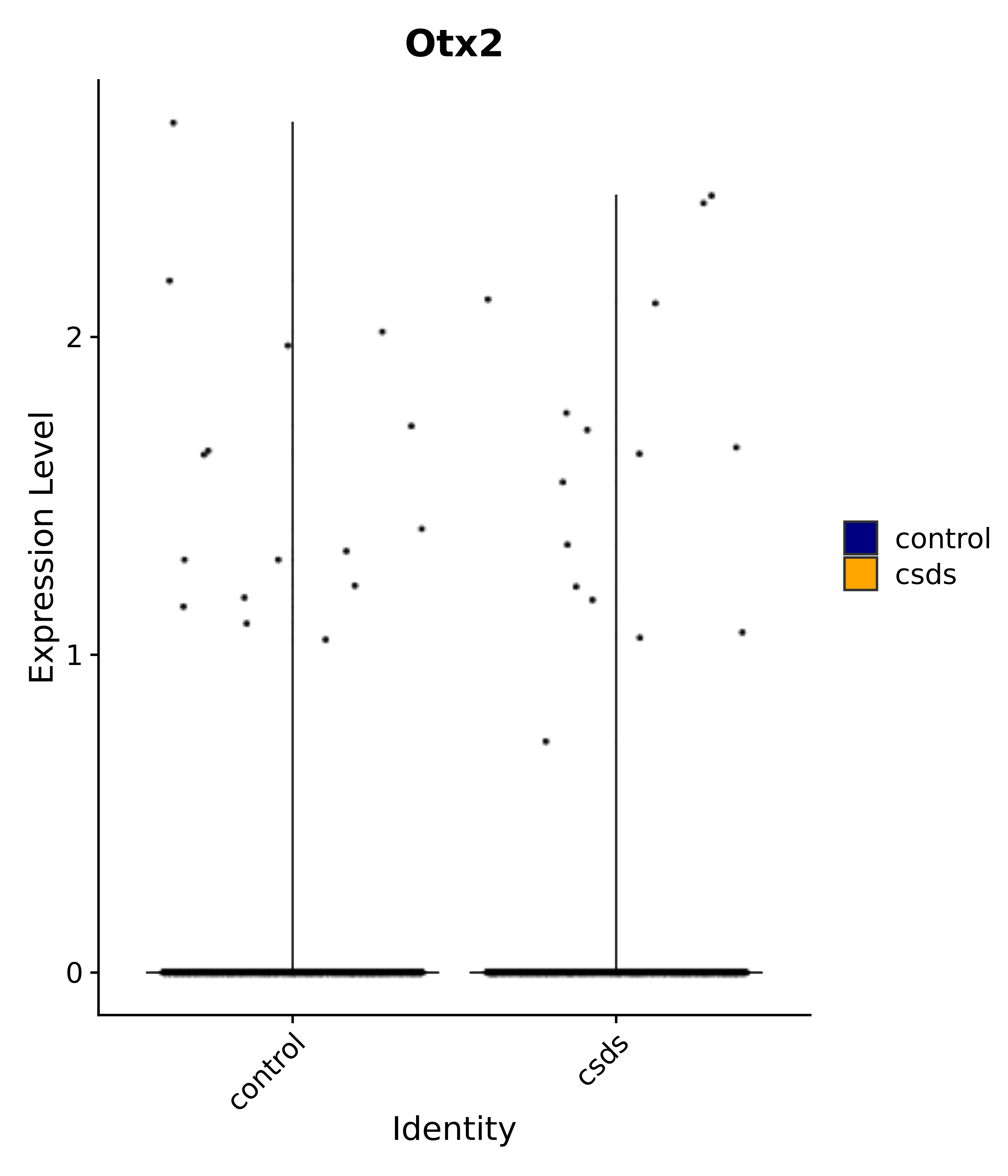
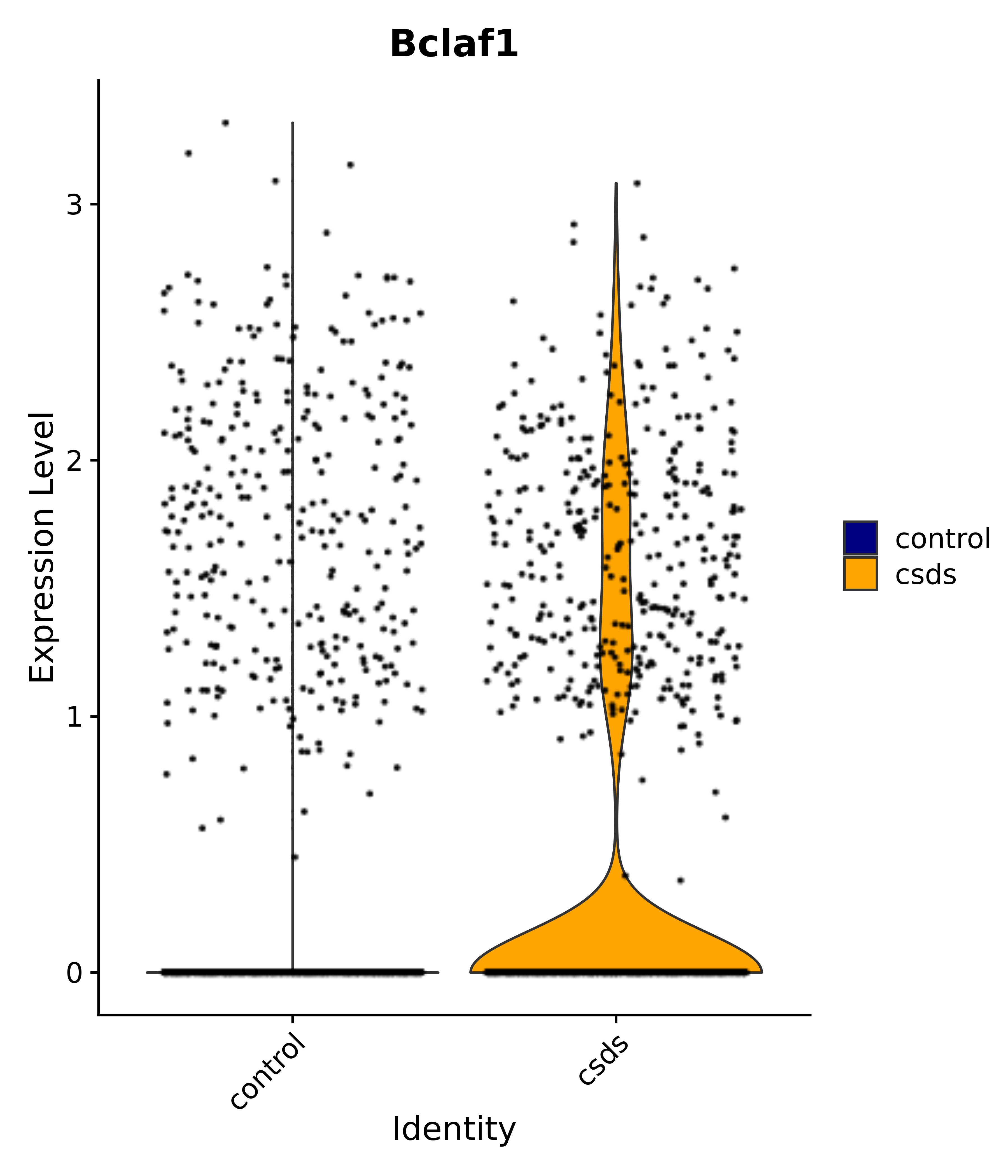
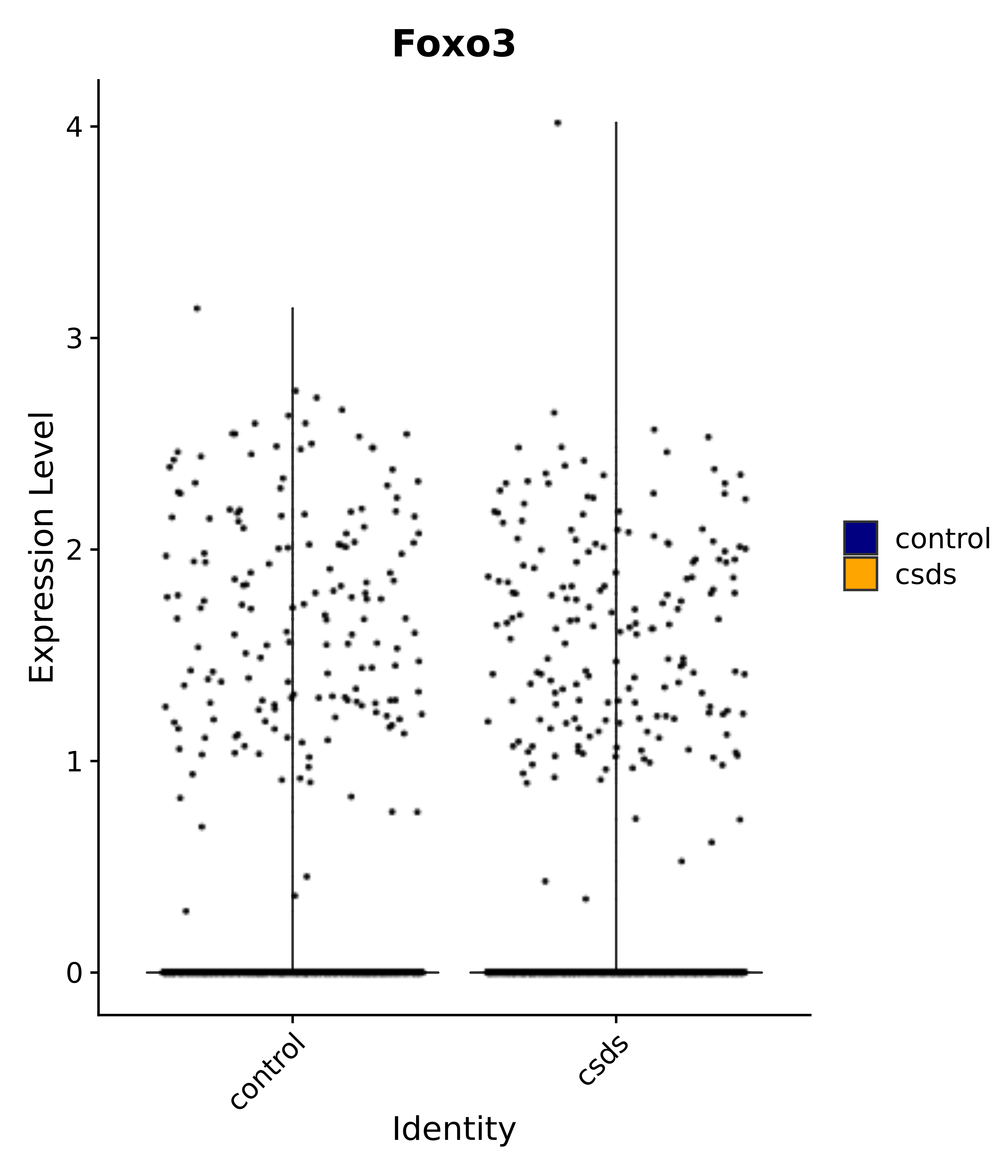
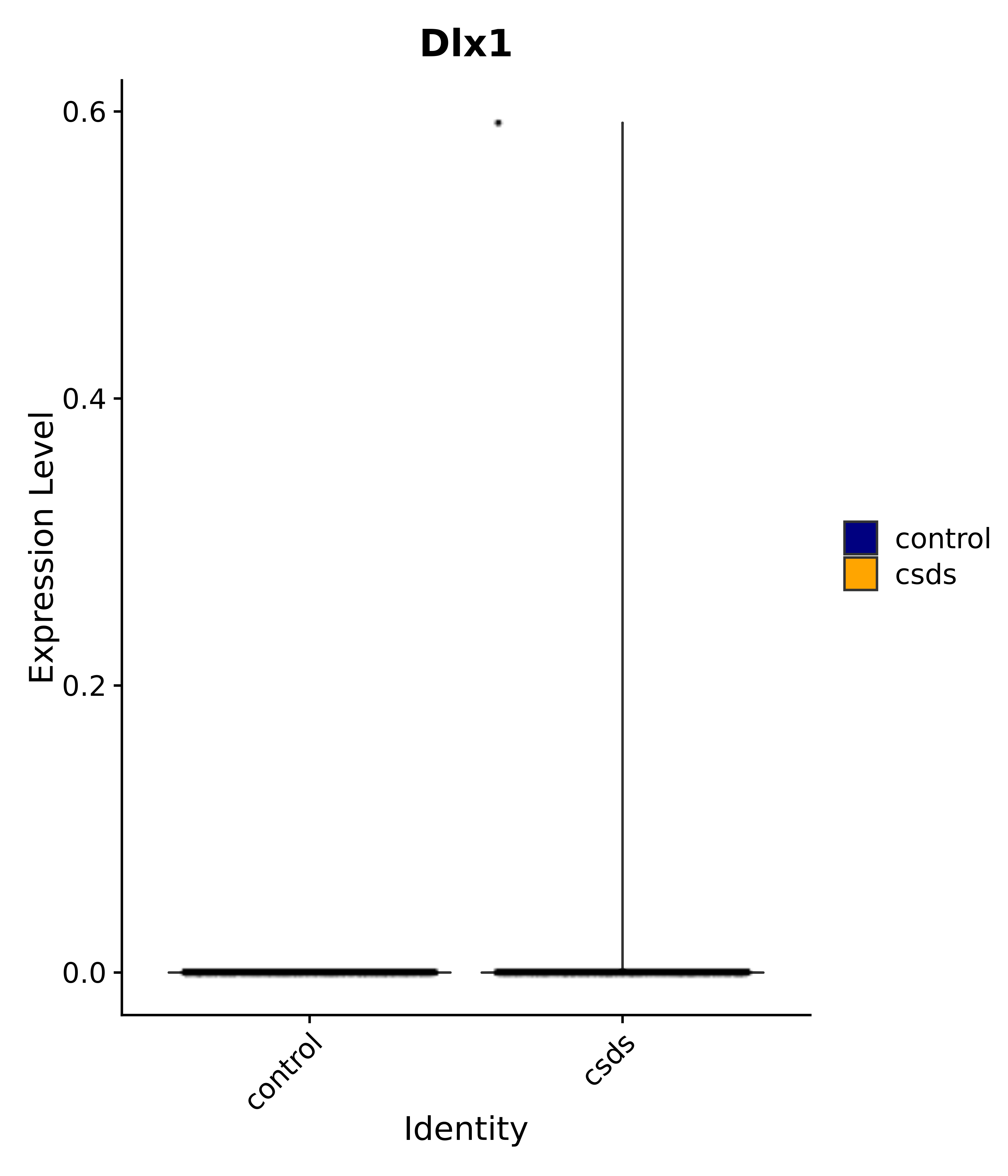
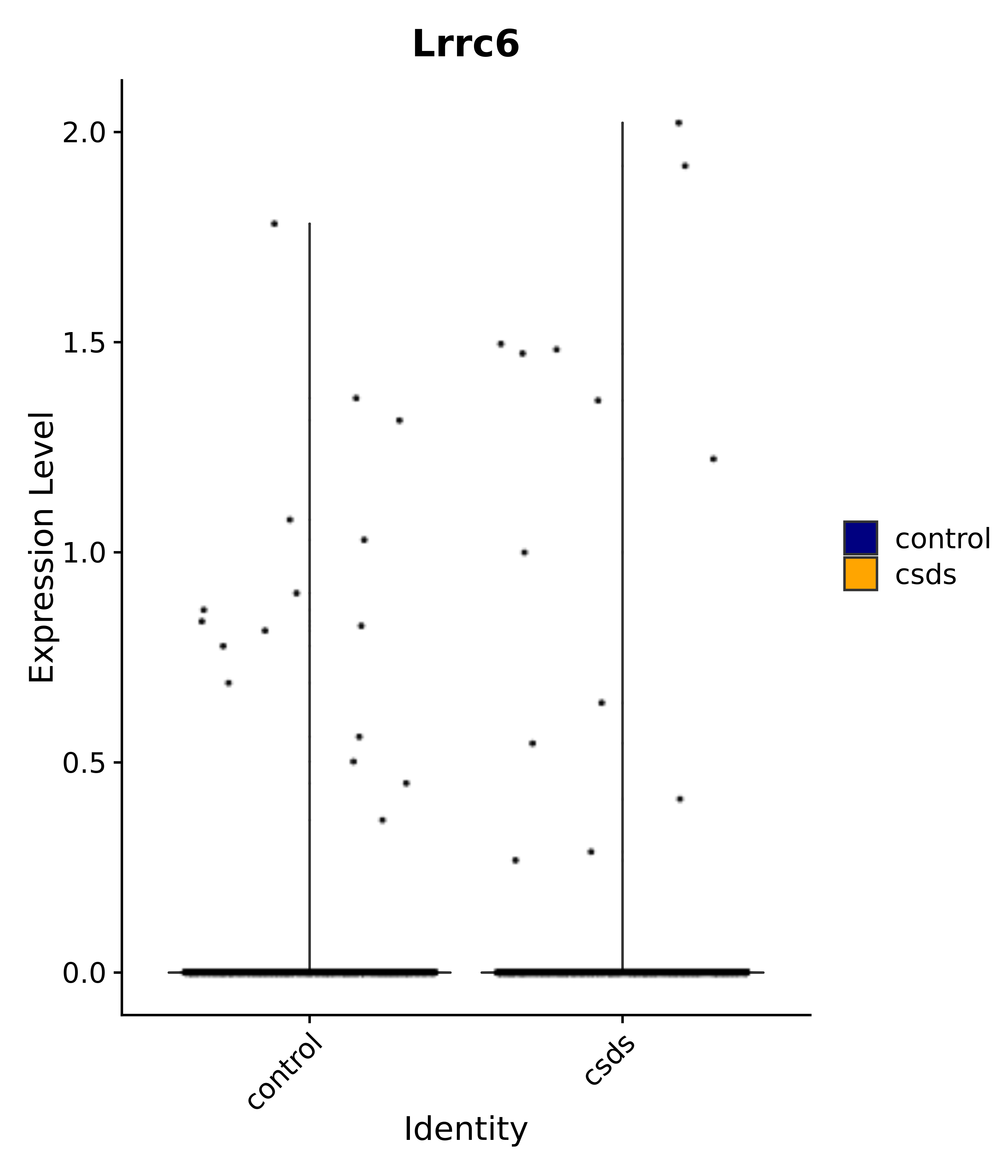
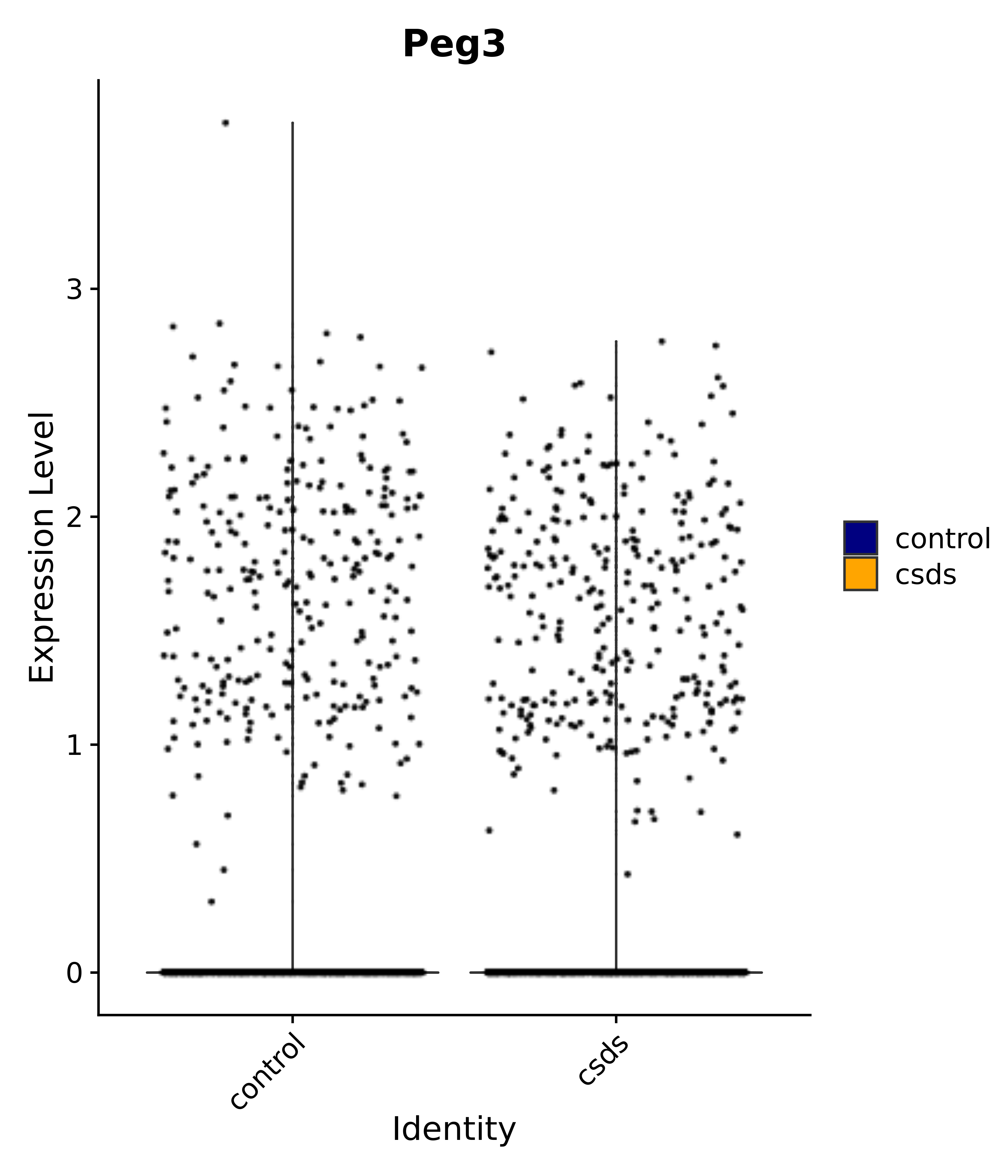
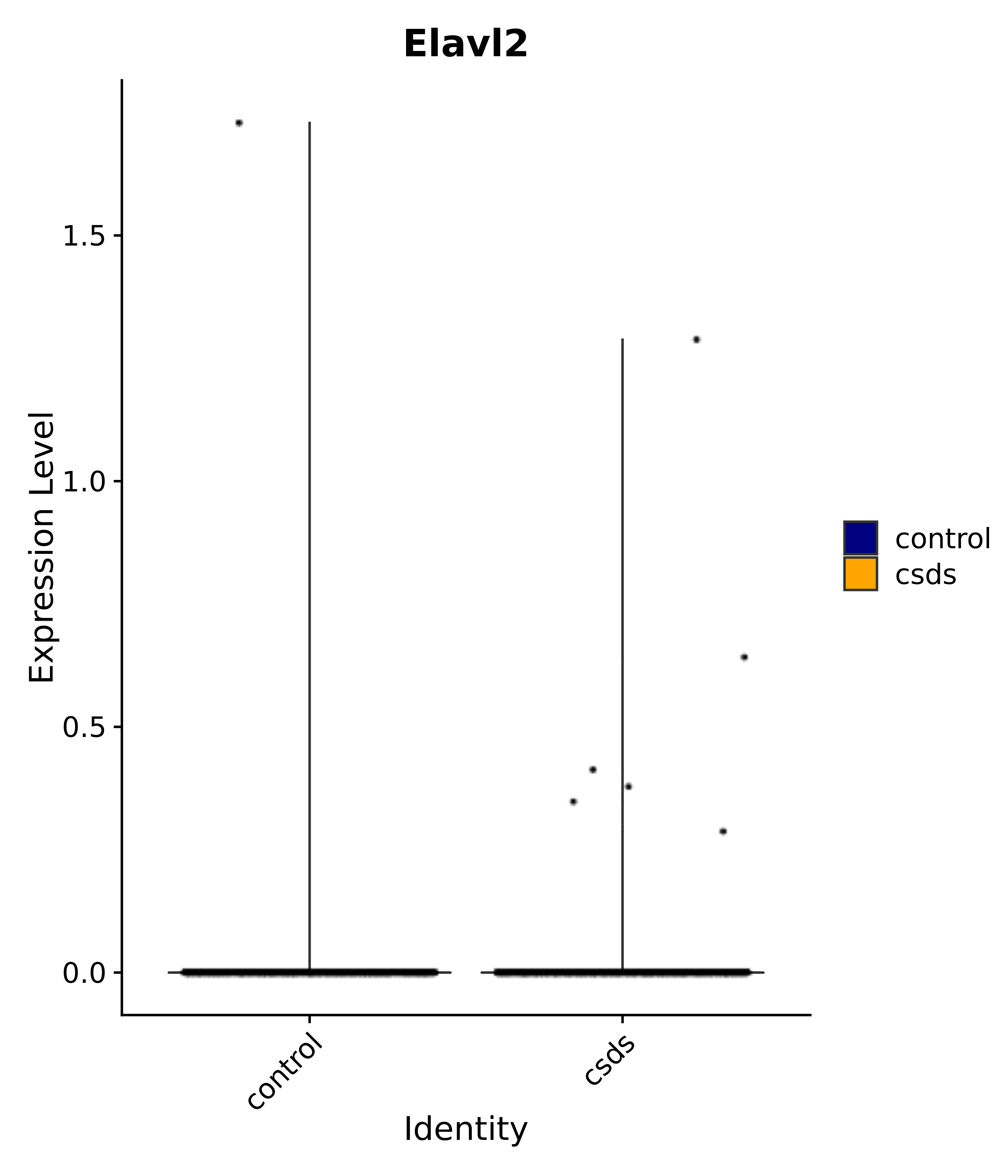
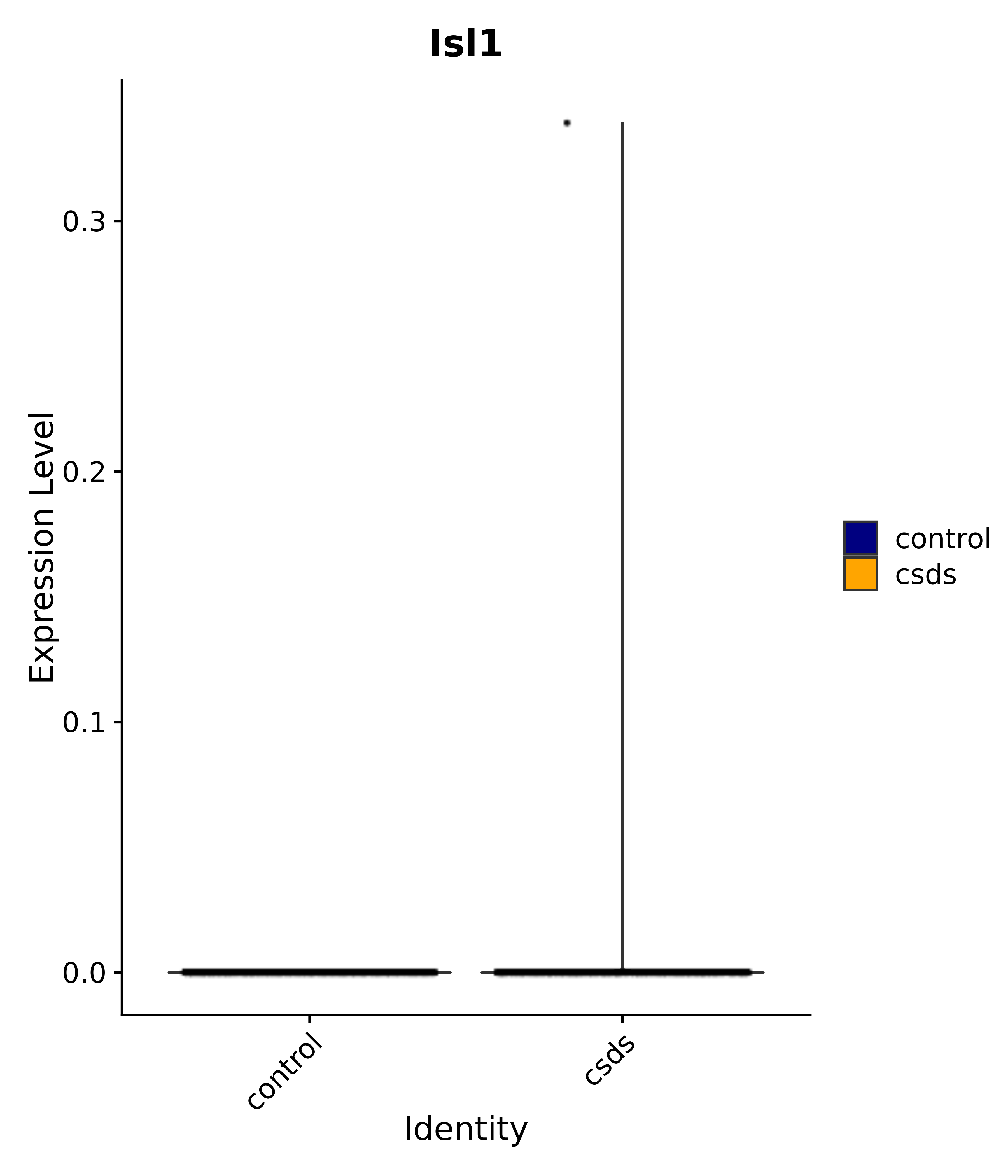
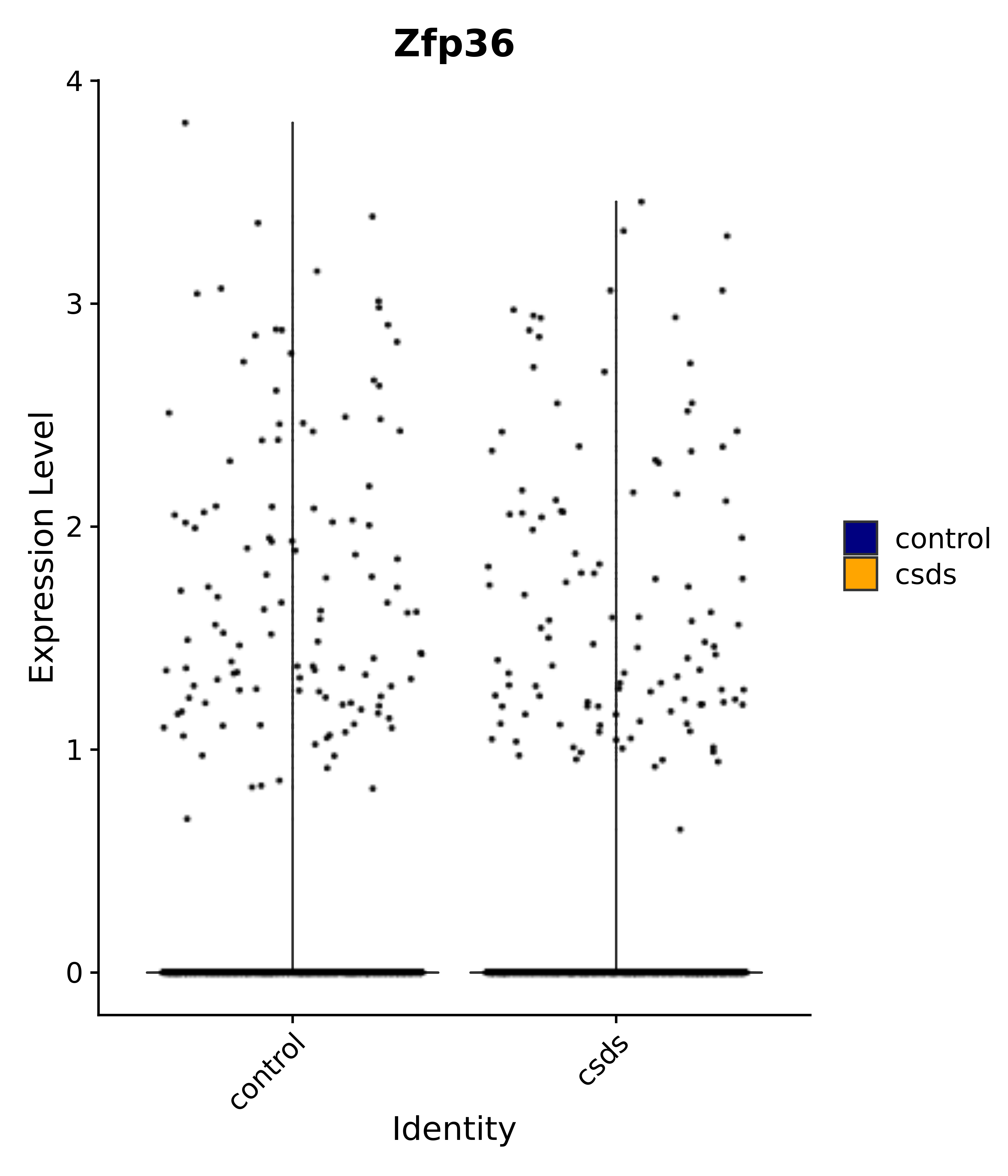
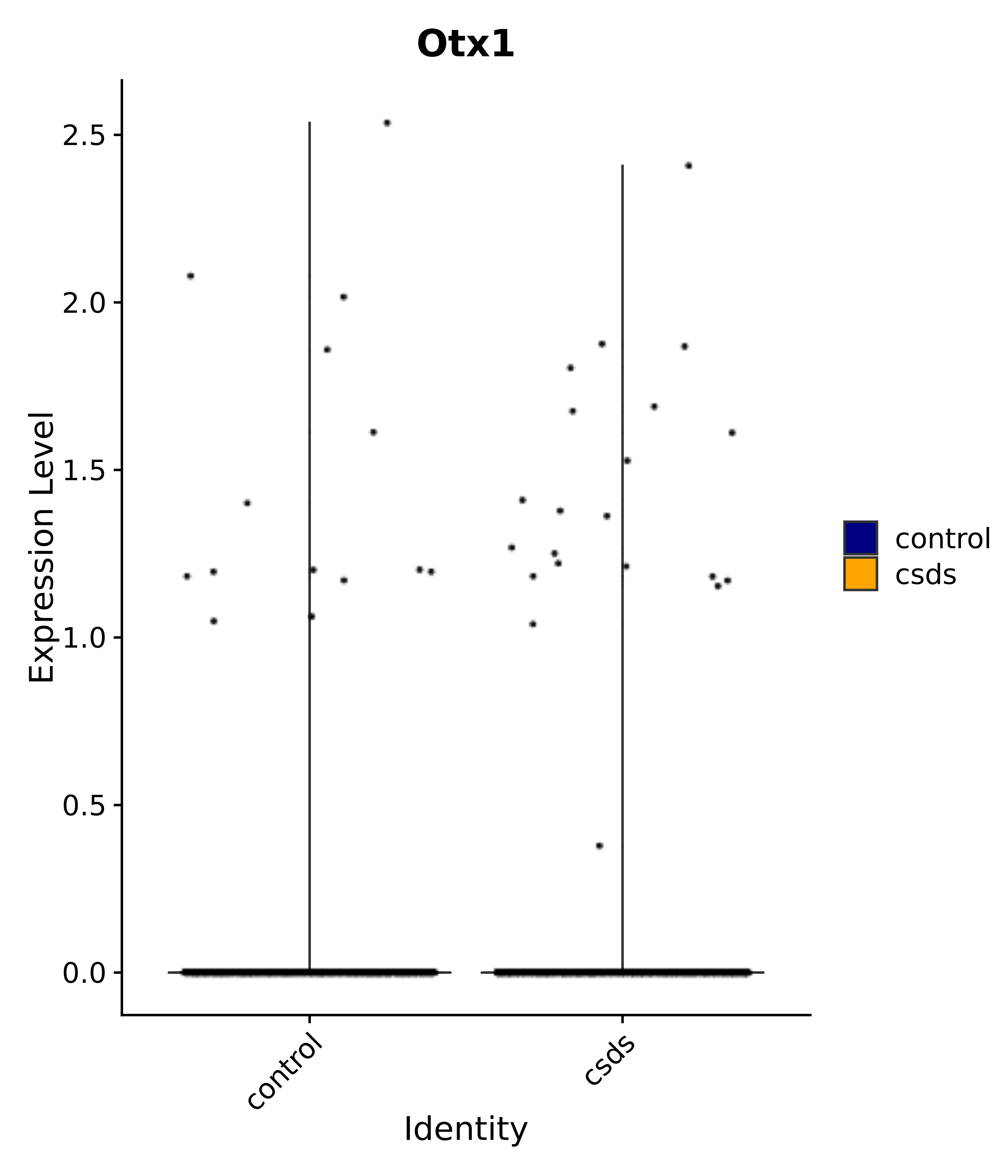
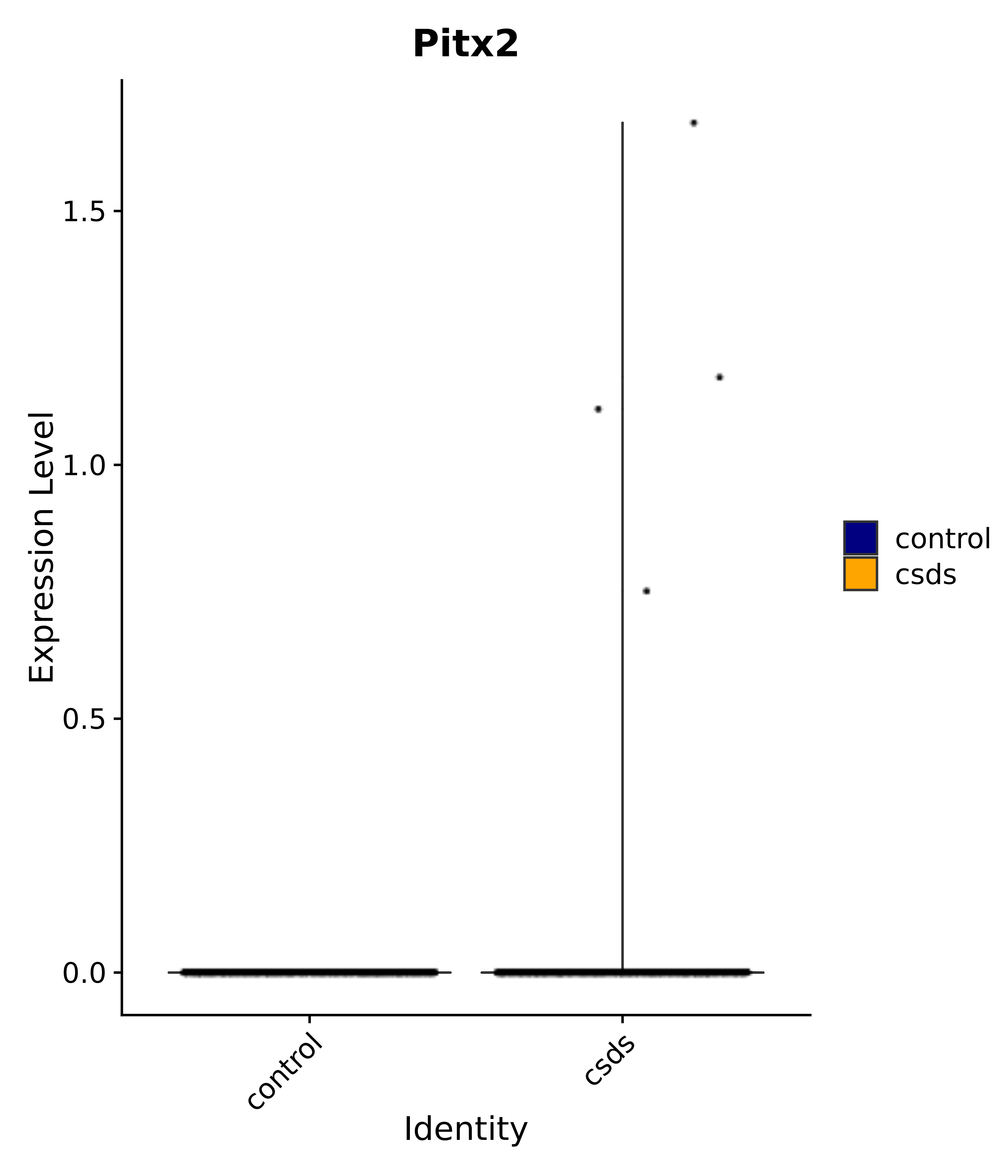
"Gfap", "Fgf1", "Fgfr3", "Hepacam", "Hif1", "Htra1", "Lxn", "Ndrg2", "Ntn1", "Nfia", "Slit2", "Aqp4", "S100a1", "S100a6", "S100b", "Slc1a2", "Slc1a3", "Slc38a1", "Vegfa", "Fos", "Fosb", "Jun", "Junb", "Jund", "Ier2", "Socs3", "Pde10a", "Fbln5", "Otp", "Nr4a1", "Six6", "Emx2", "Myt1l", "Adcyap1r1", "Ghr", "Ntrk2", "Npy1r", "Gria1", "Gria2", "Grin2b", "Grin3a", "Grm3", "Grm7", "Gabrb1", "Gabbr1", "Gabbr2", "Rfx3", "Nr5a1", "Nkx2-1", "Otx2", "Bclaf1", "Foxo3", "Dlx1", "Lrrc6", "Peg3", "Elavl2", "Isl1", "Zfp36", "Otx1", "Pitx2",
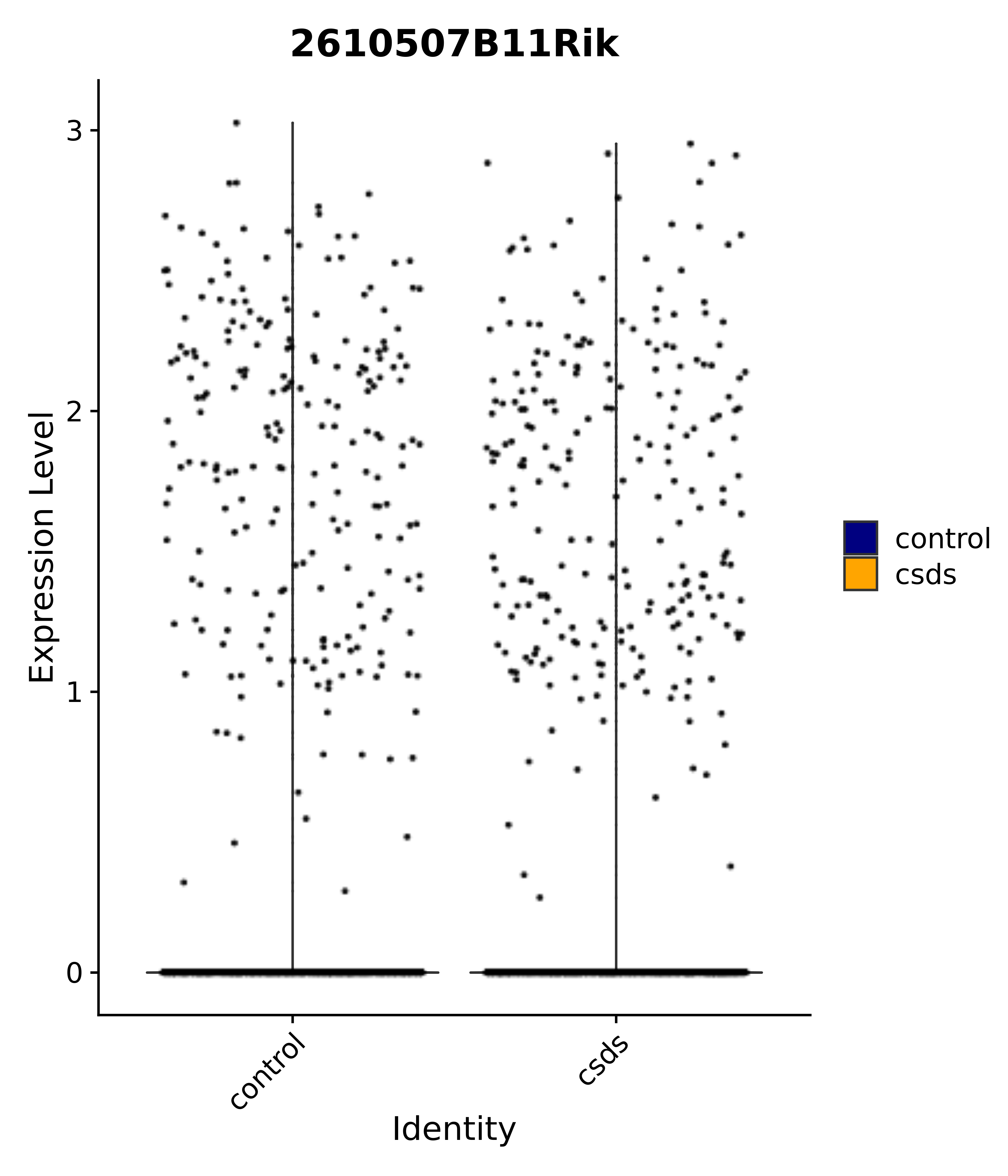
"2610507B11Rik",
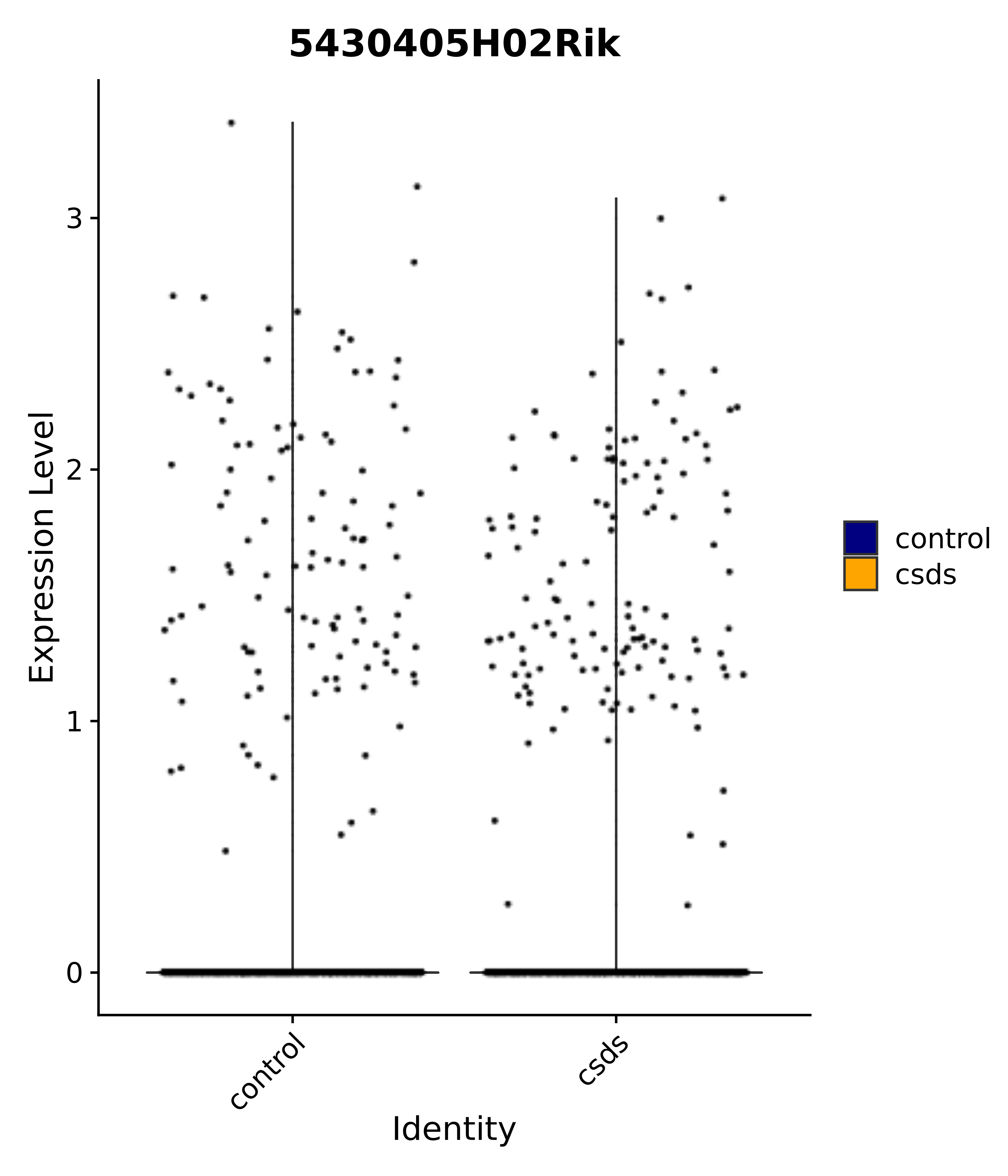
"5430405H02Rik",
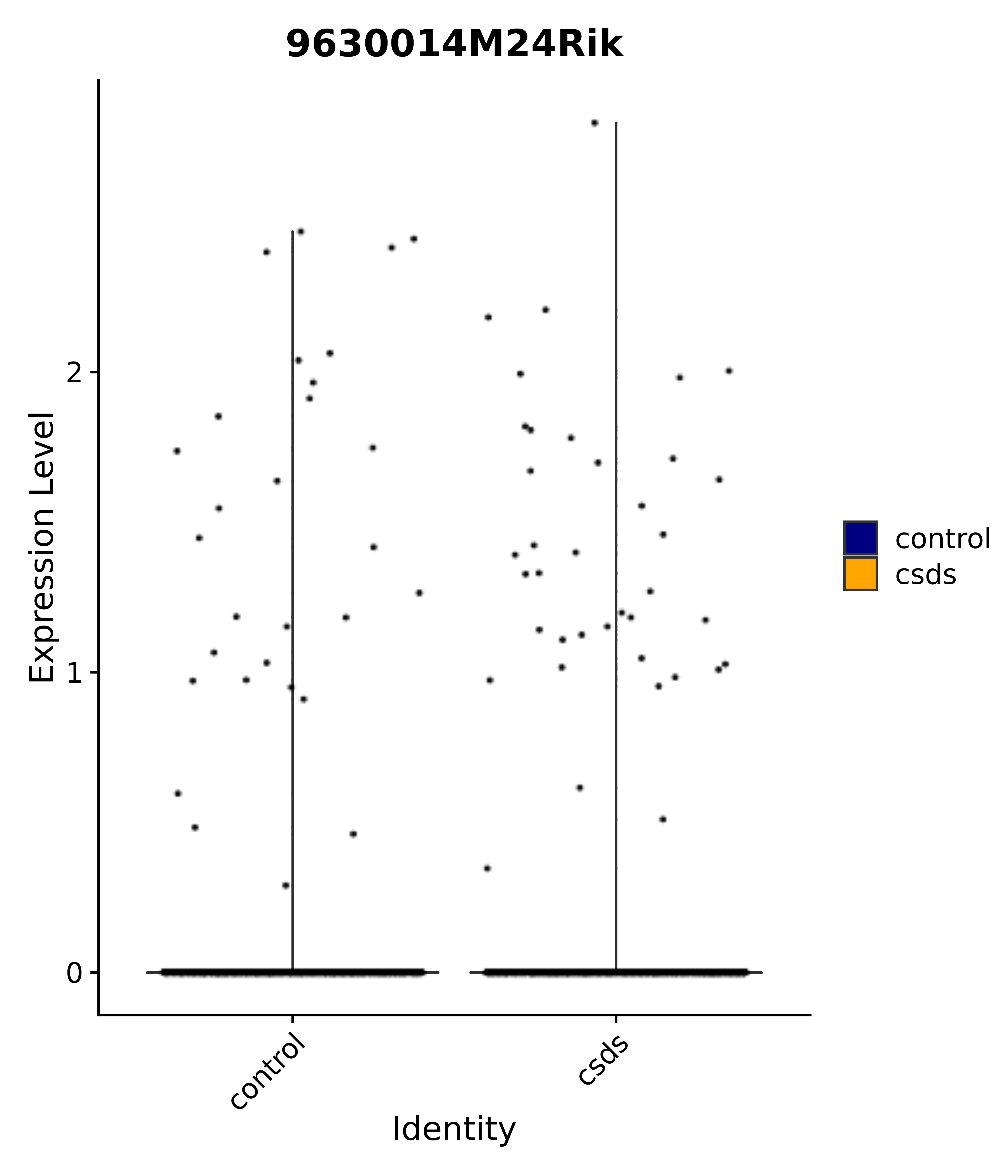
"9630014M24Rik",
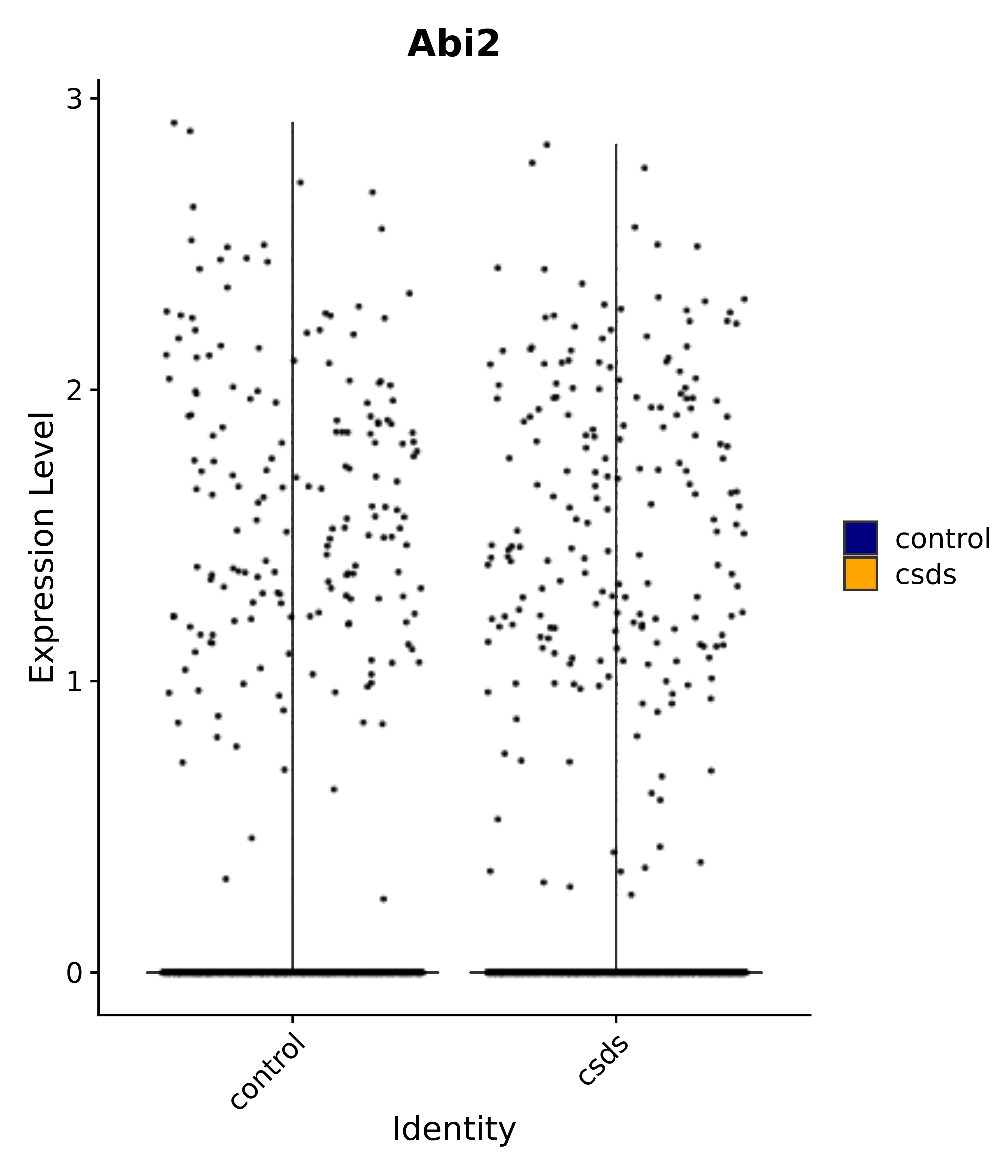
"Abi2",
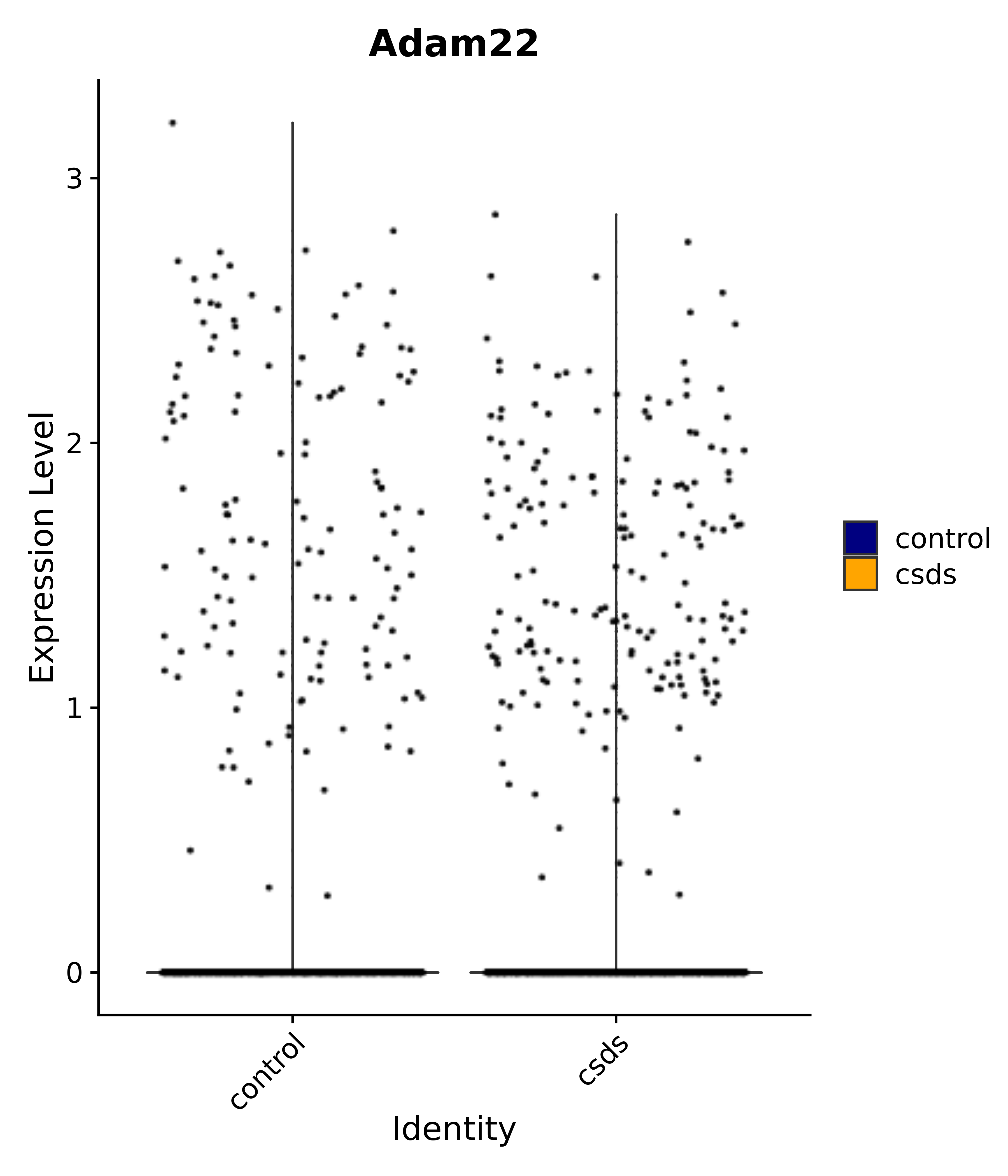
"Adam22",
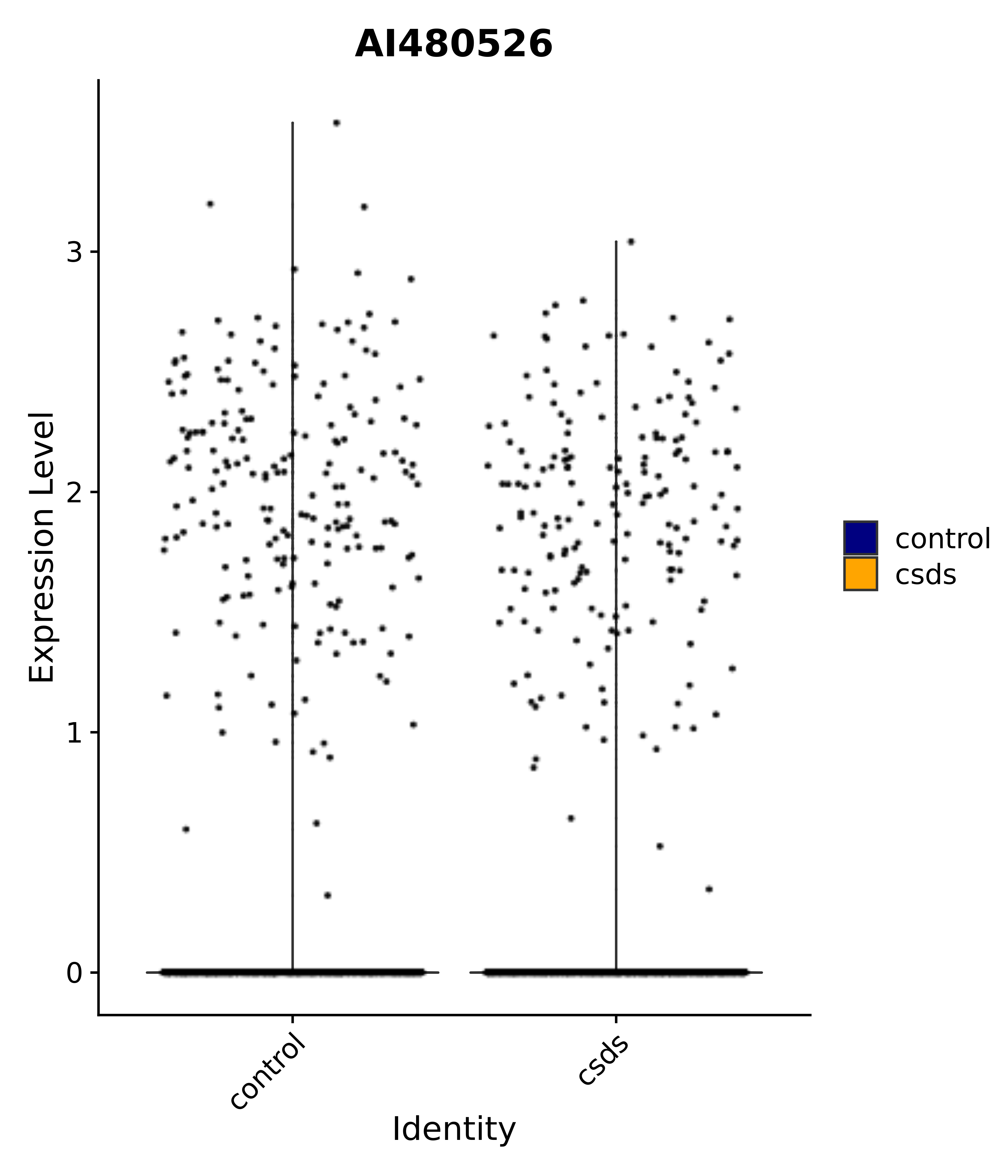
"AI480526",
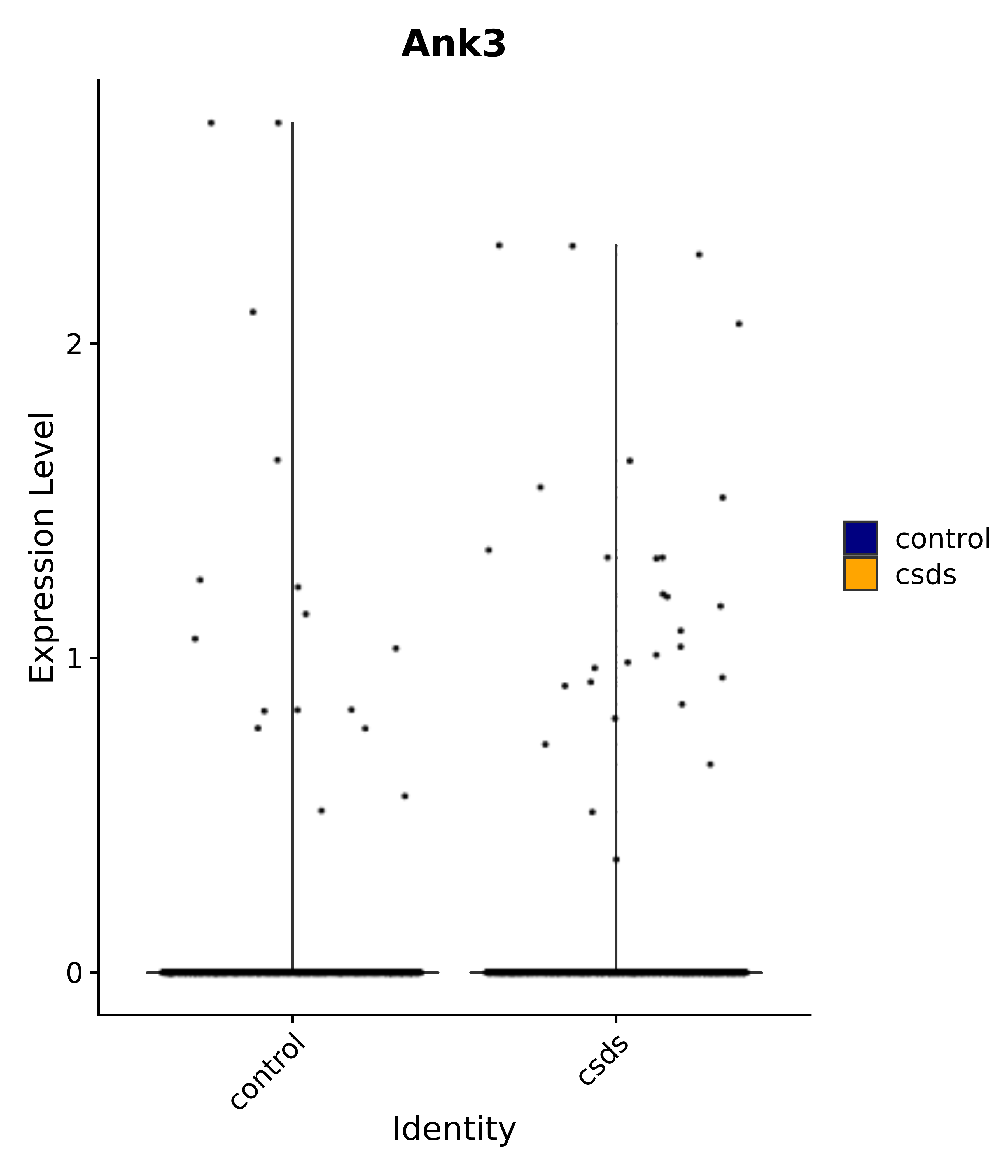
"Ank3",
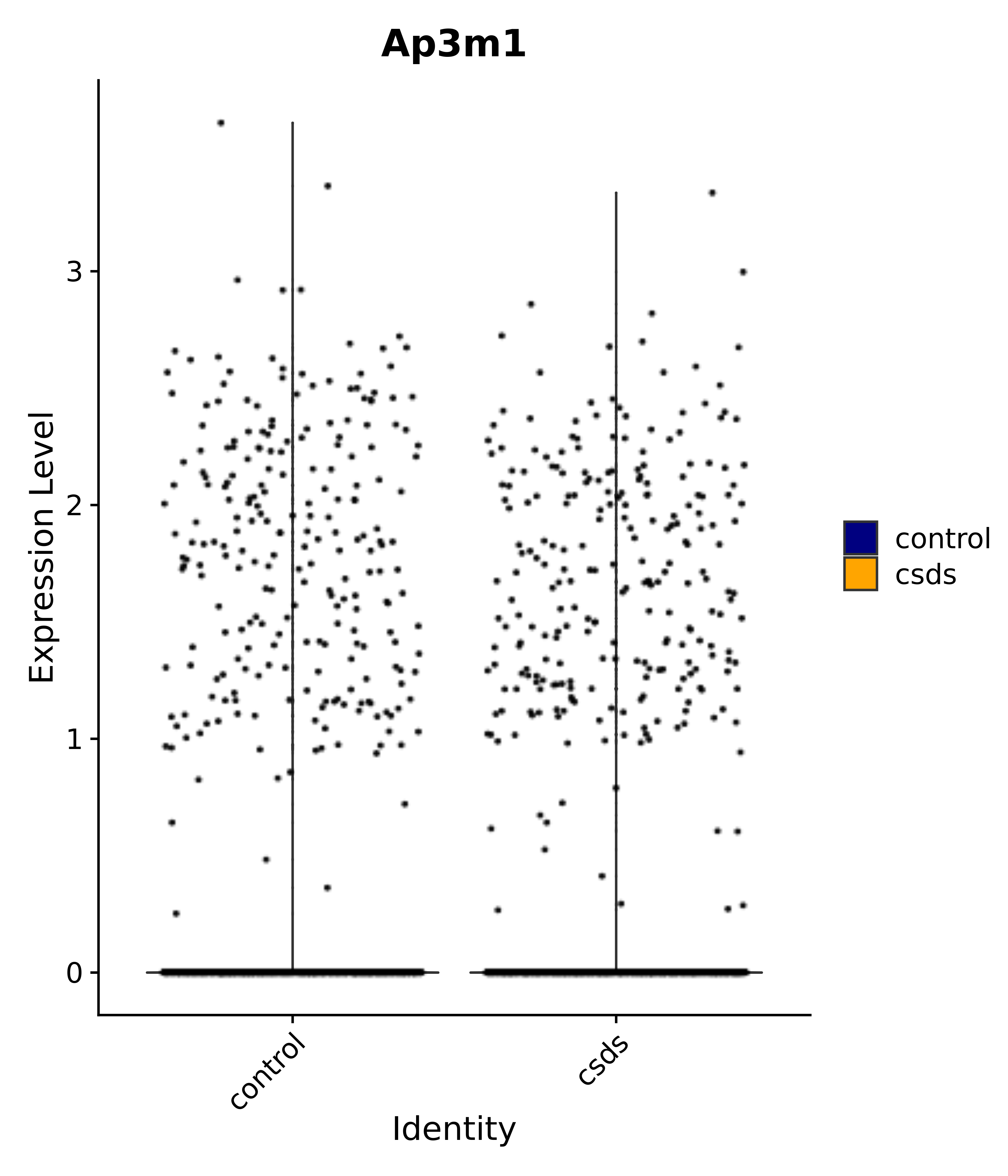
"Ap3m1",
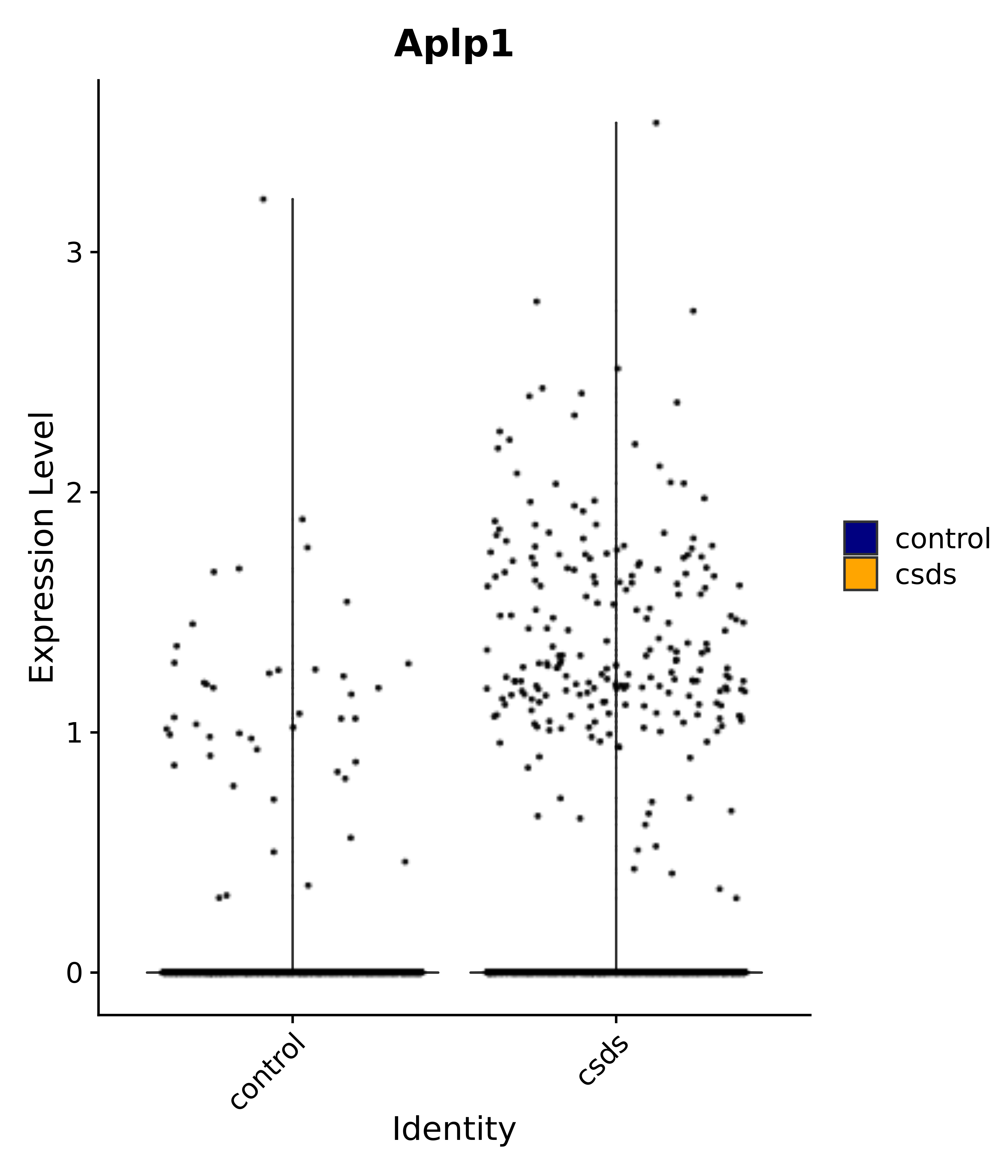
"Aplp1",
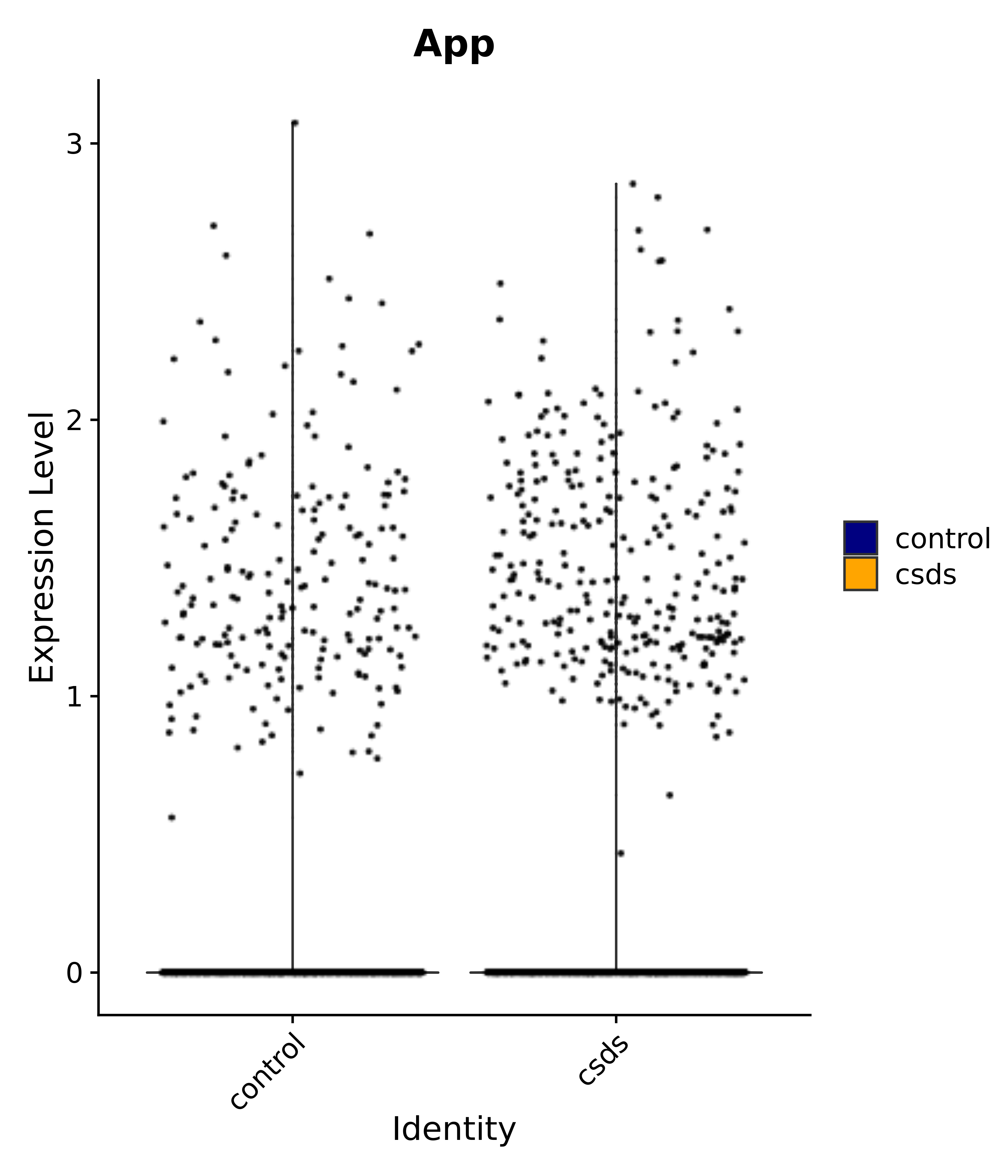
"App",
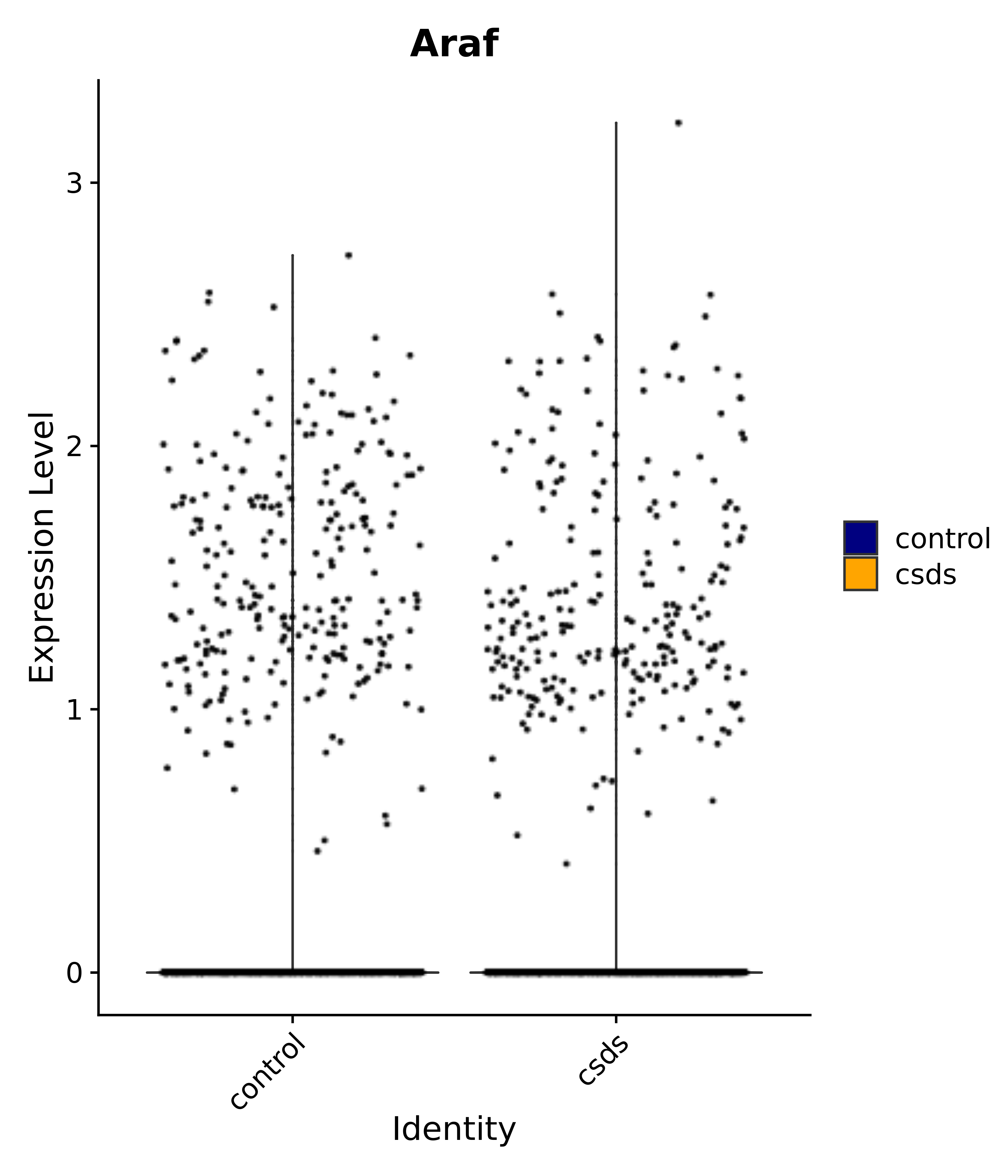
"Araf",
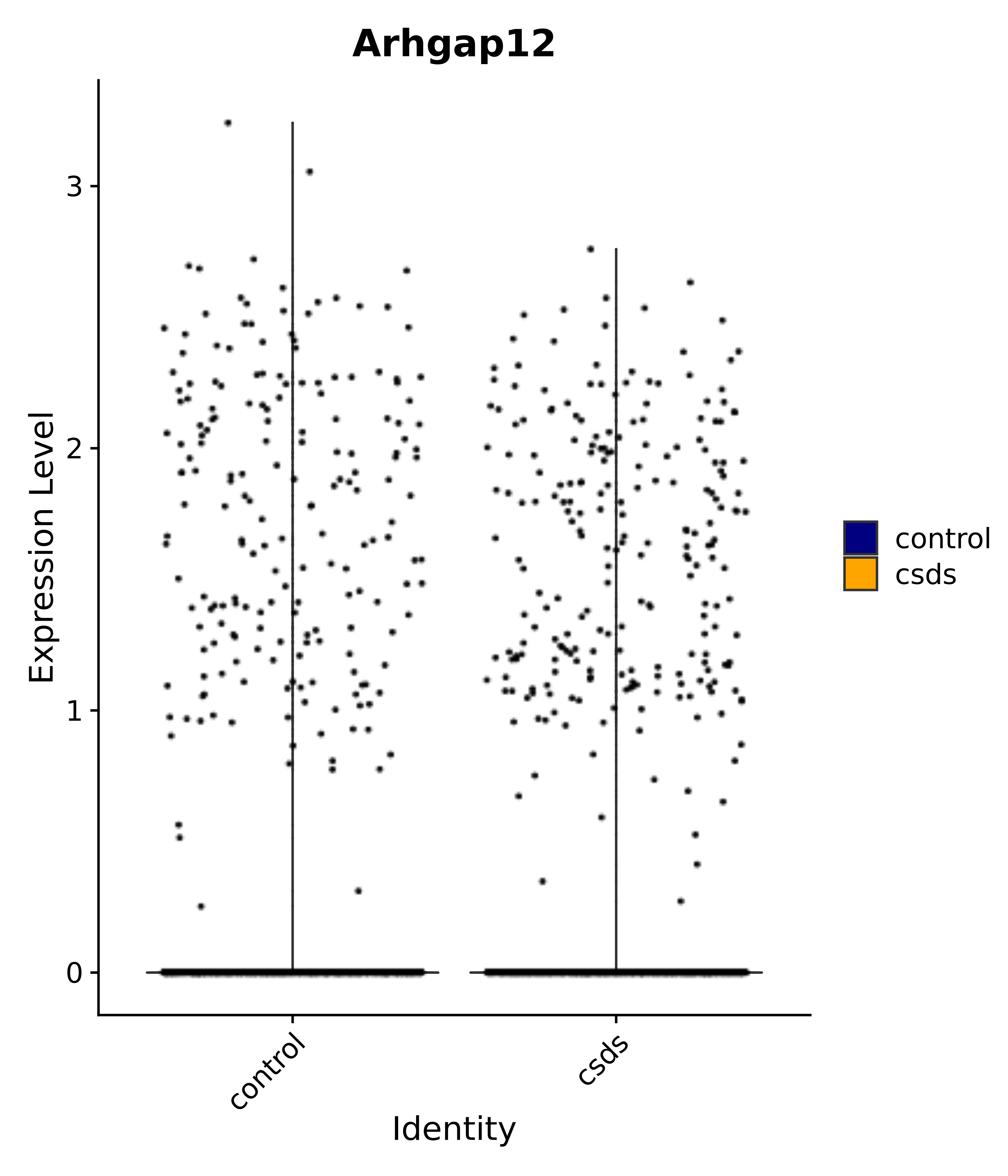
"Arhgap12",
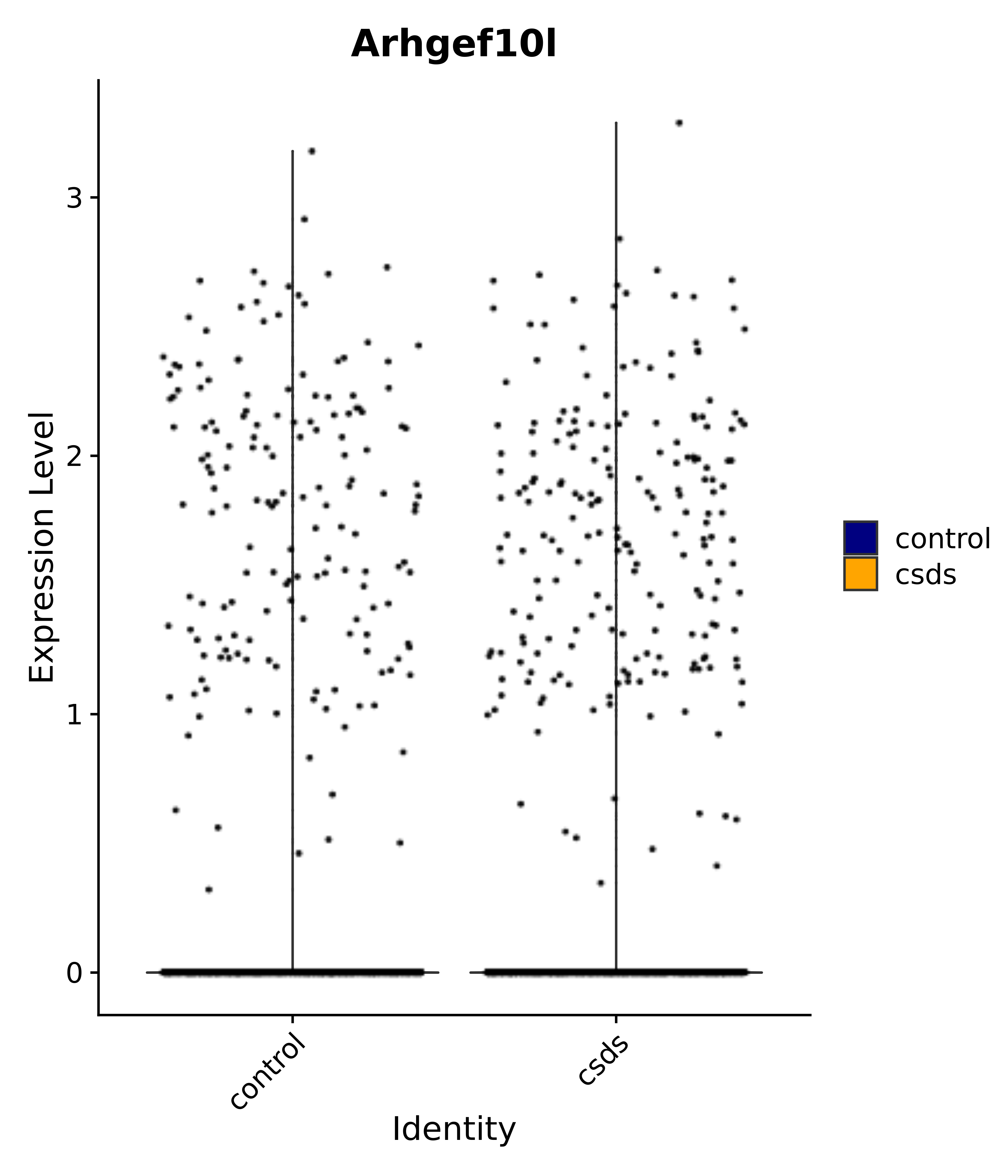
"Arhgef10l",
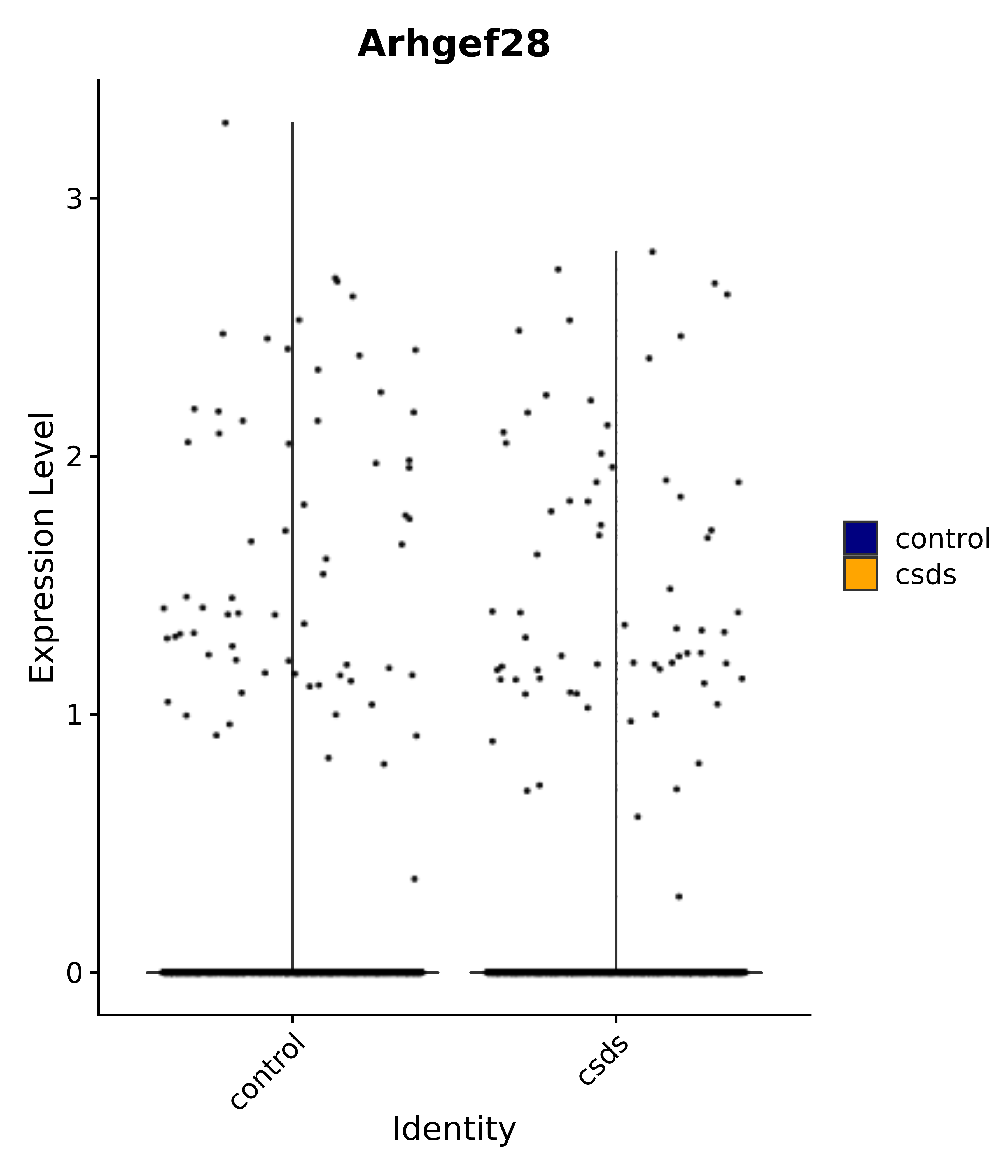
"Arhgef28",
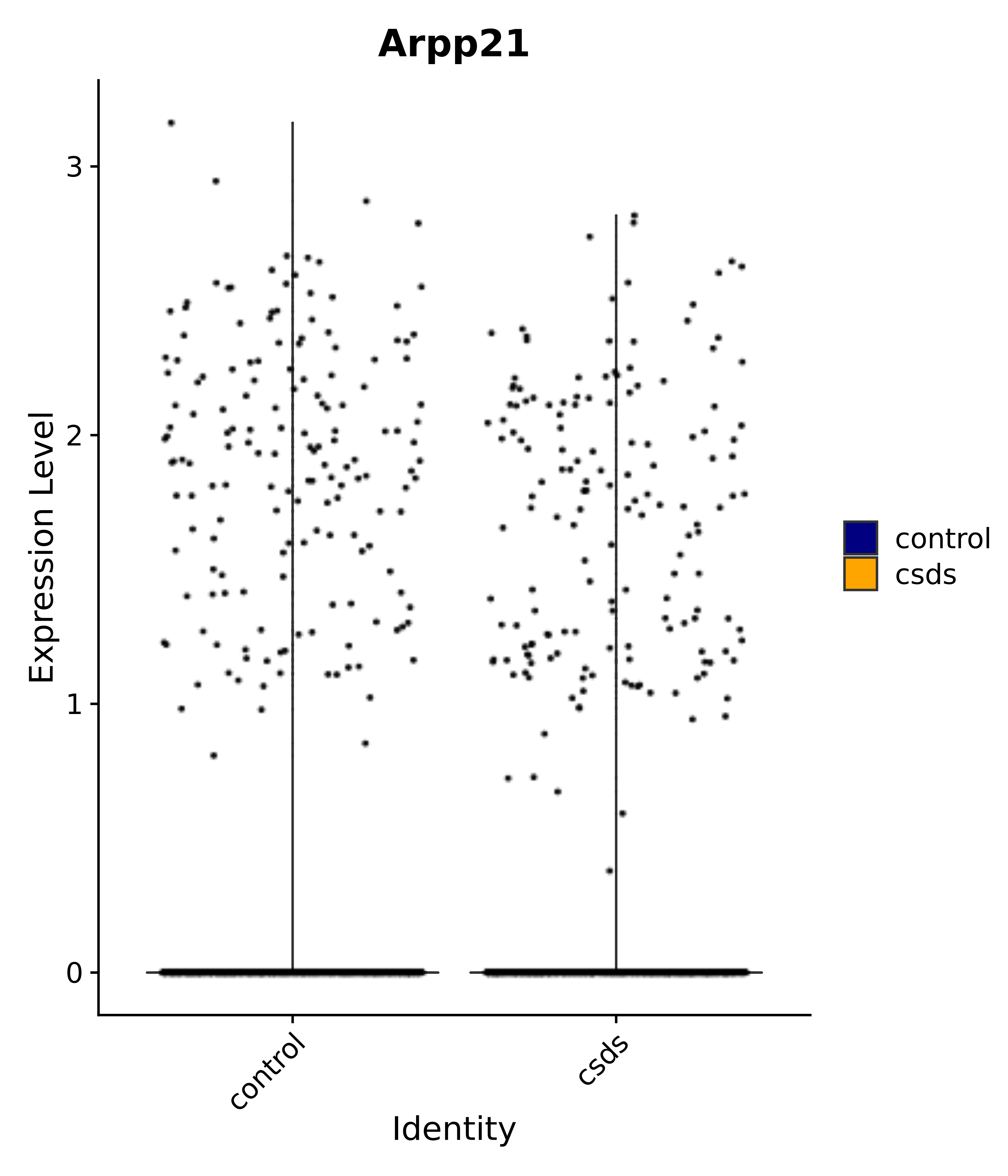
"Arpp21",
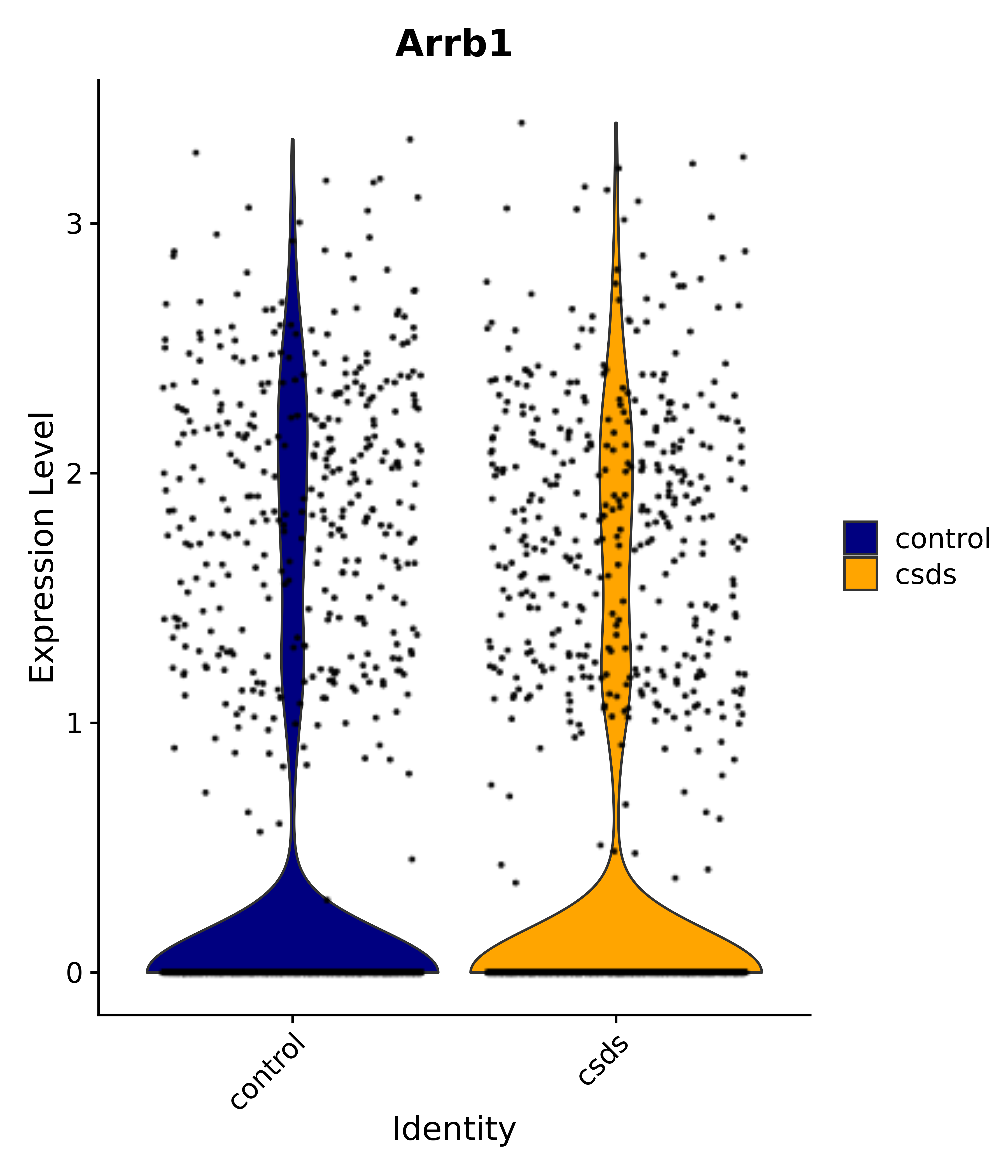
"Arrb1",
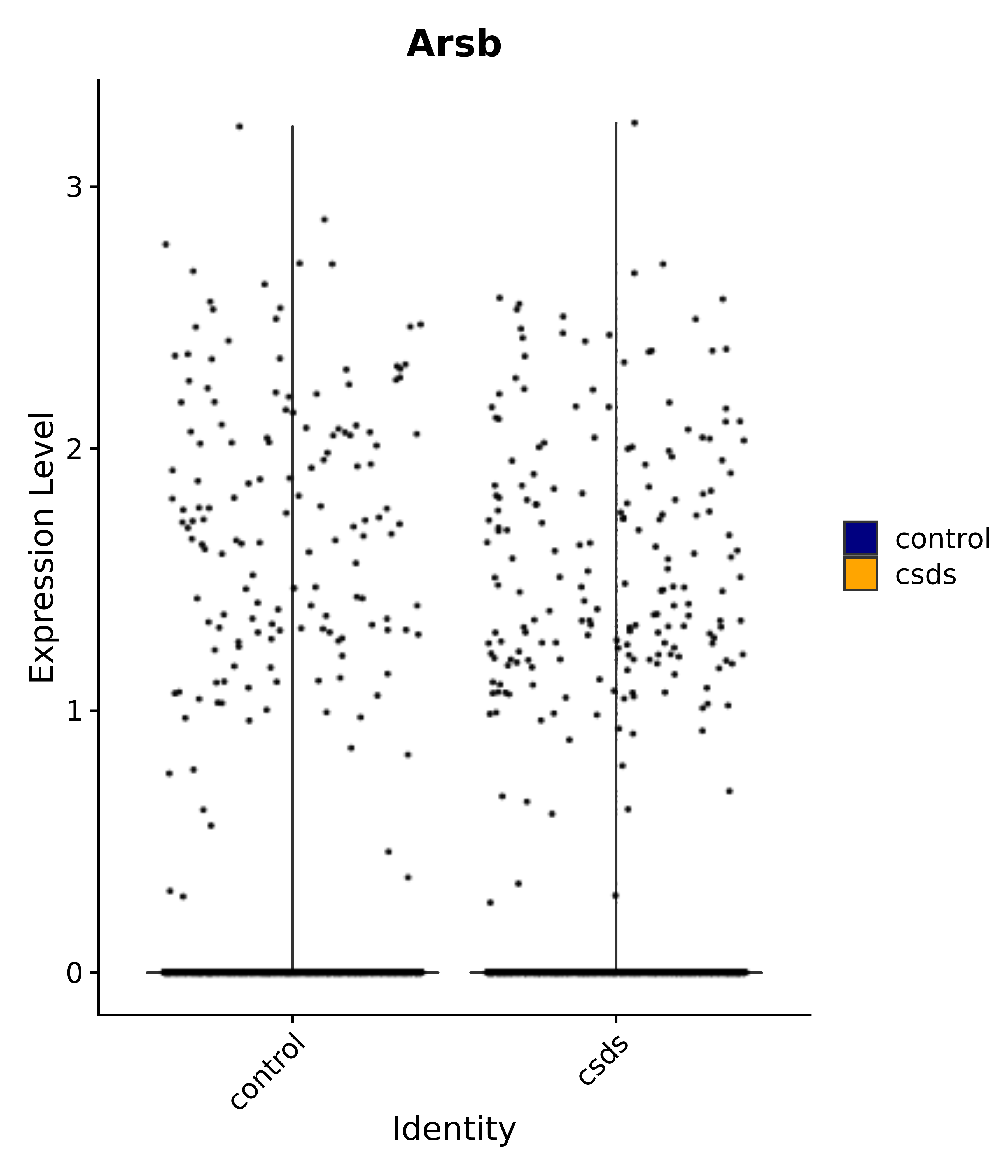
"Arsb",
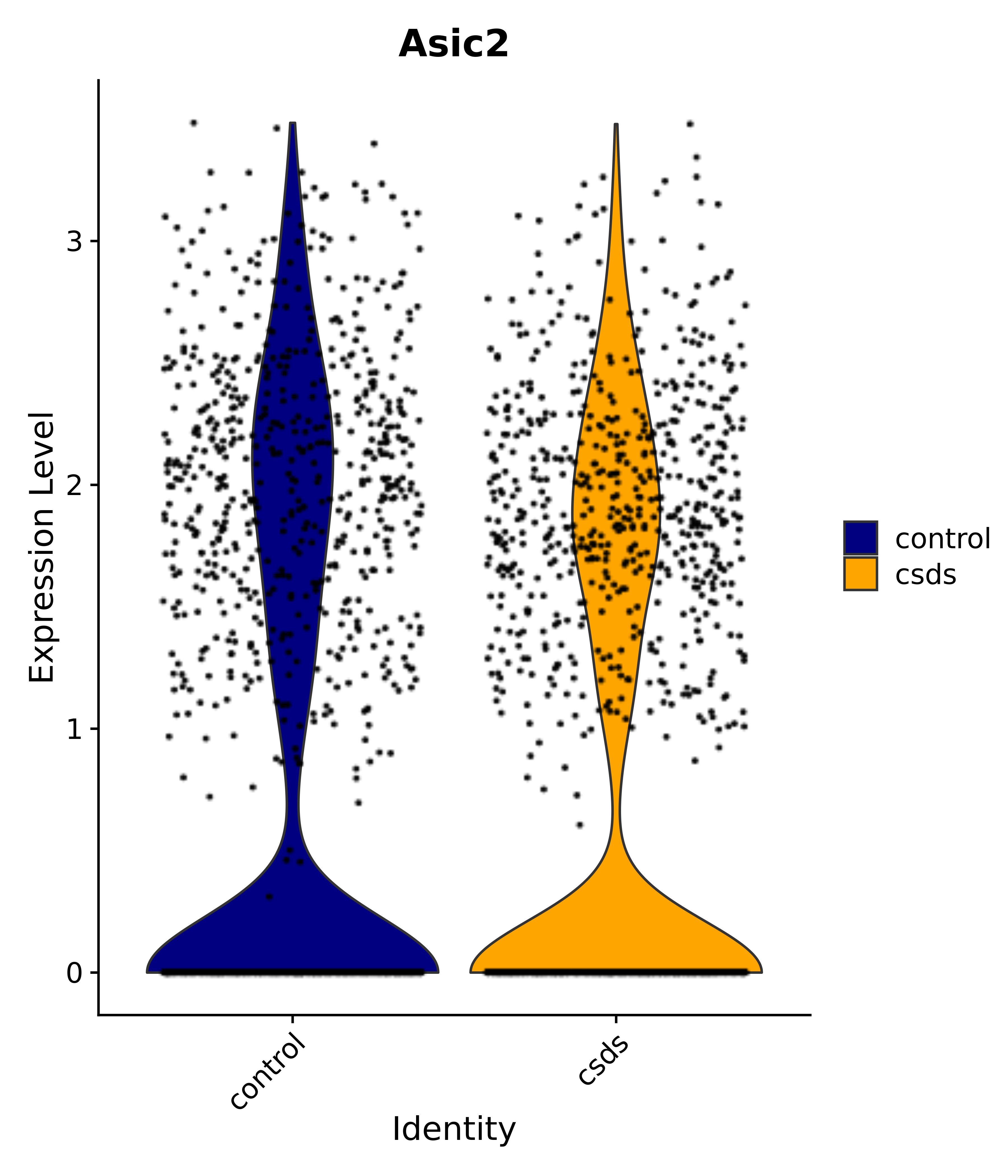
"Asic2",
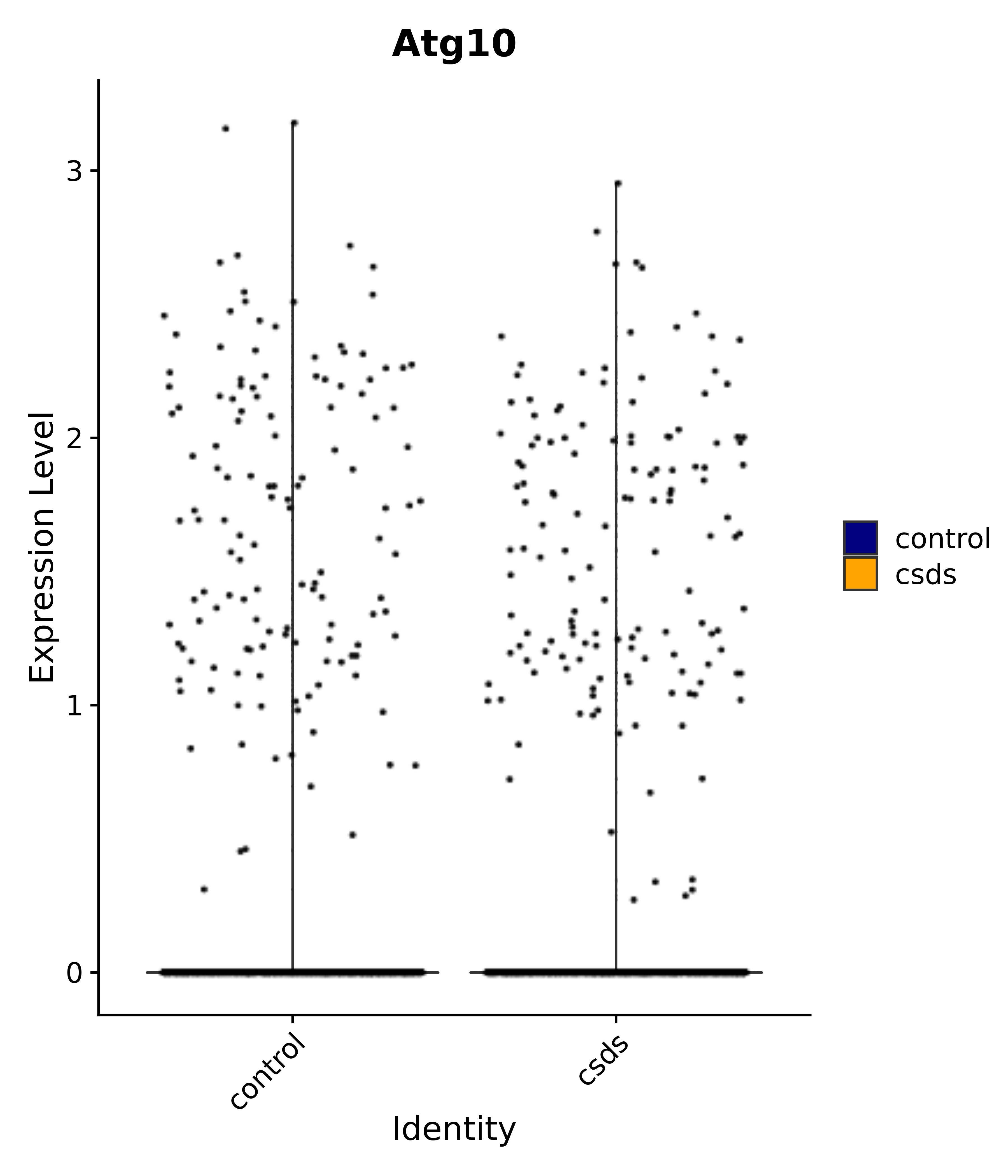
"Atg10",
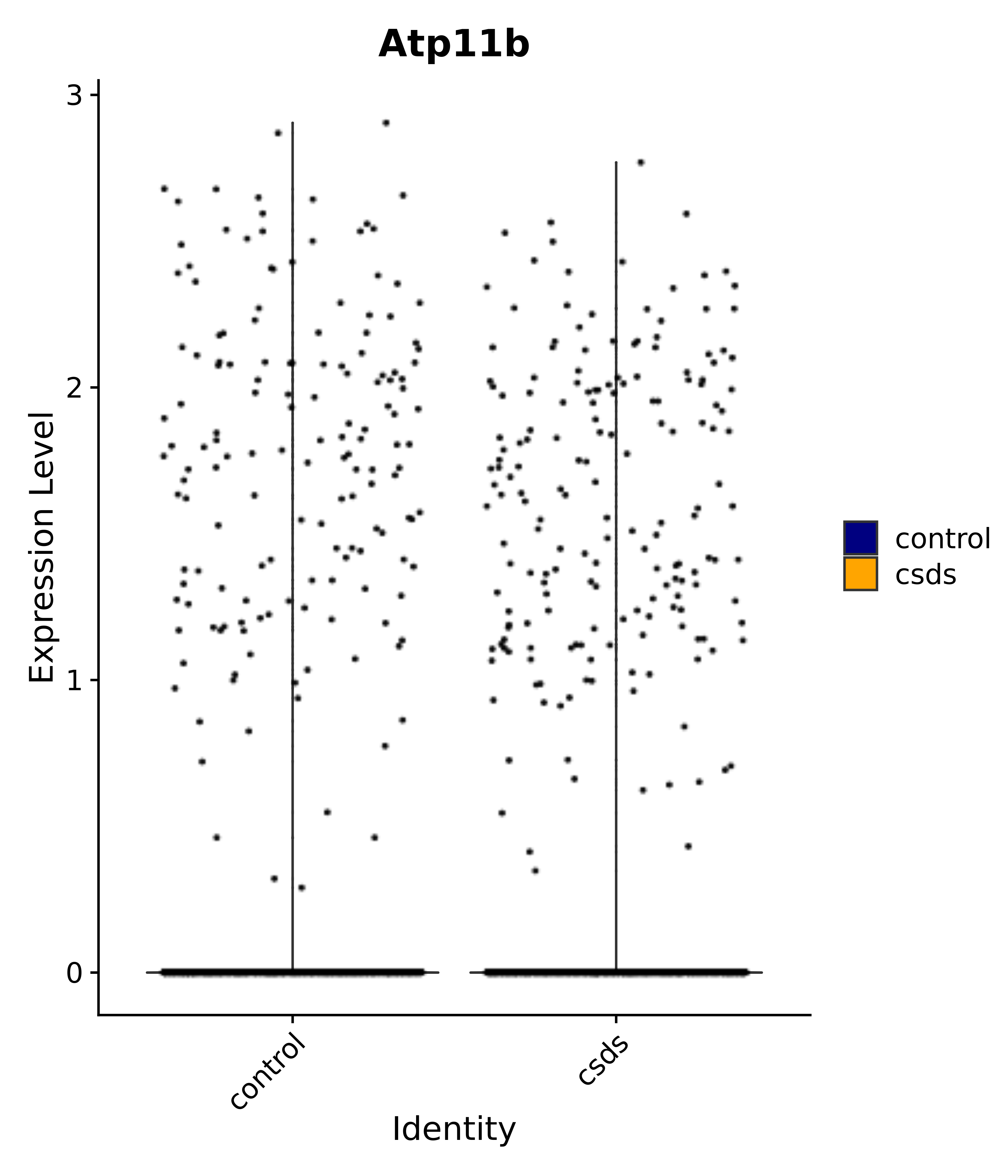
"Atp11b",
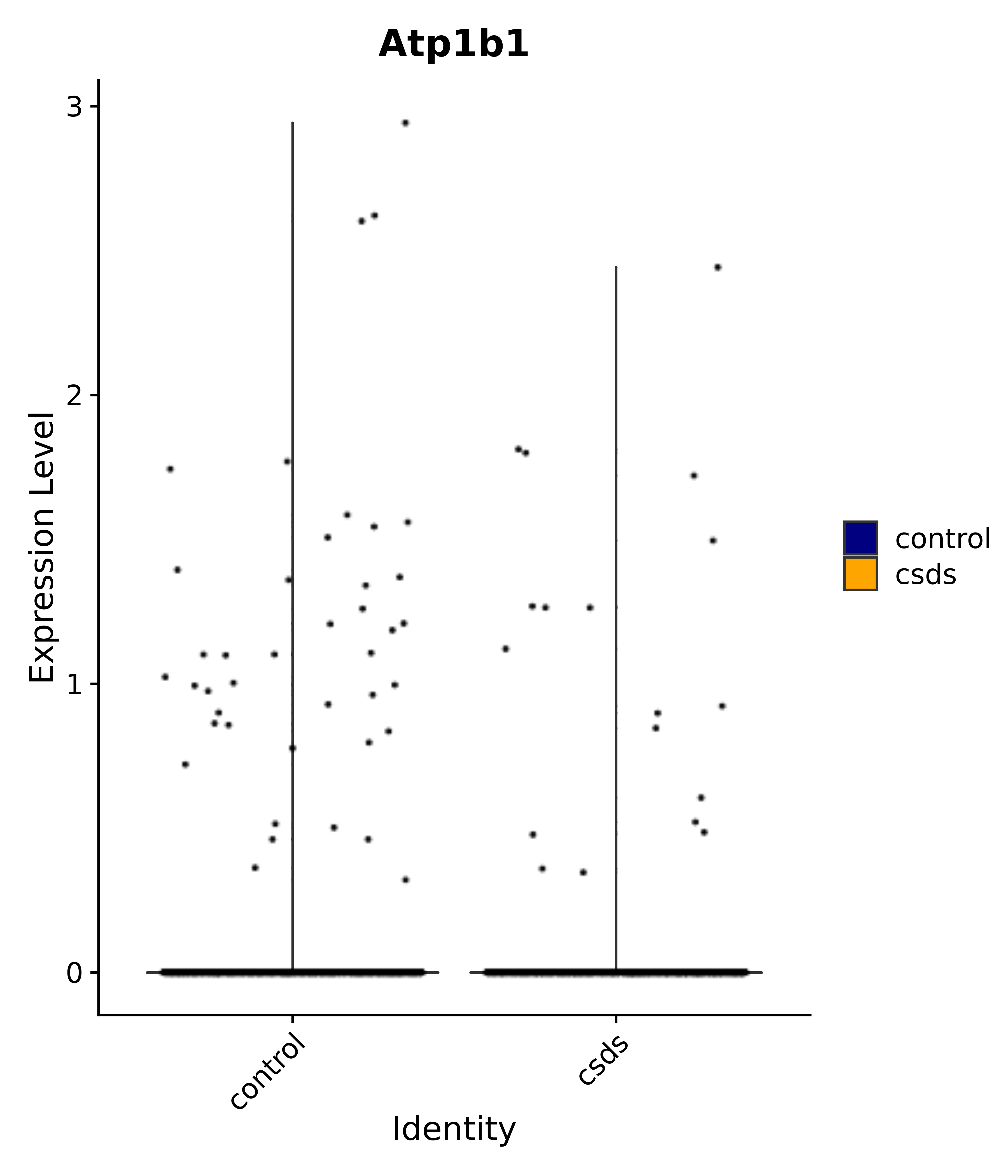
"Atp1b1",
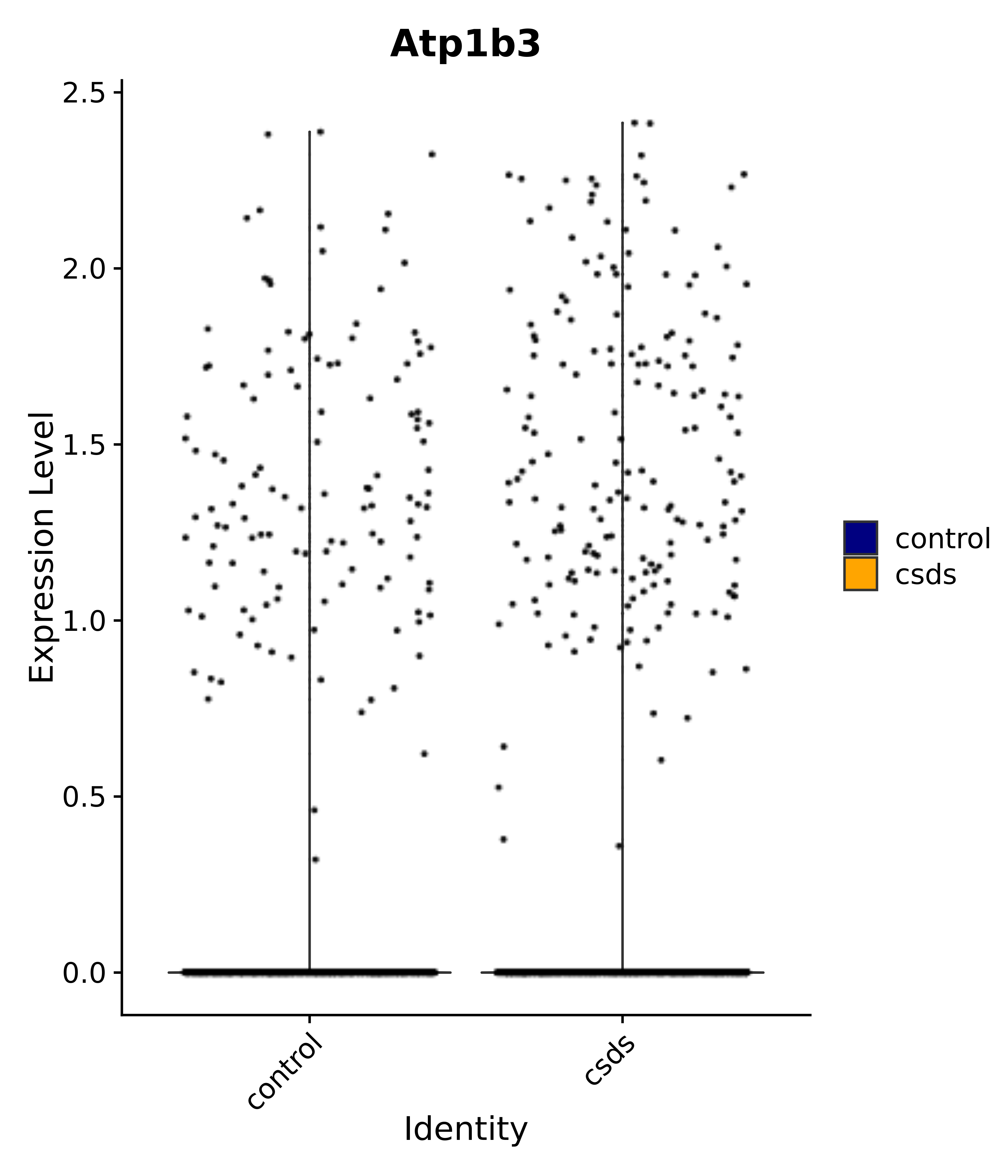
"Atp1b3",
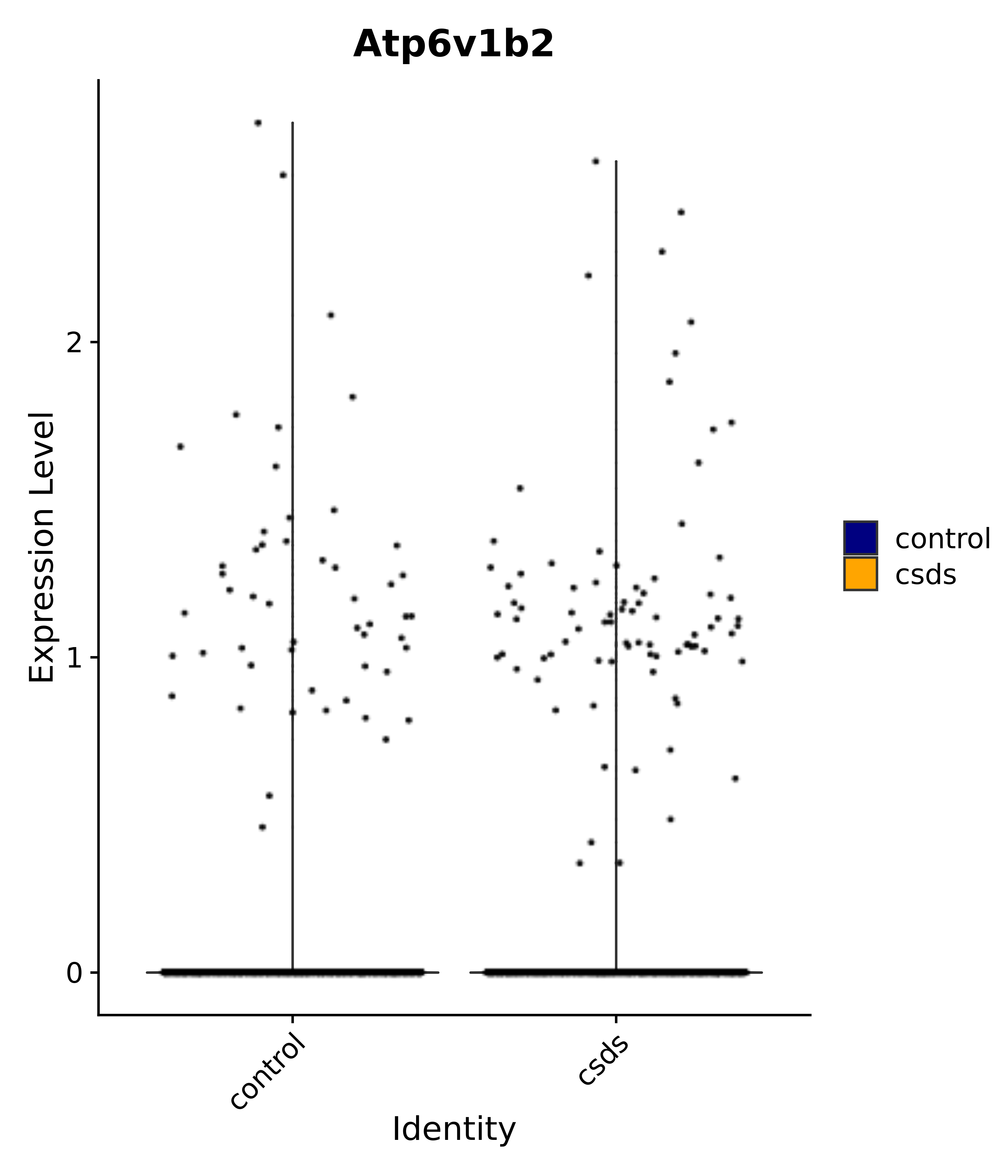
"Atp6v1b2",
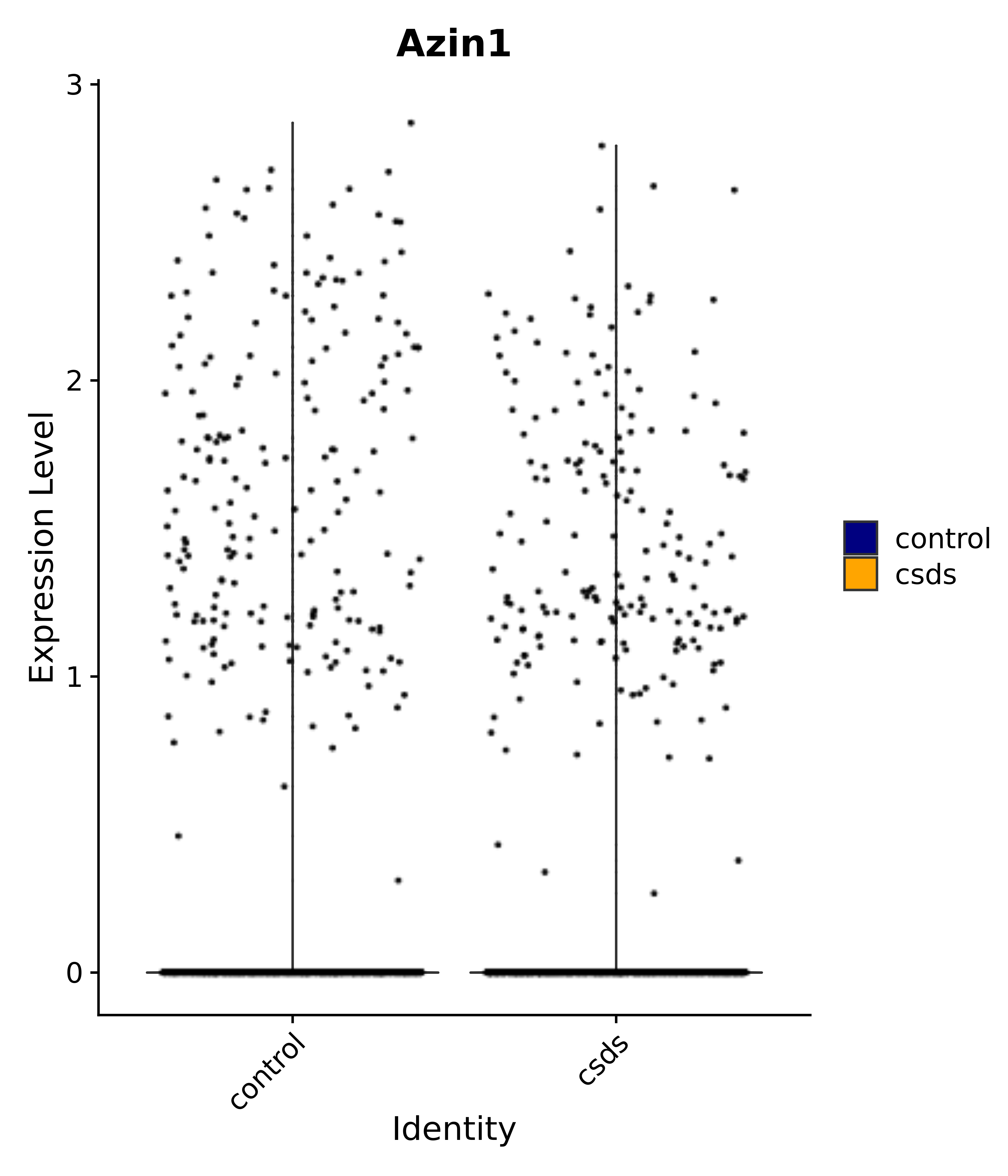
"Azin1",
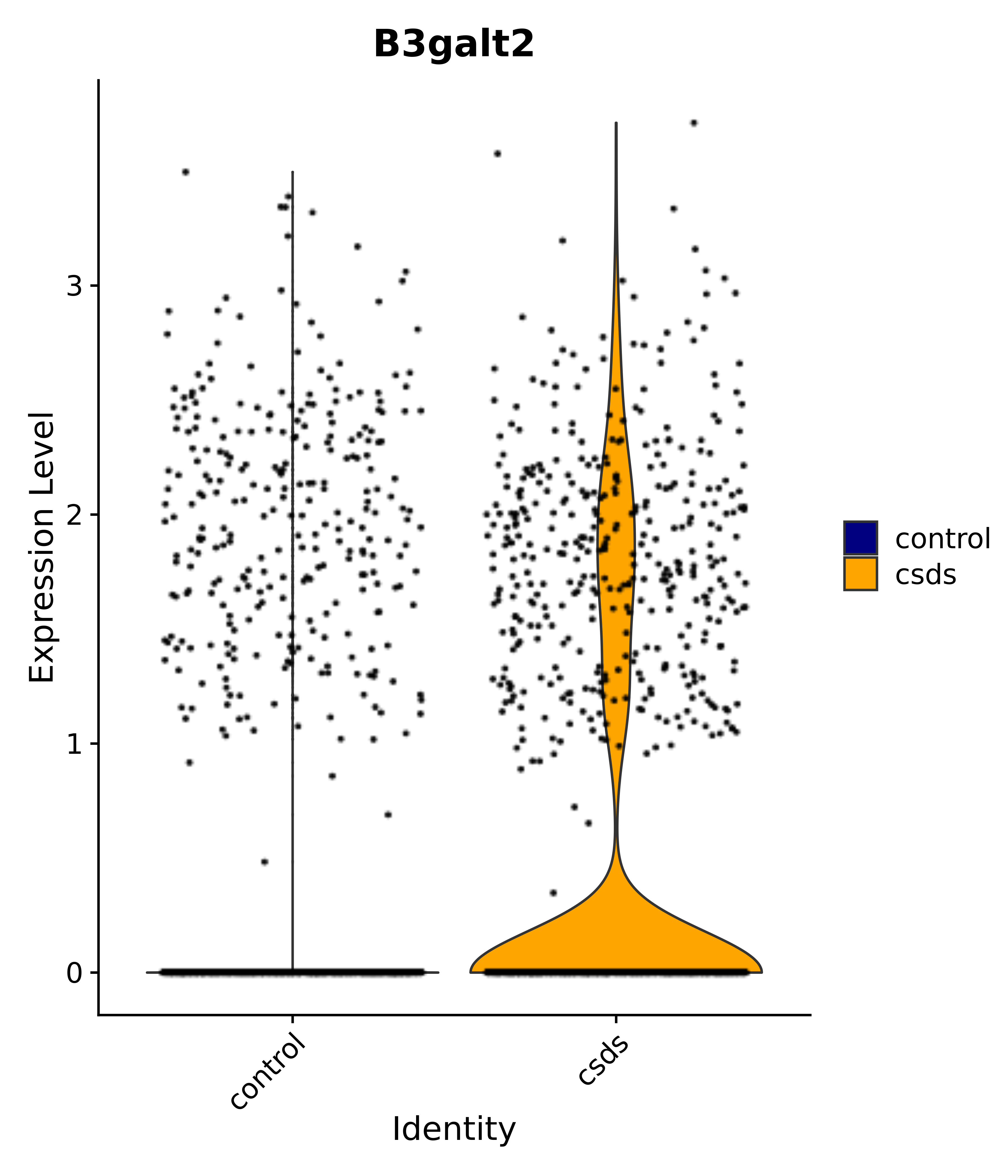
"B3galt2",
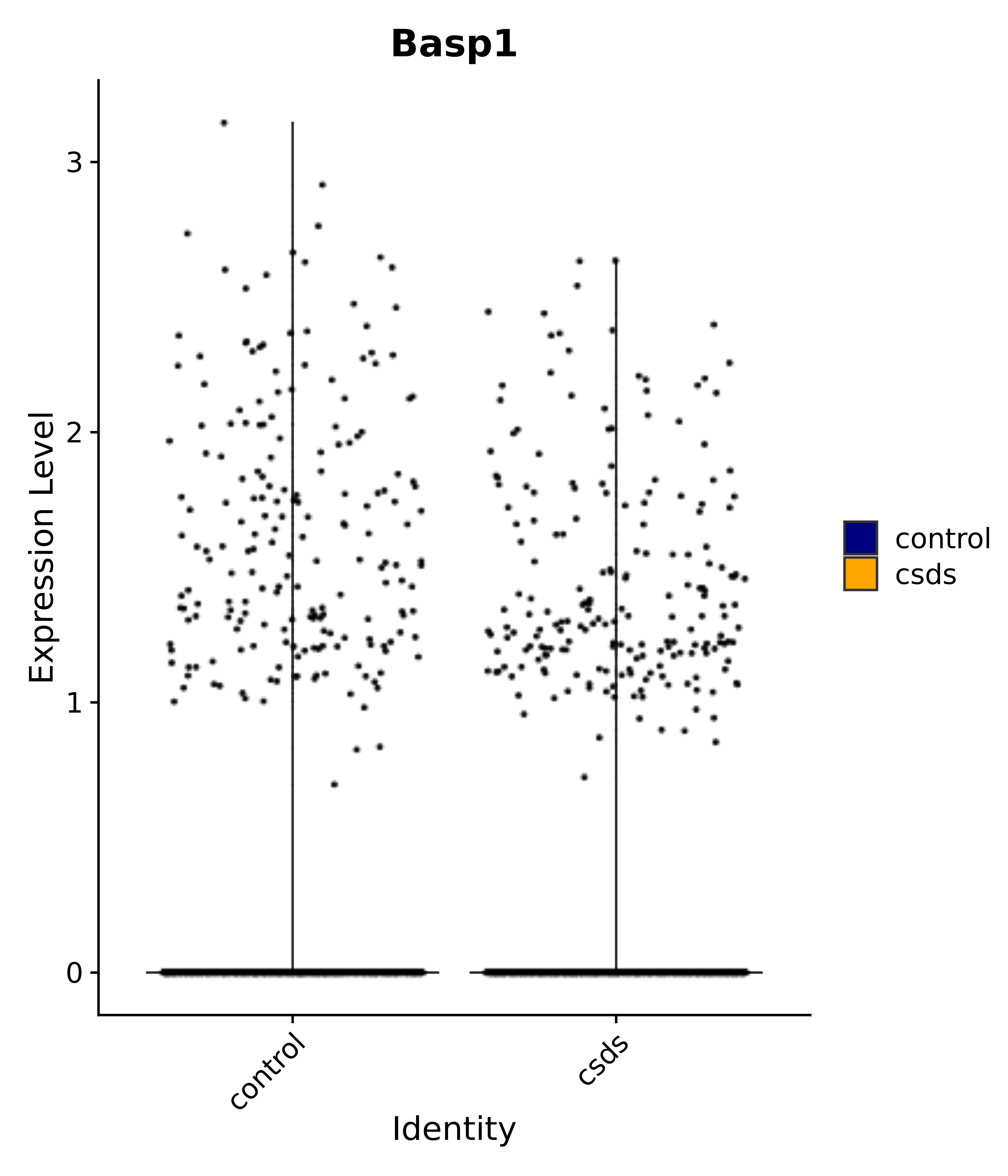
"Basp1",
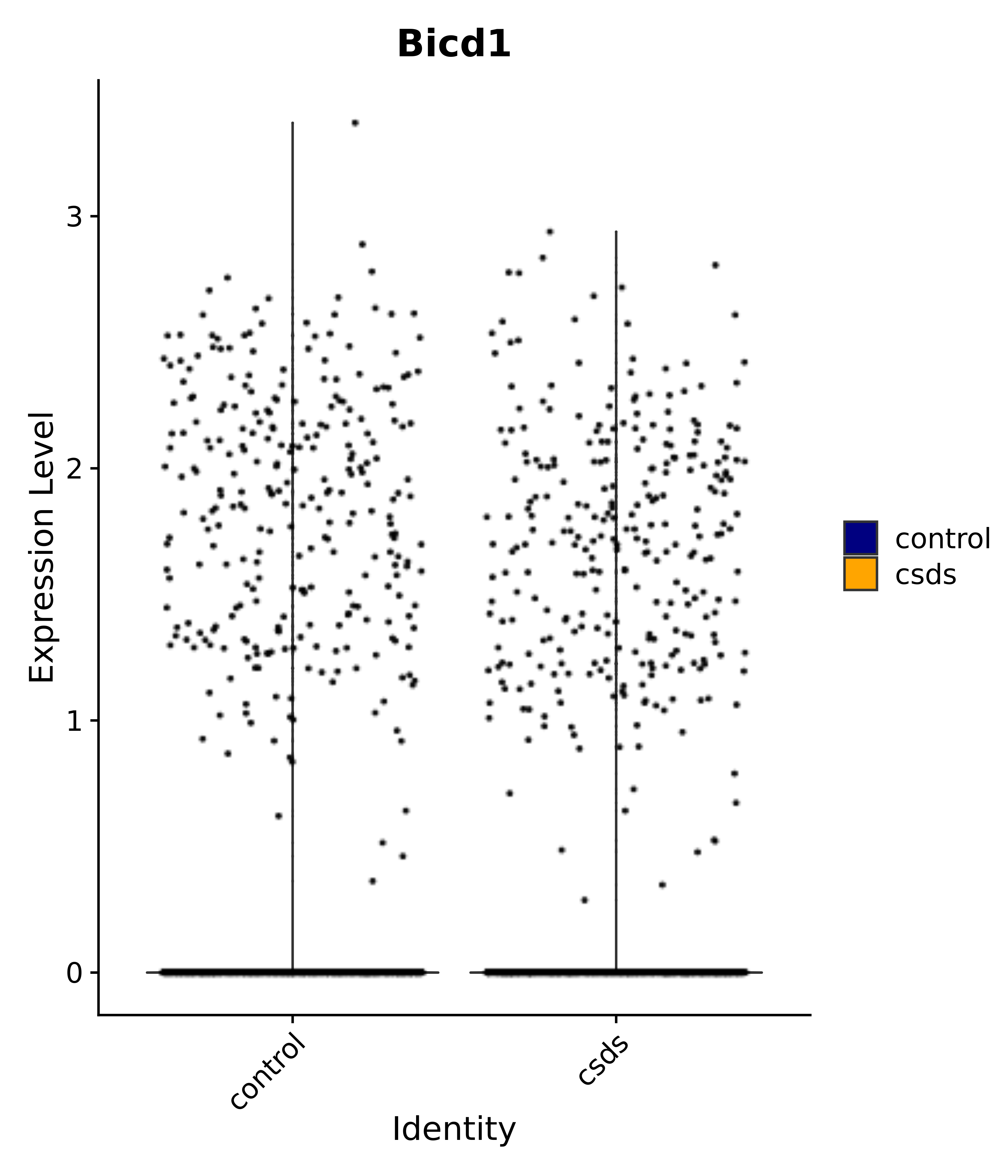
"Bicd1",
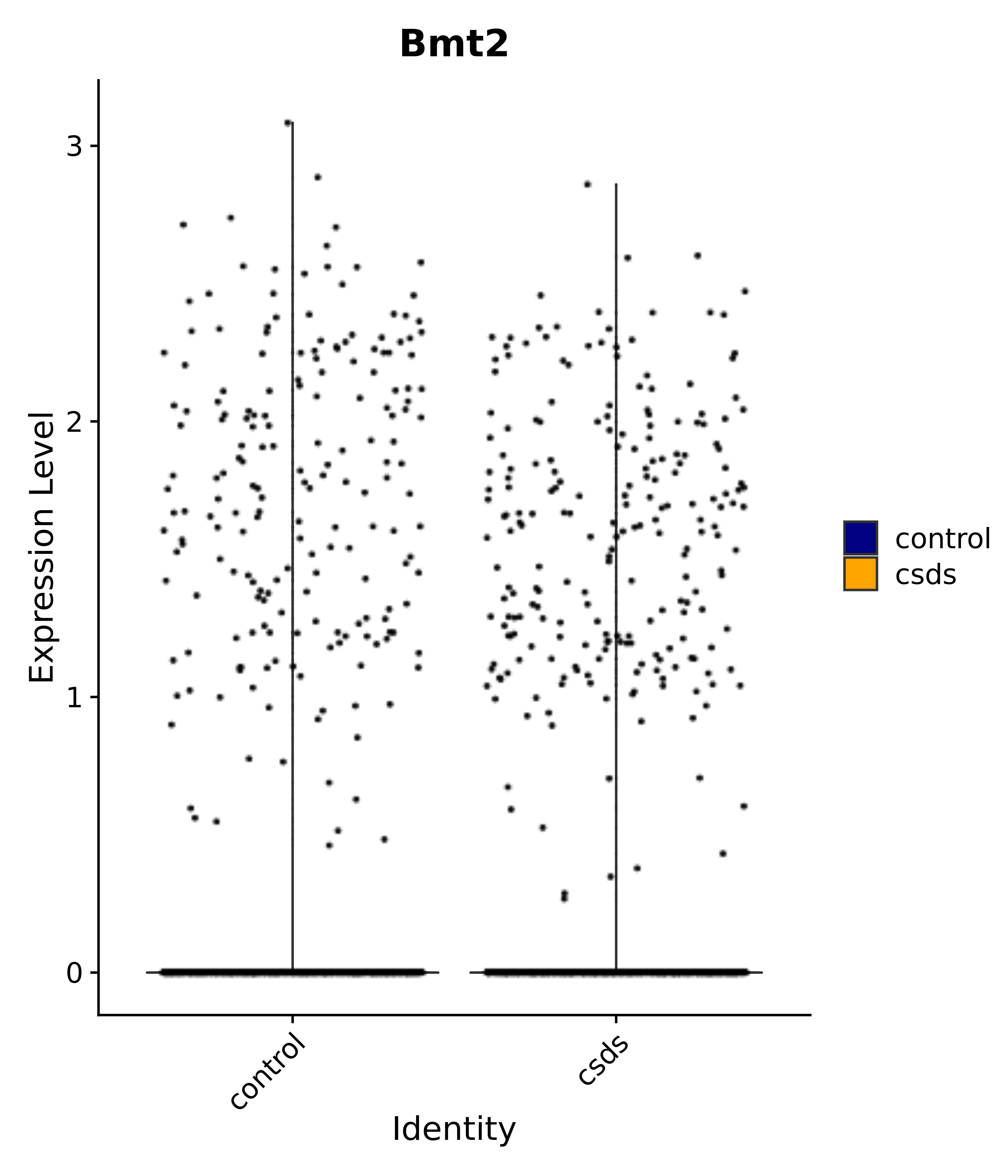
"Bmt2",
"Btg2",
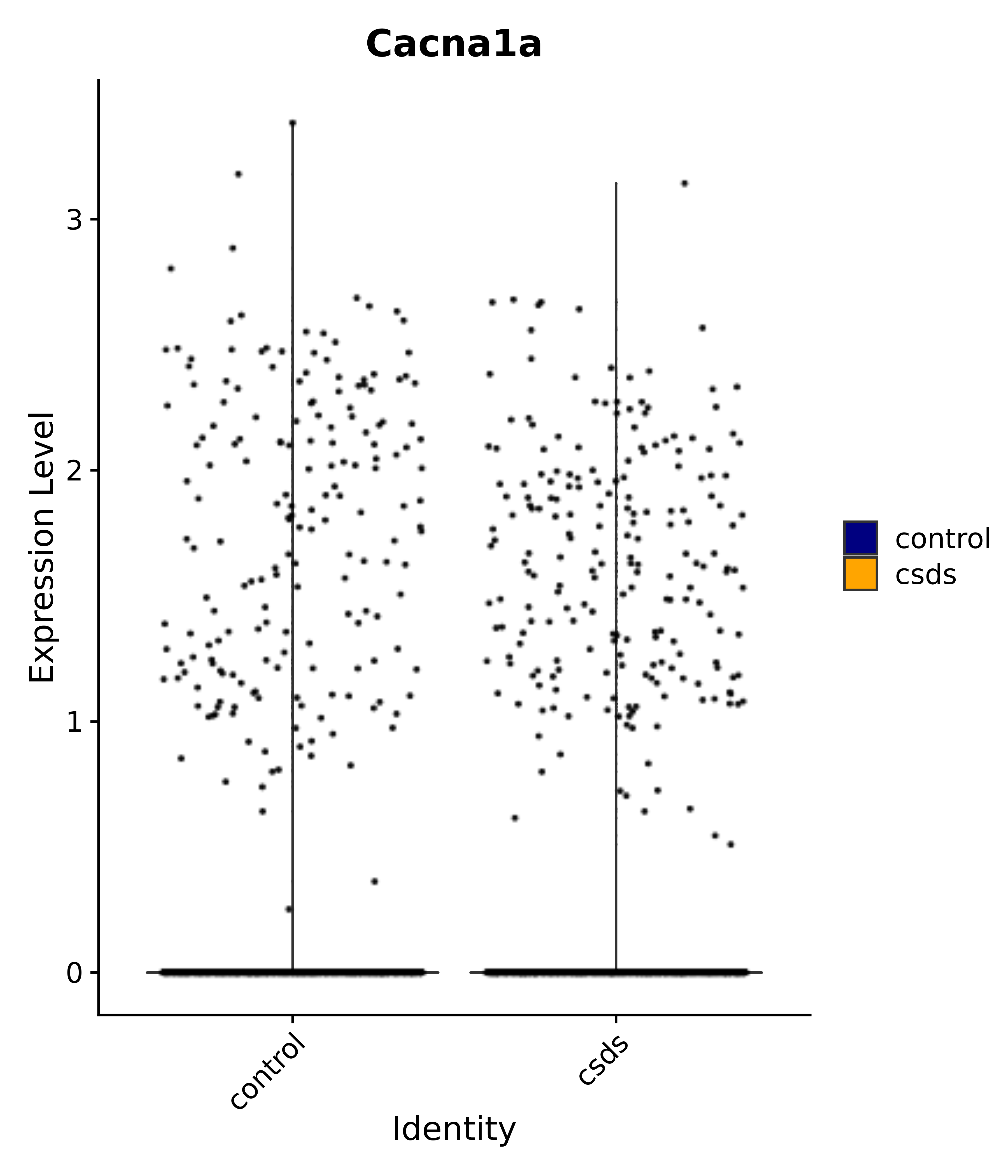
"Cacna1a",
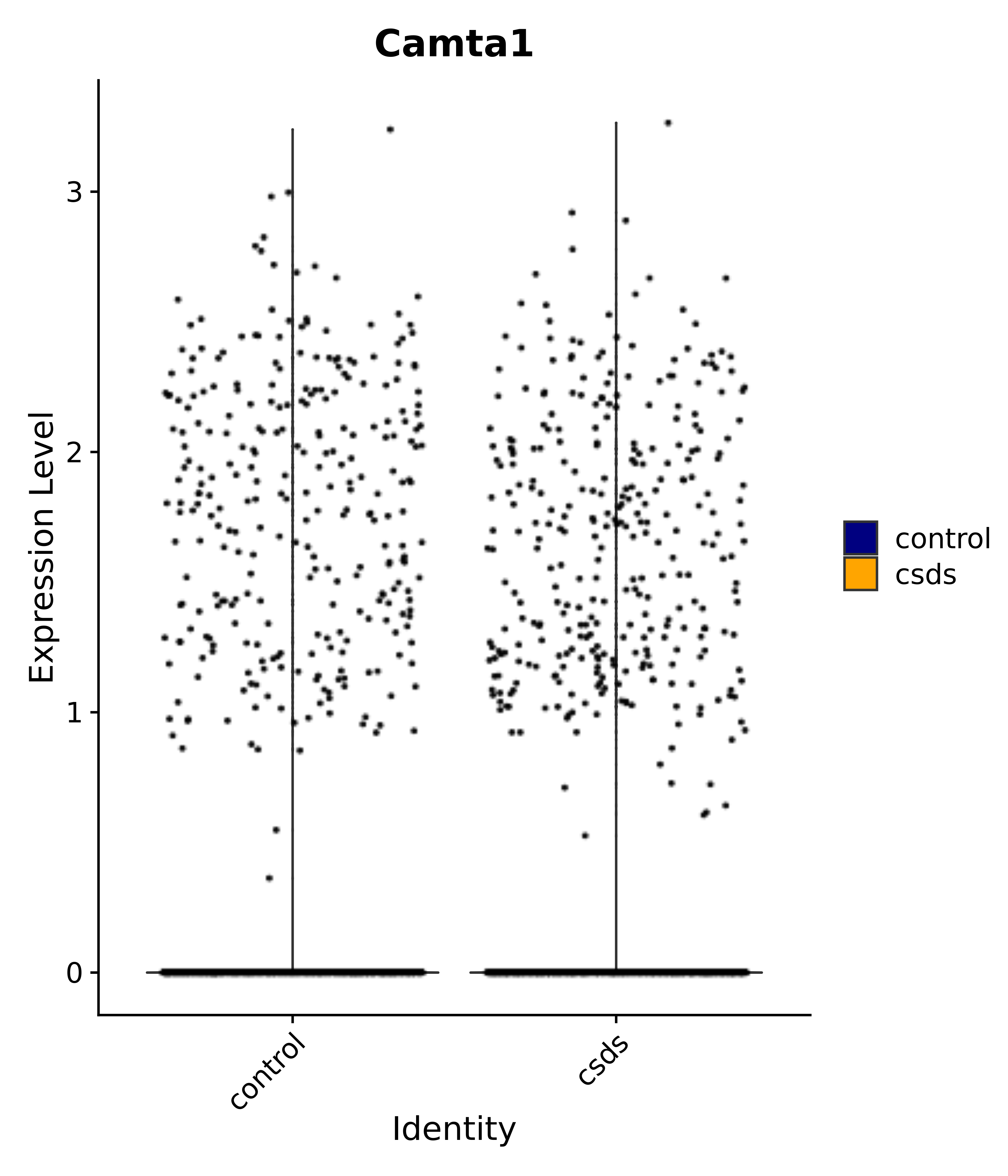
"Camta1",
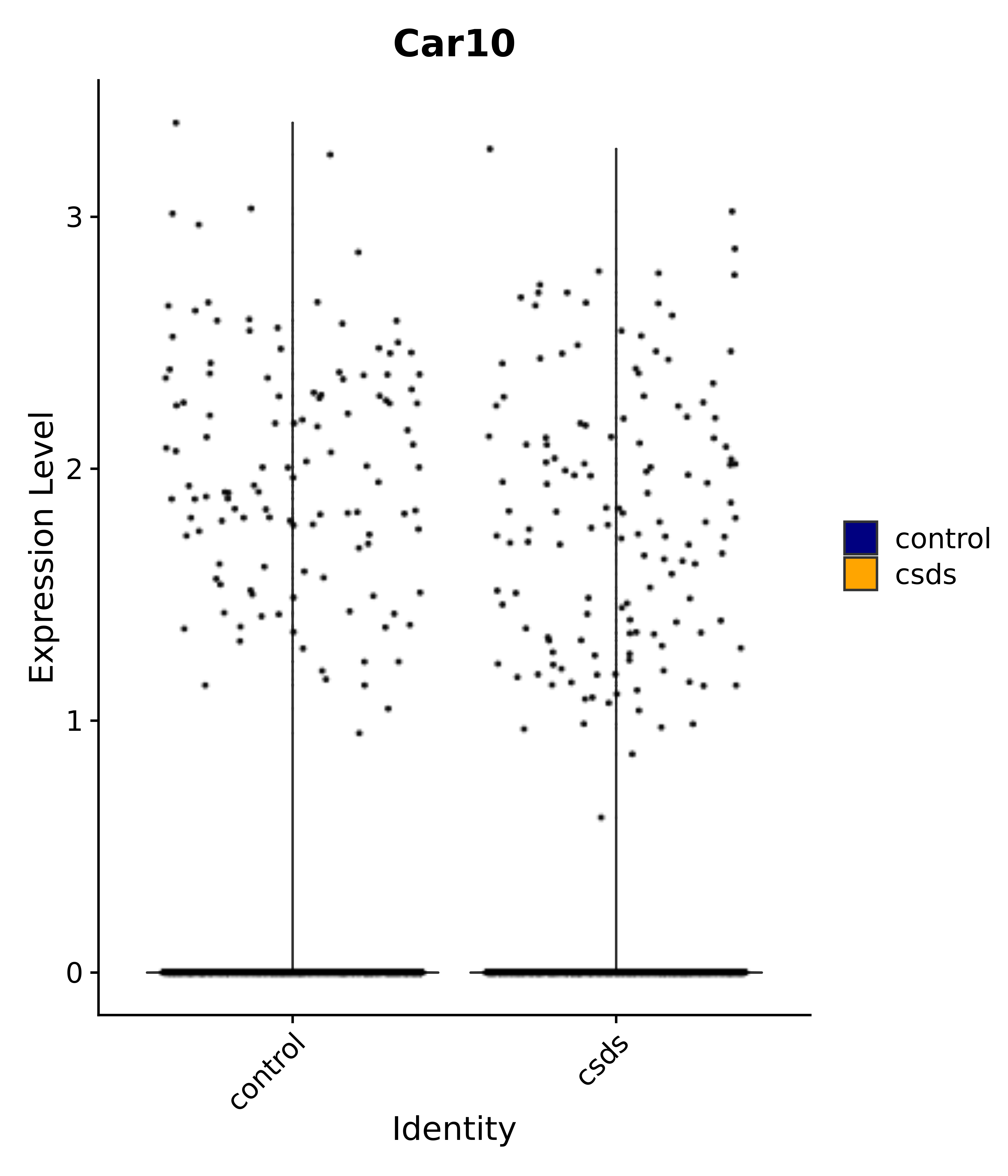
"Car10",
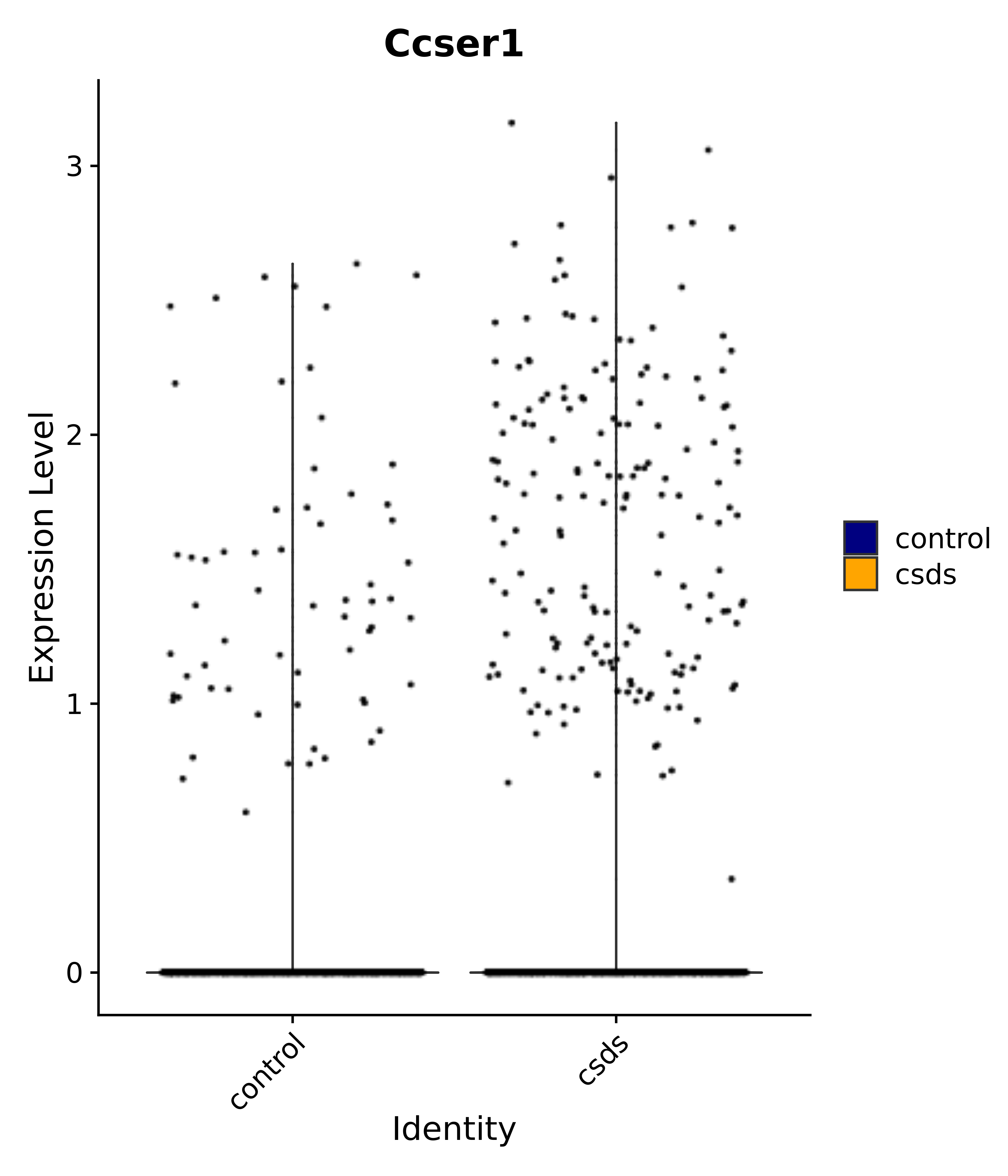
"Ccser1",
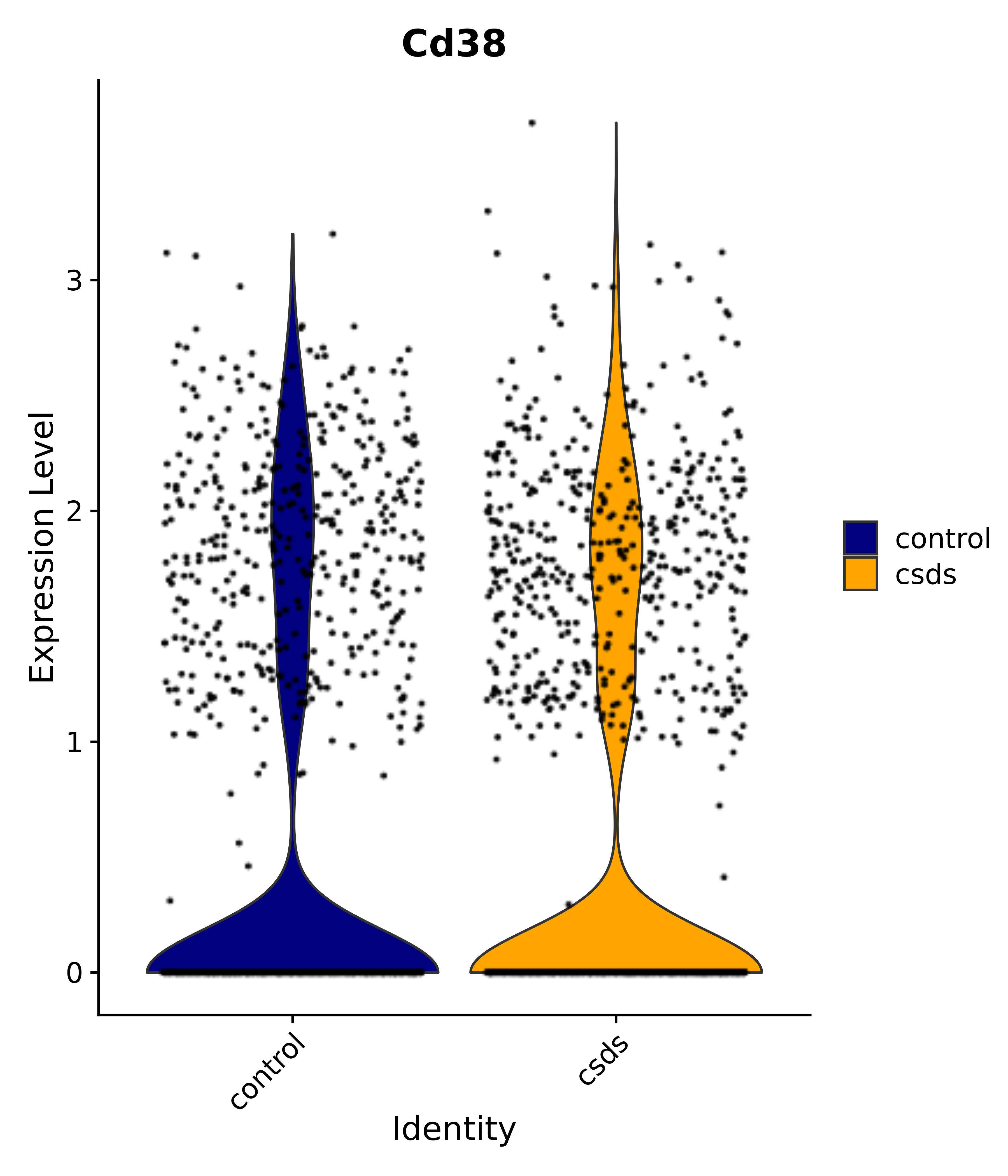
"Cd38",
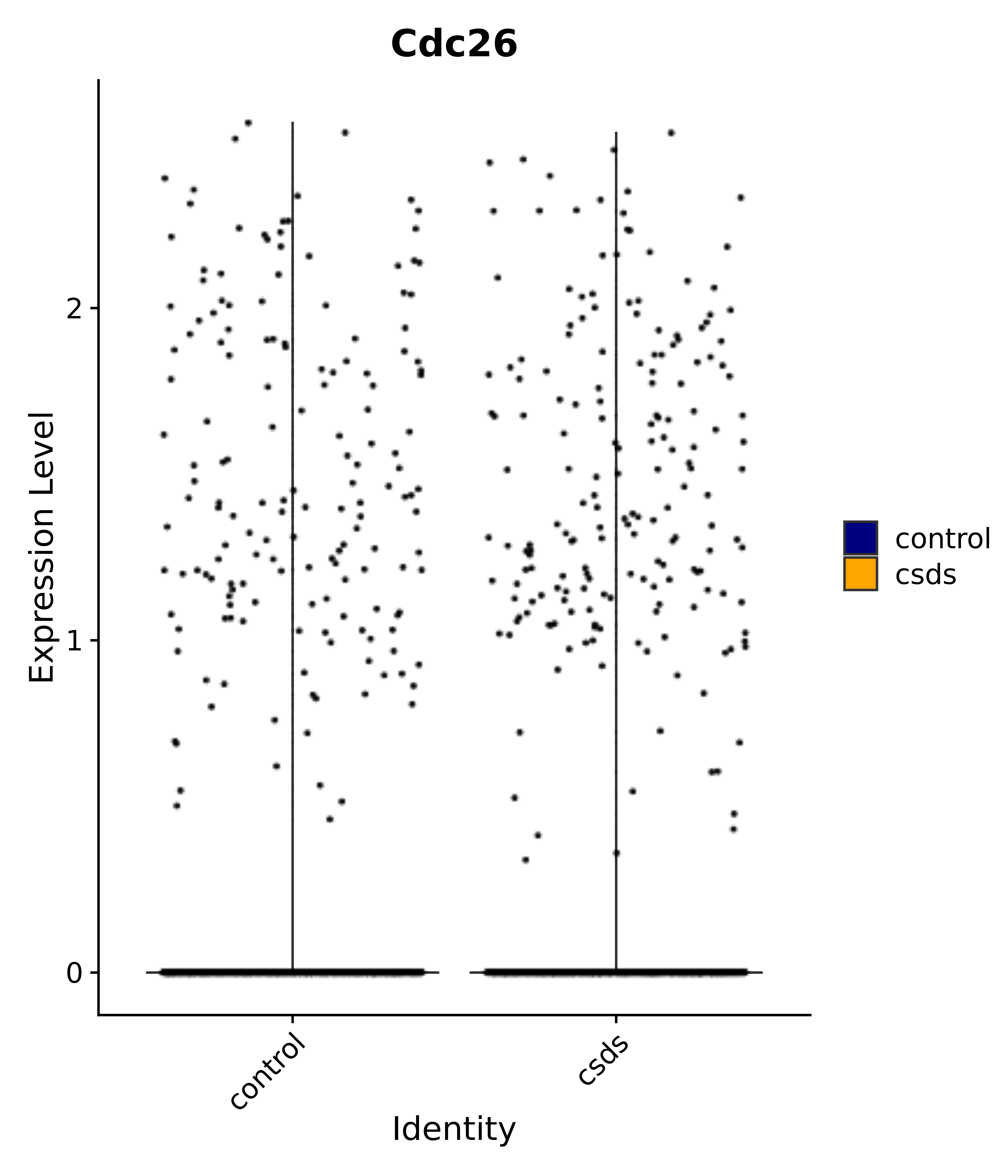
"Cdc26",
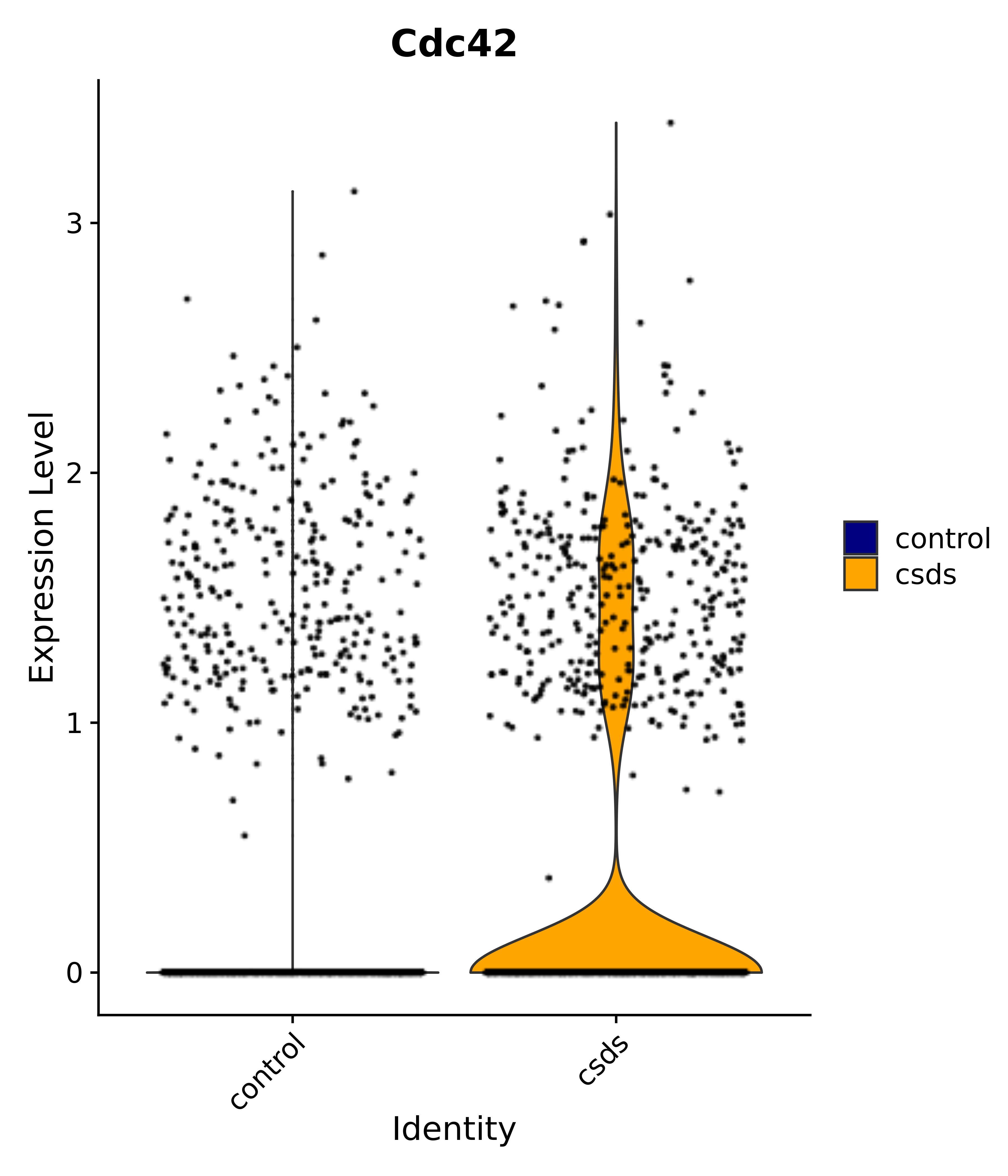
"Cdc42",
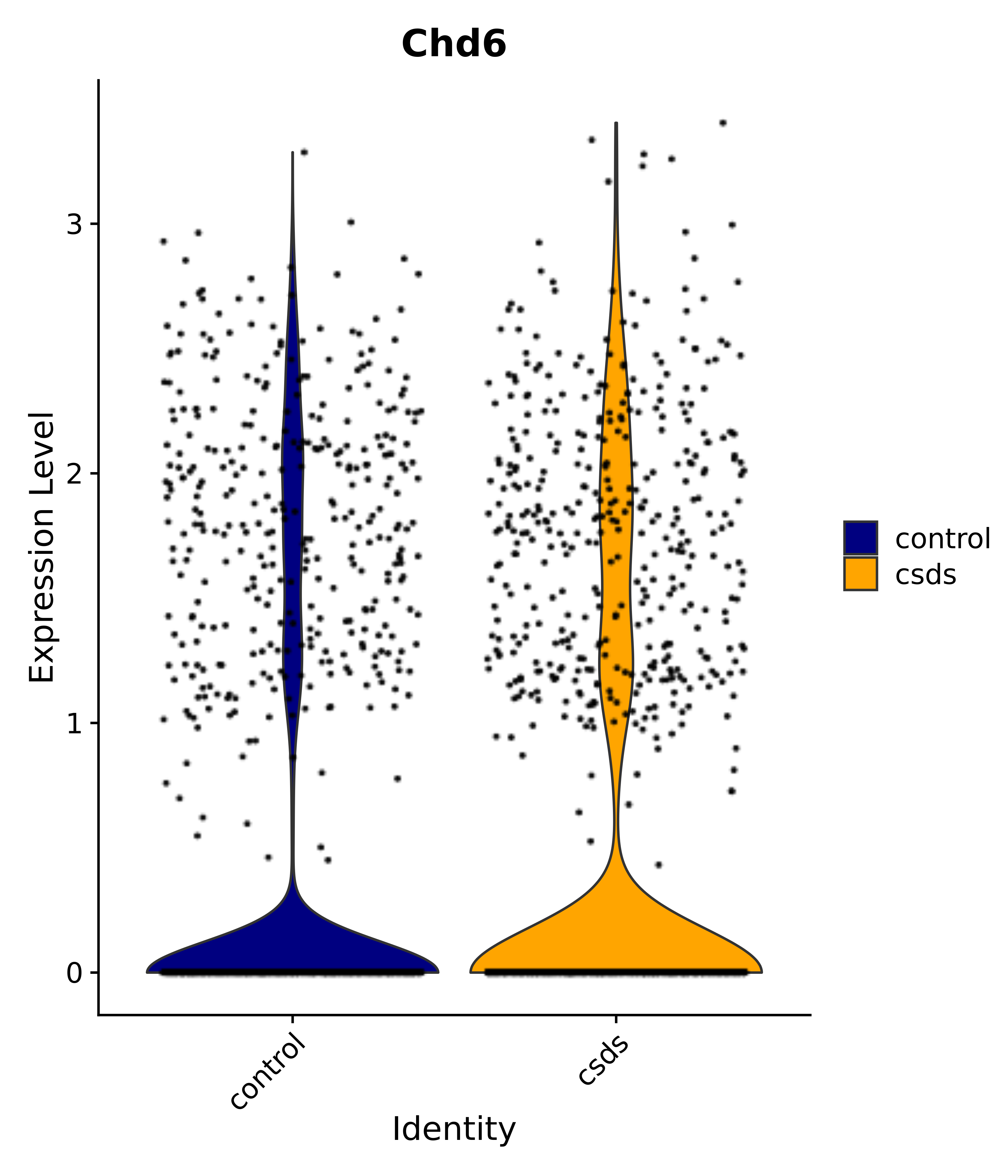
"Chd6",
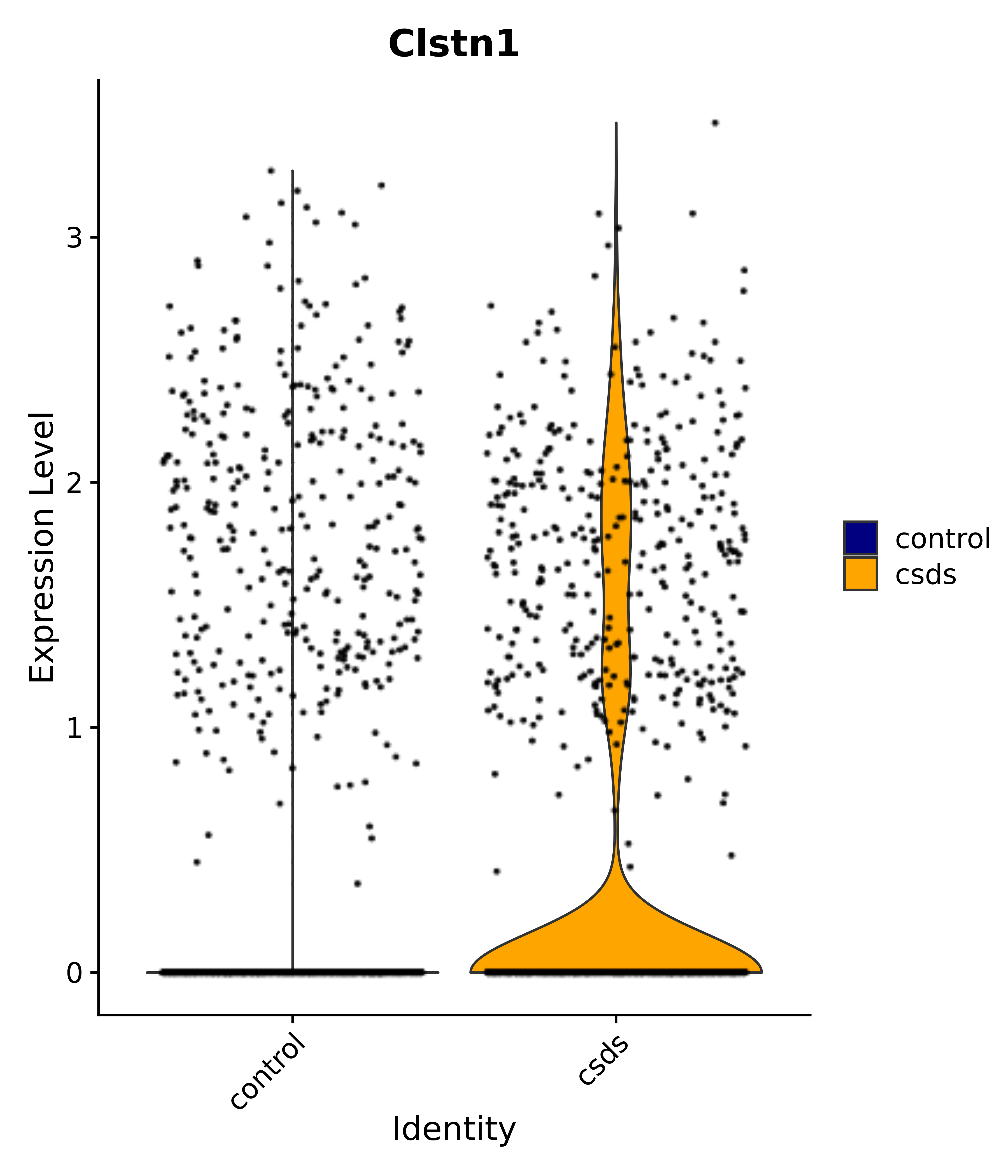
"Clstn1",
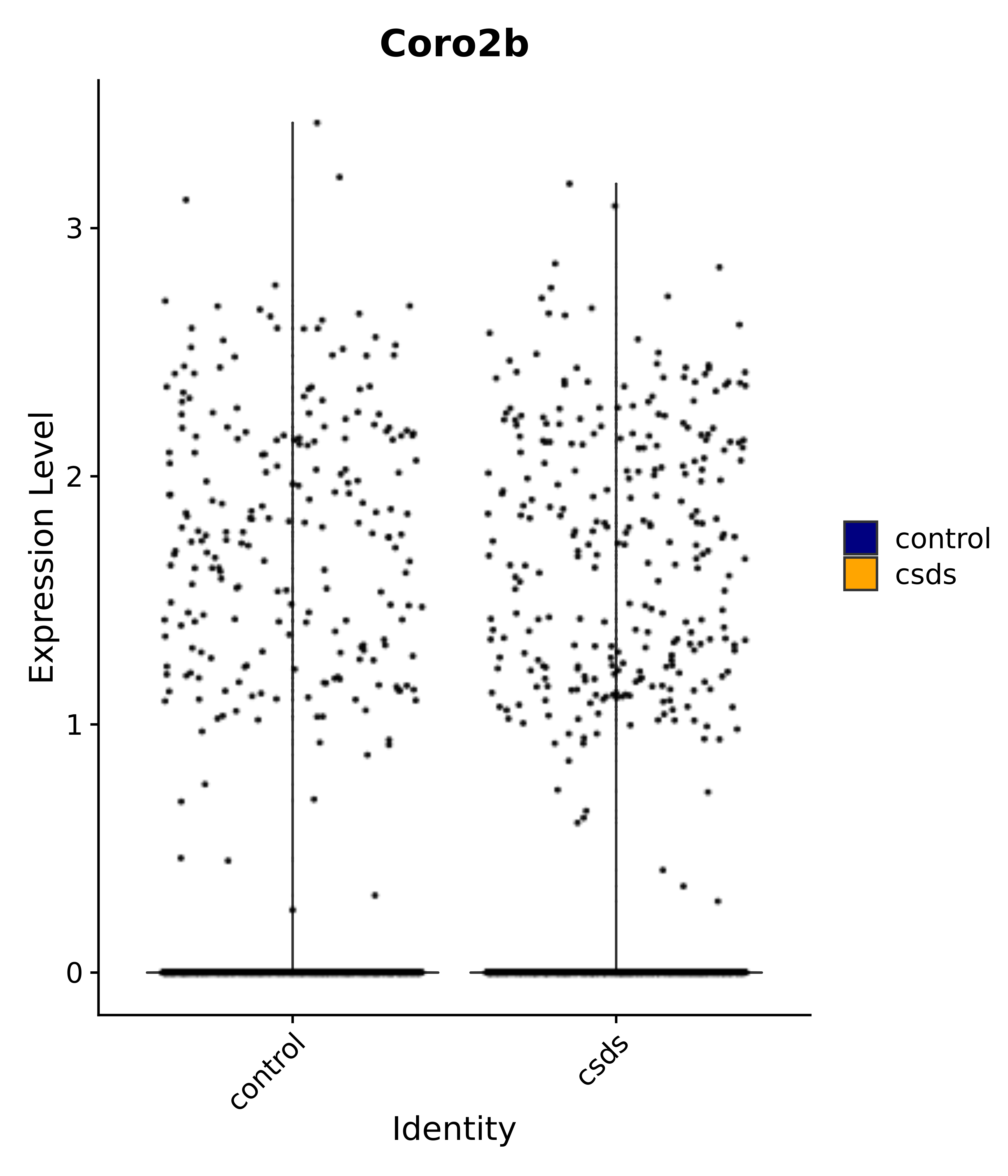
"Coro2b",
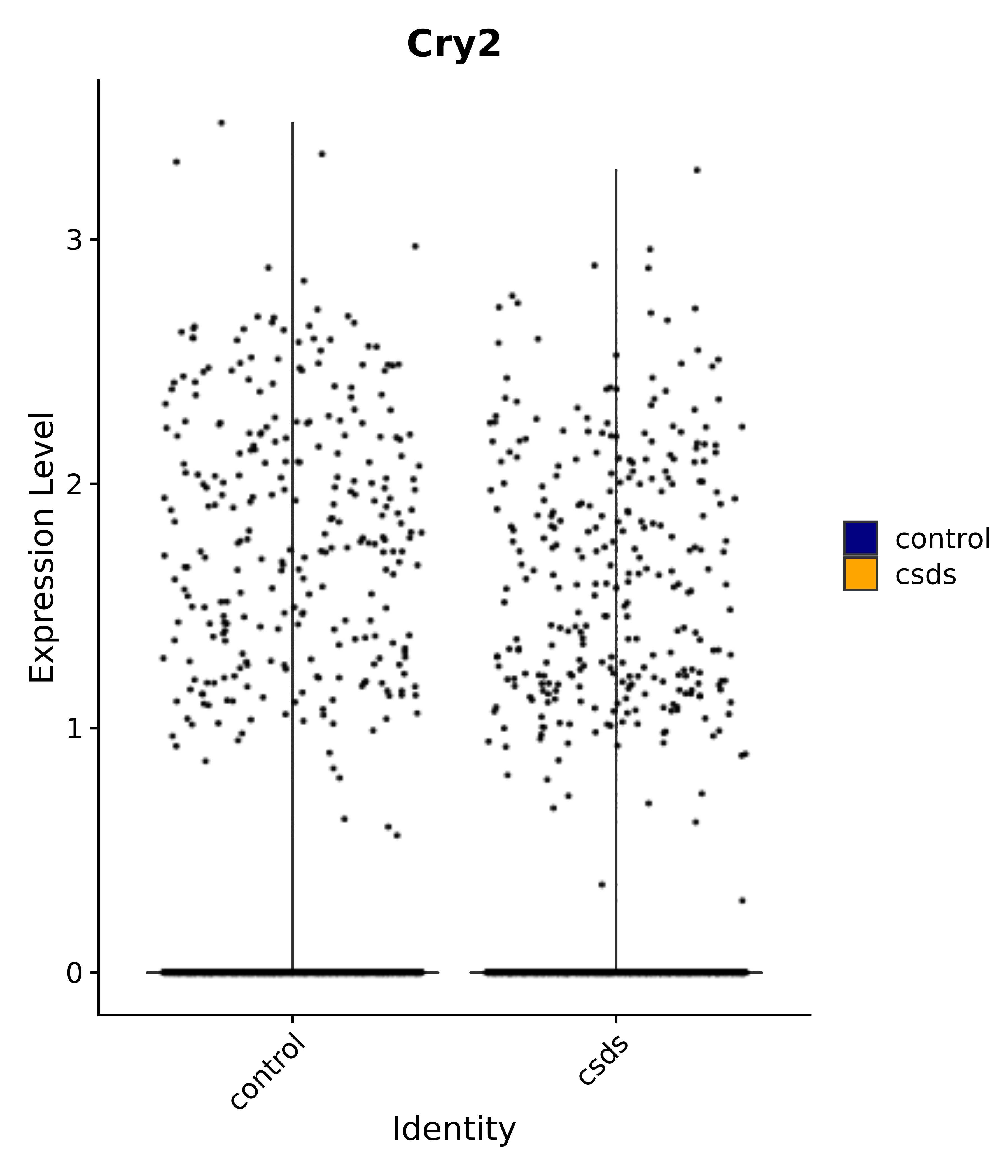
"Cry2",
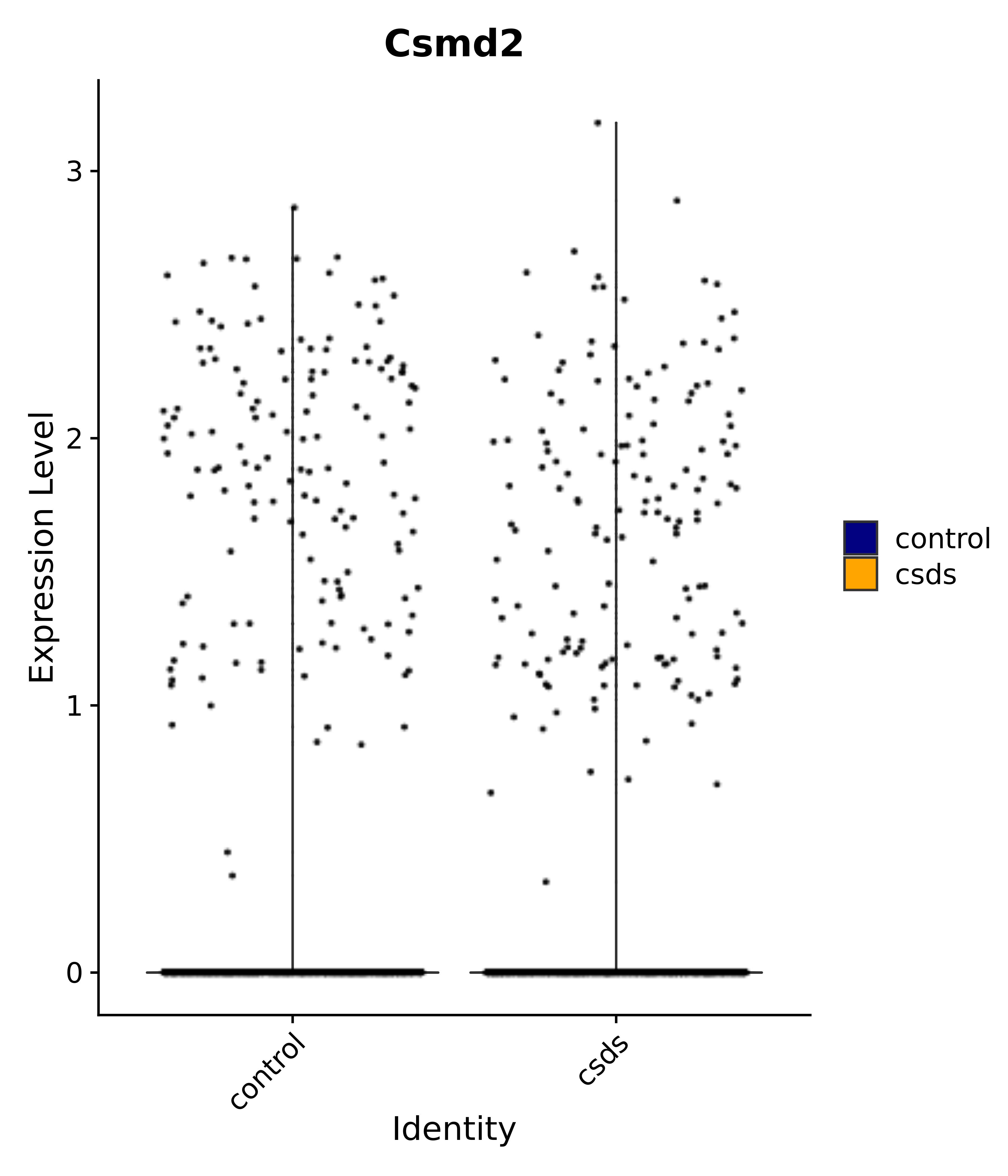
"Csmd2",
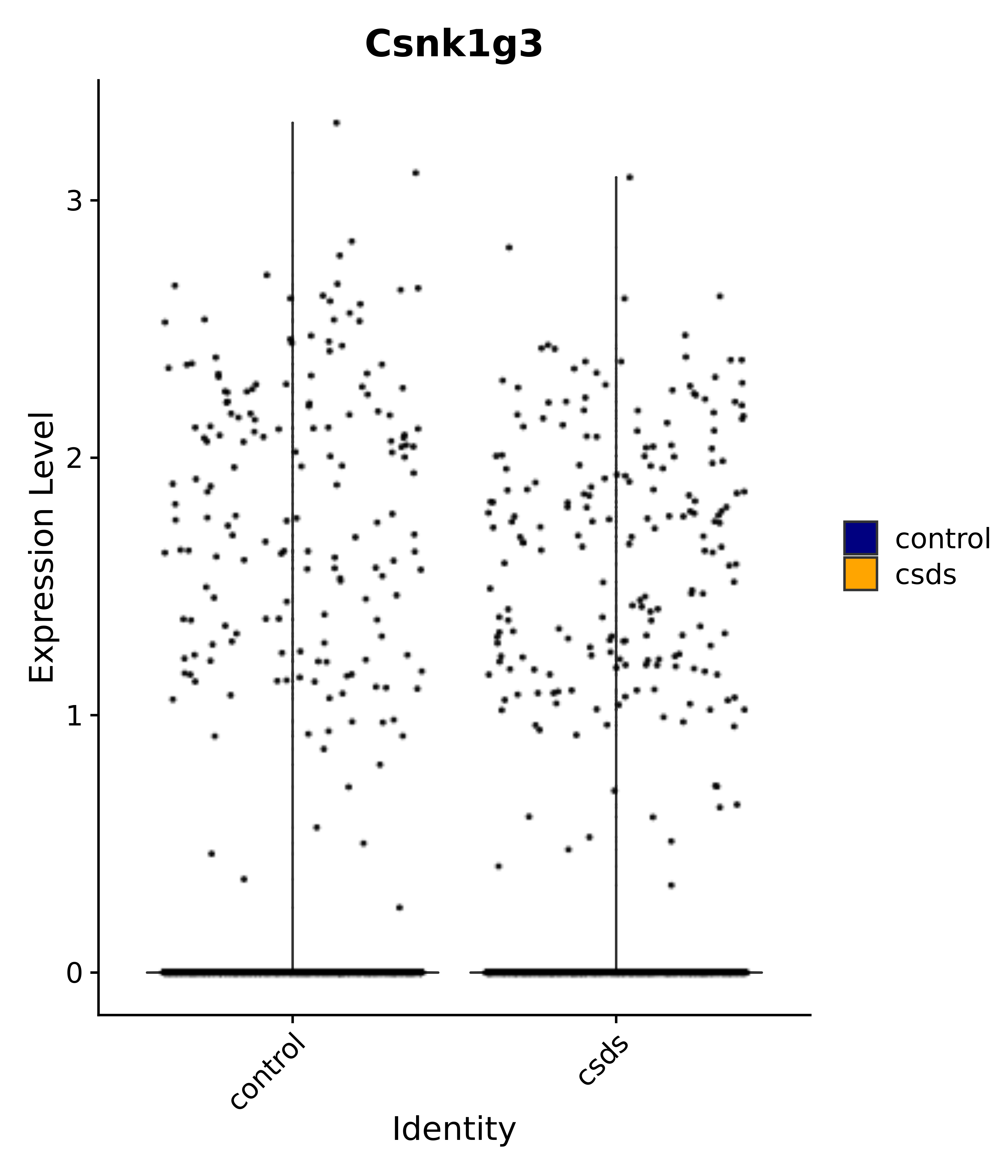
"Csnk1g3",
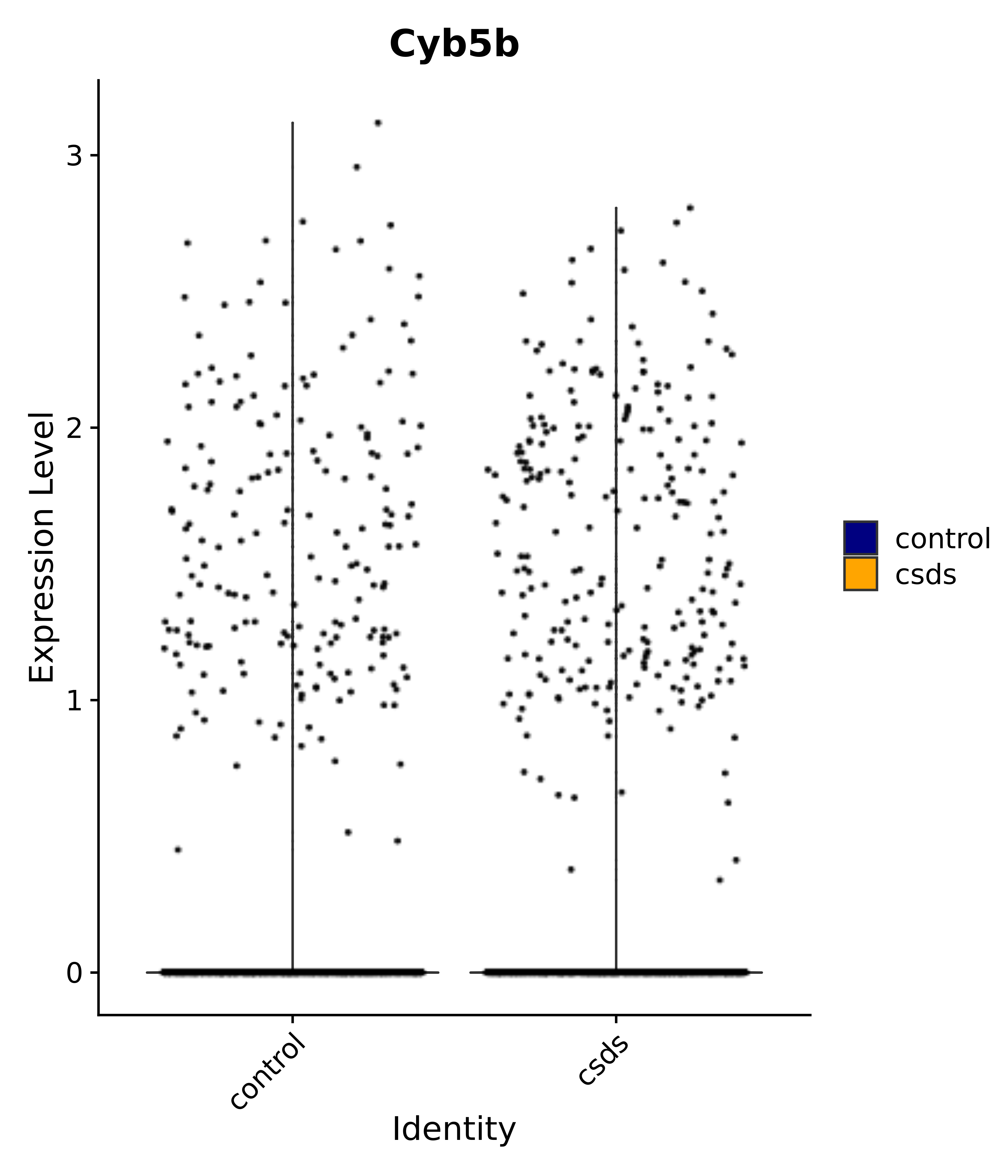
"Cyb5b",
"Dcc",
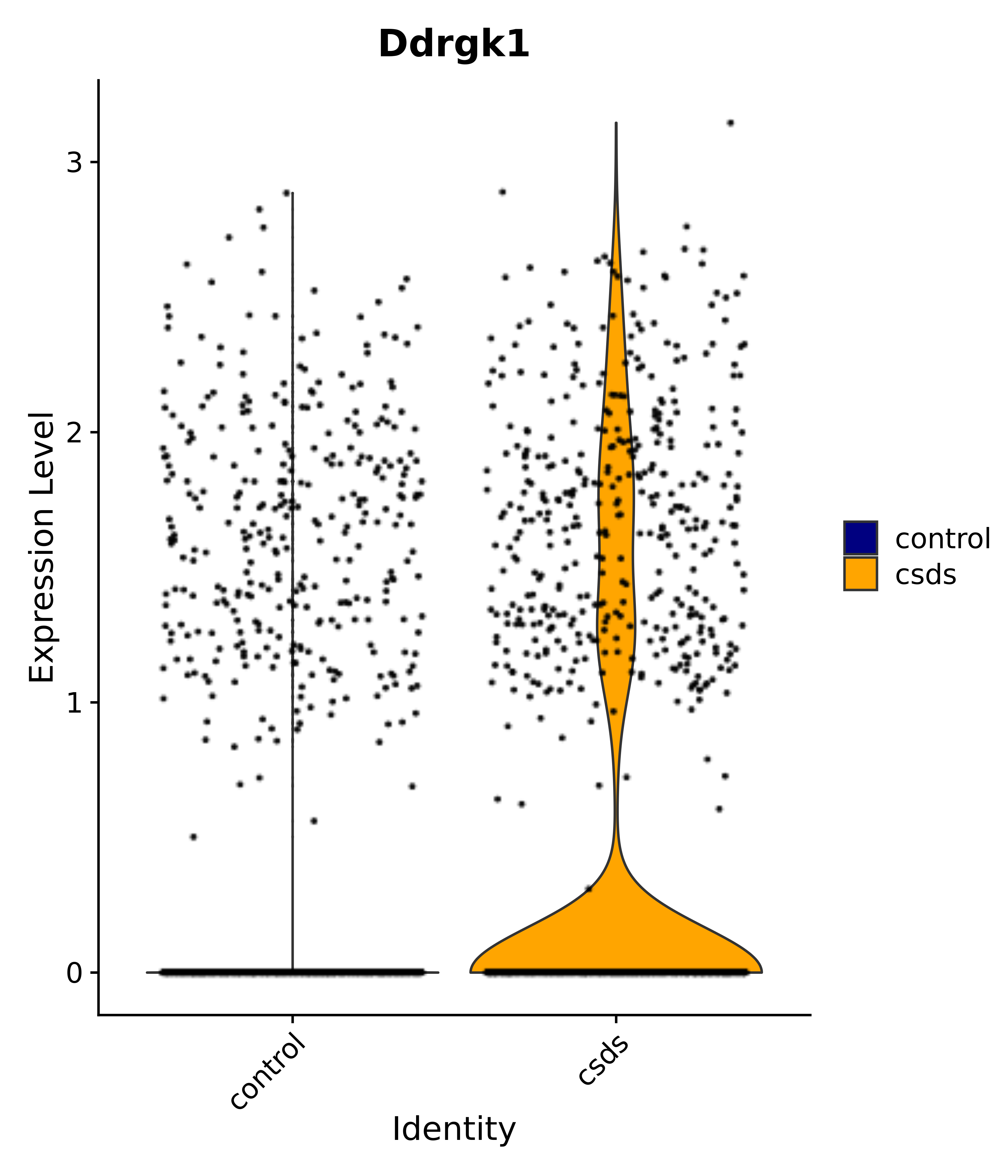
"Ddrgk1",
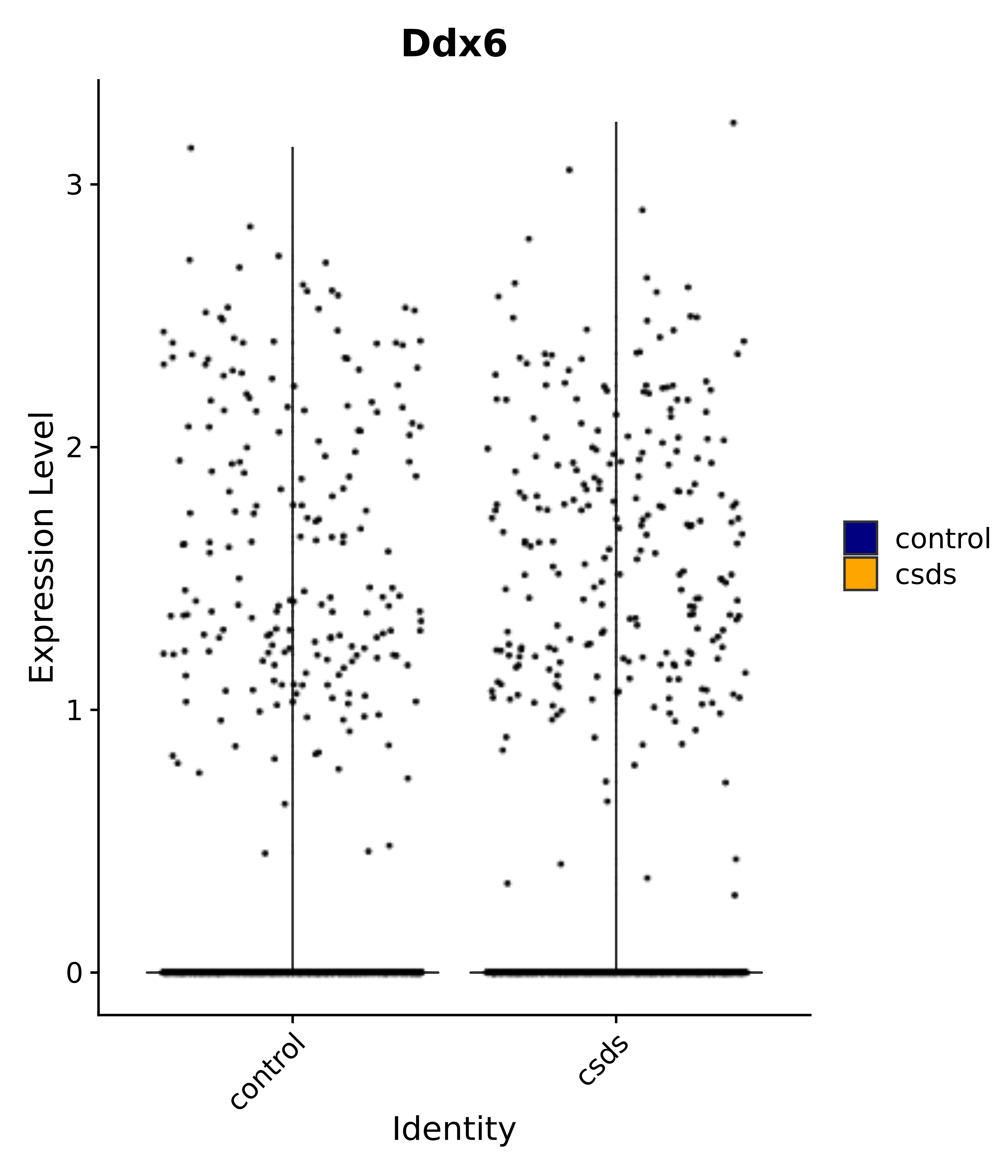
"Ddx6",
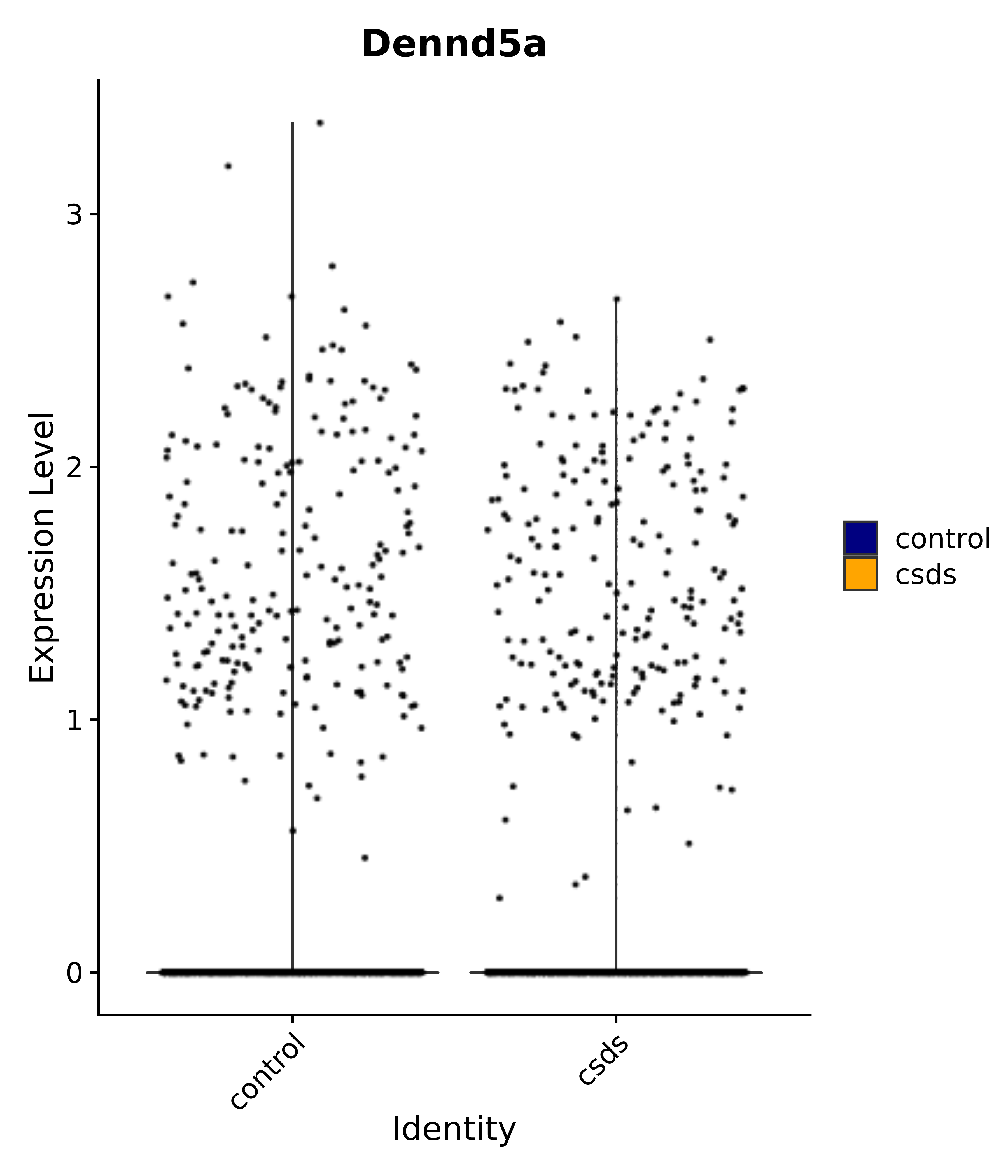
"Dennd5a",
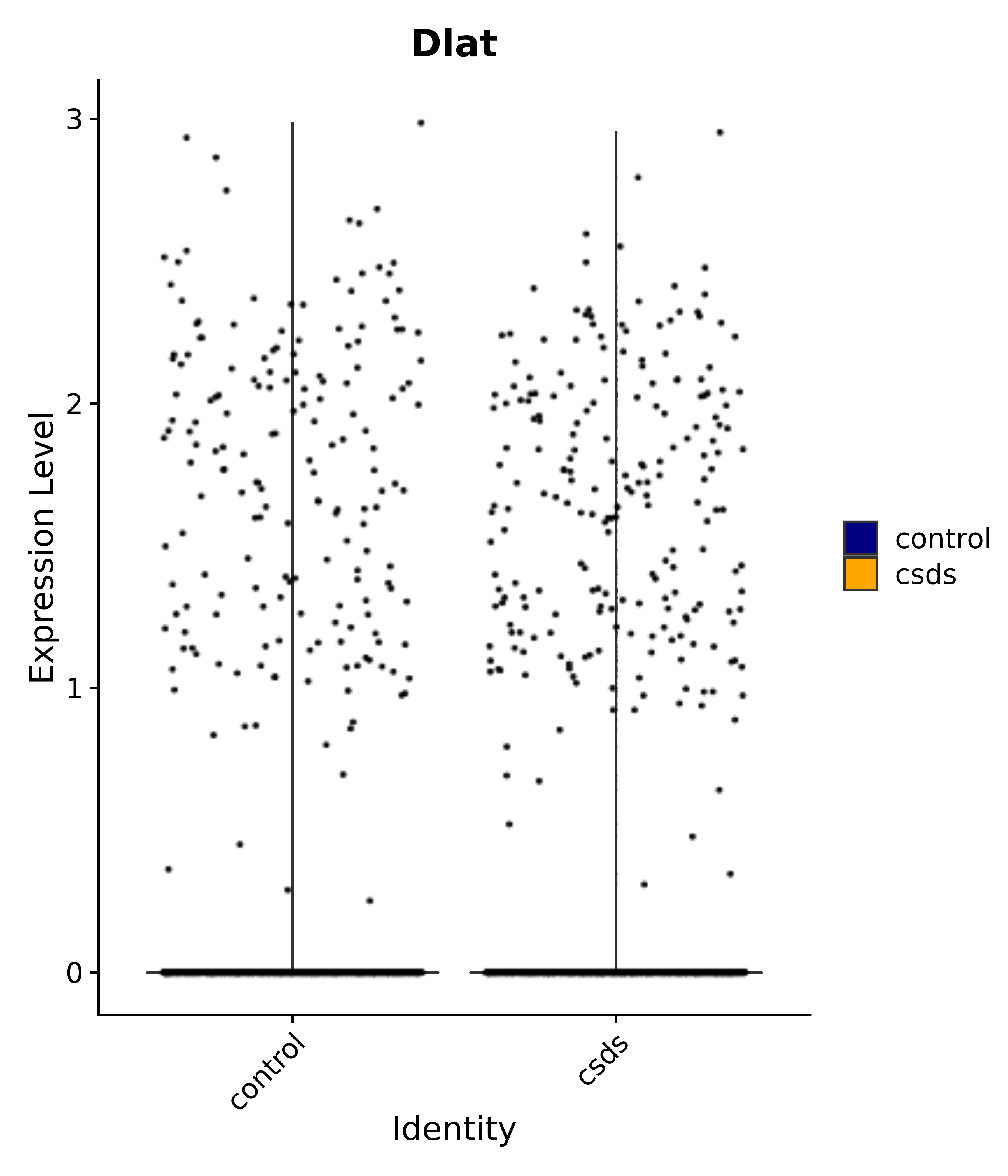
"Dlat",
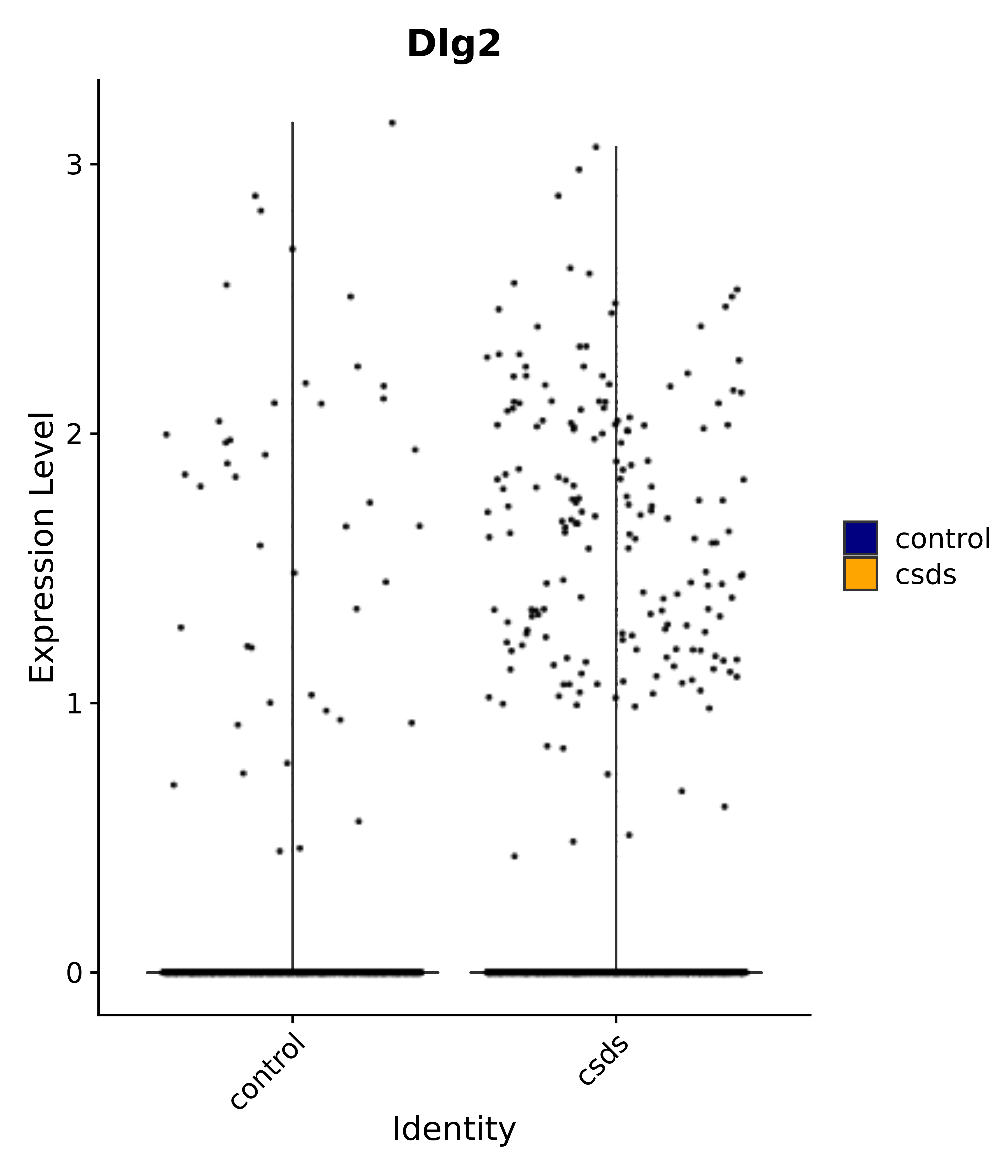
"Dlg2",
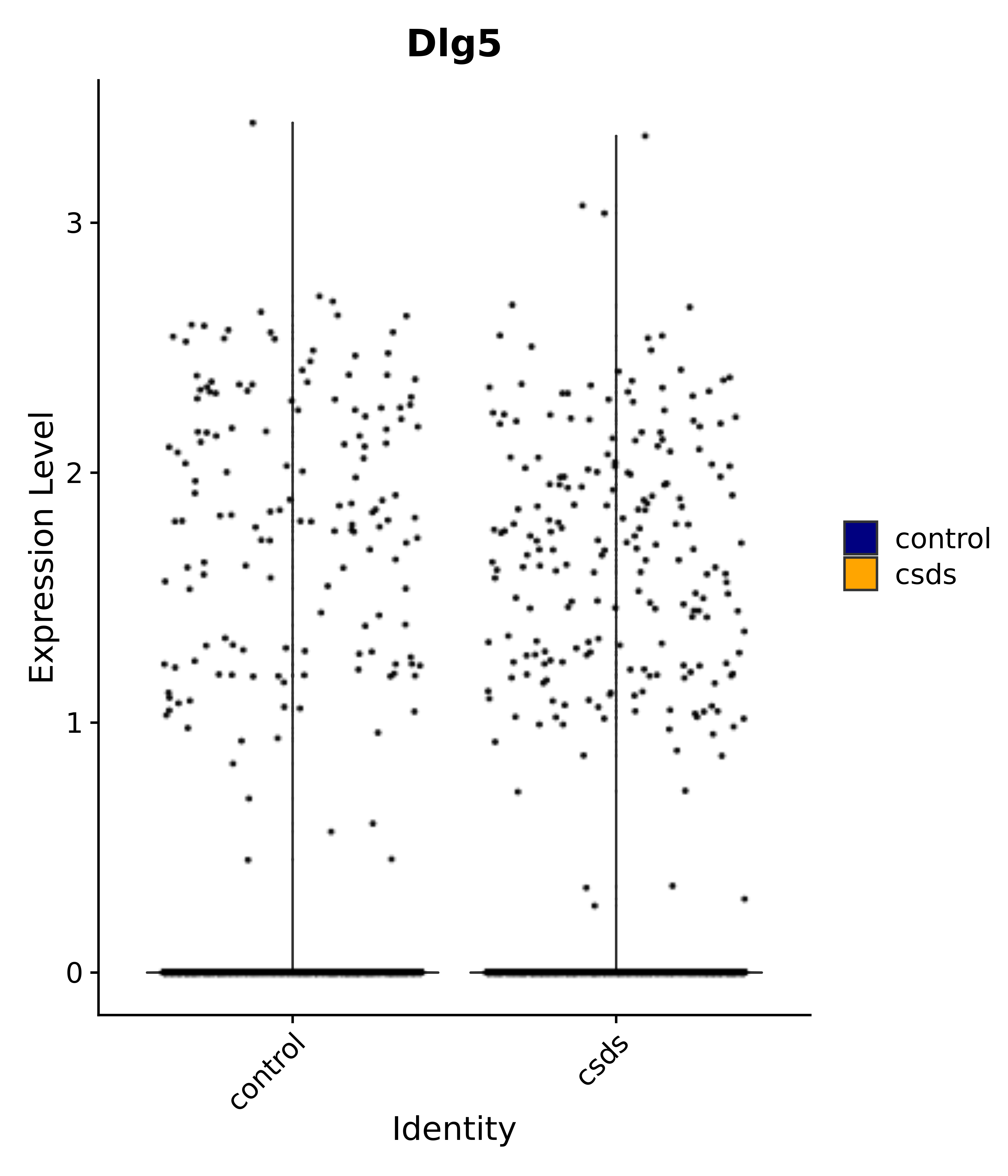
"Dlg5",
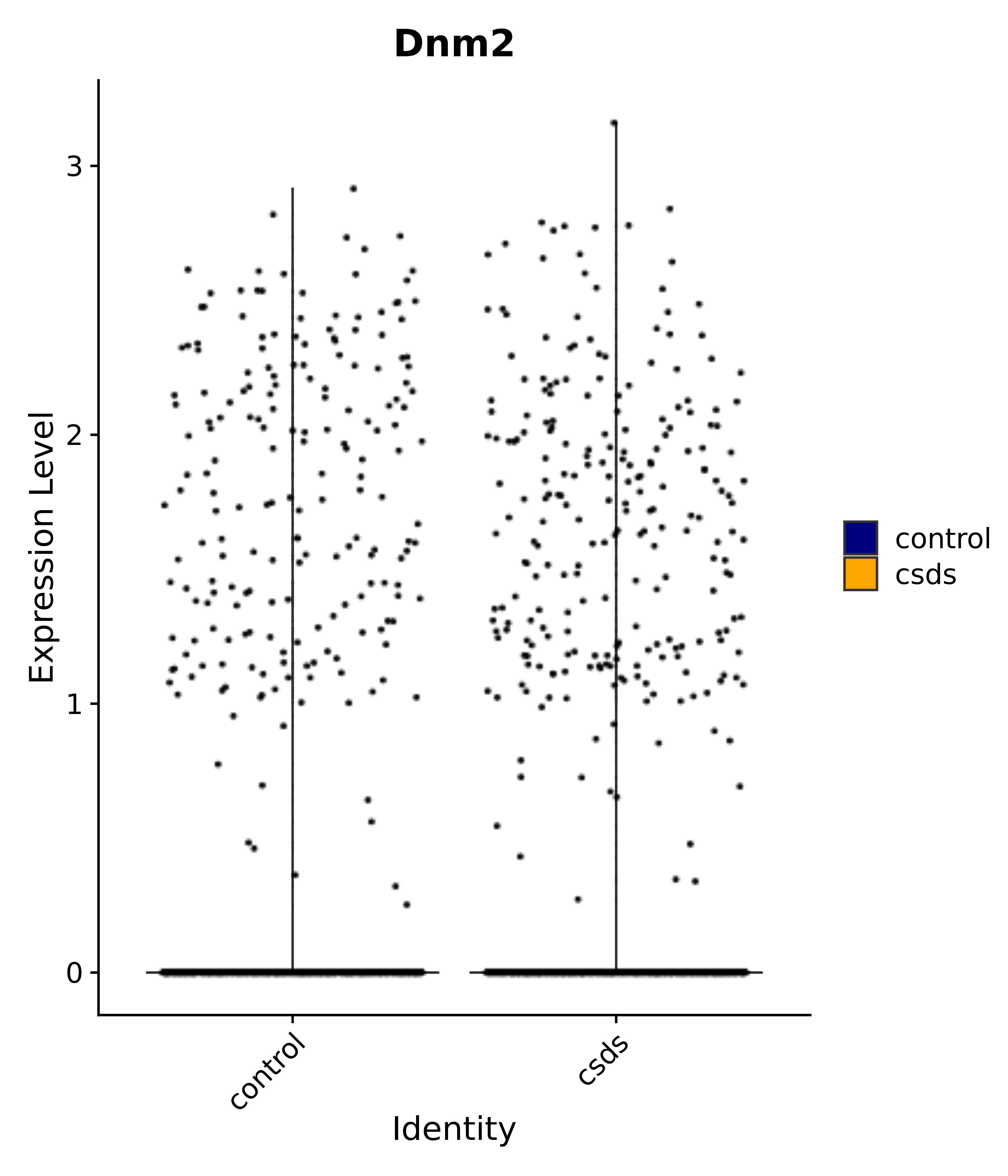
"Dnm2",
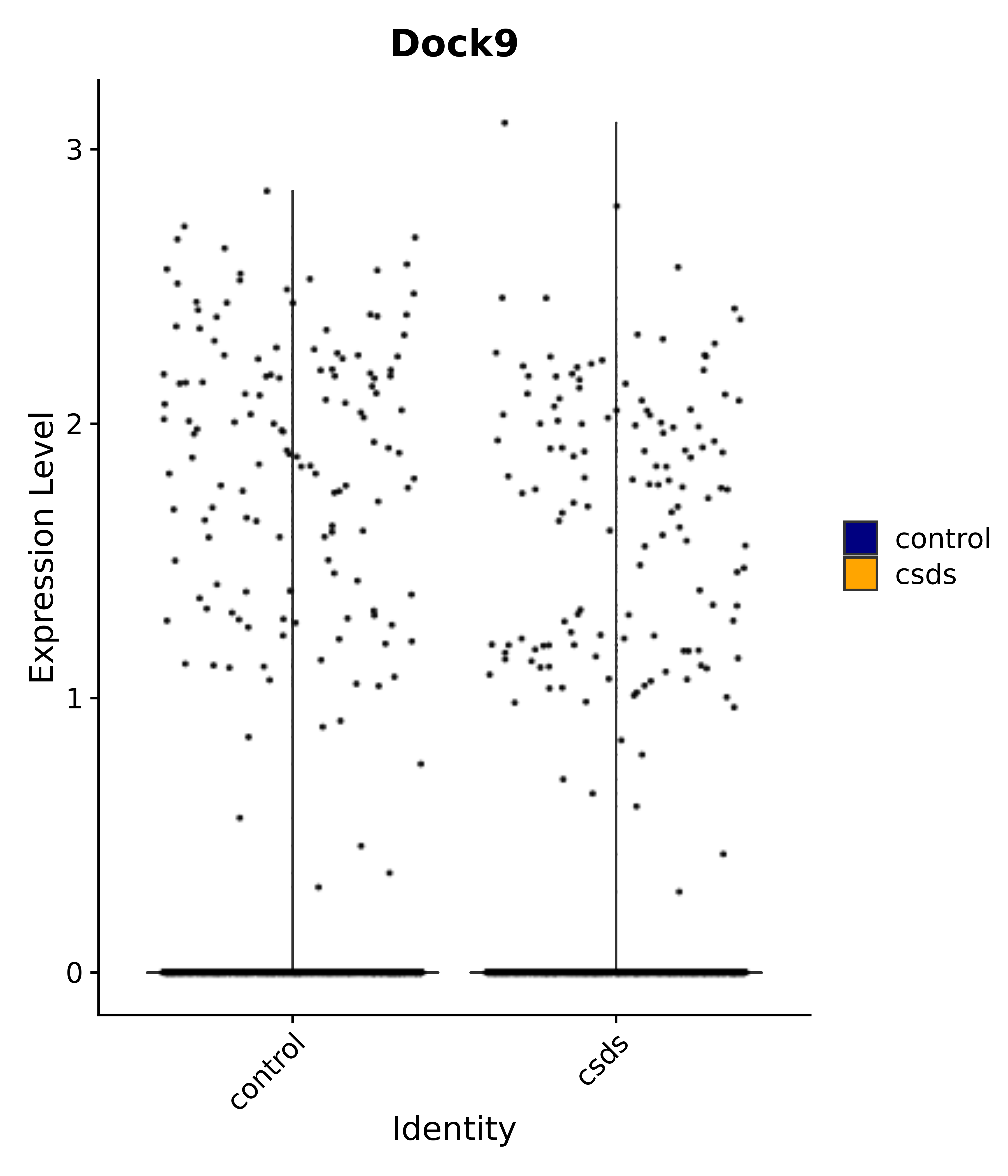
"Dock9",
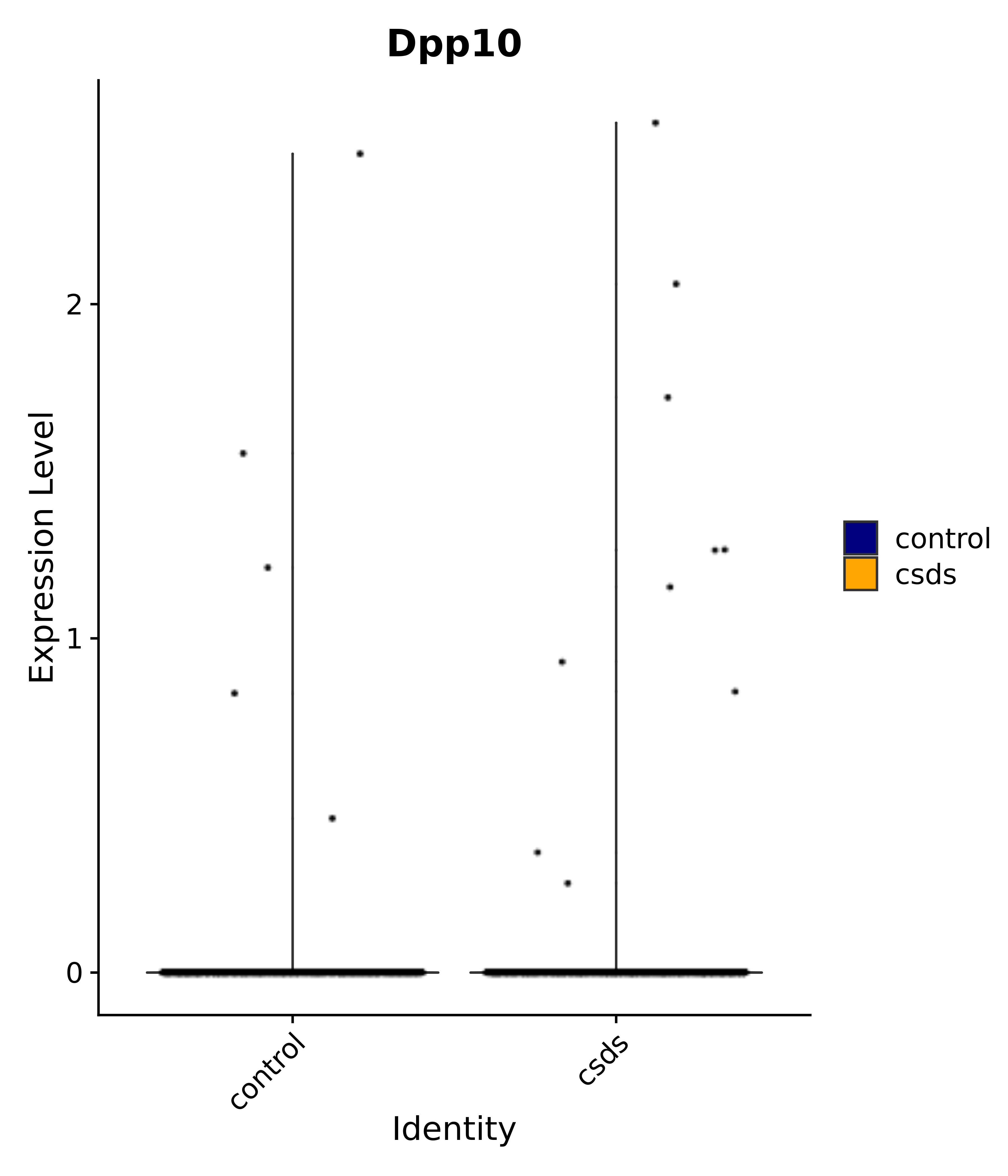
"Dpp10",
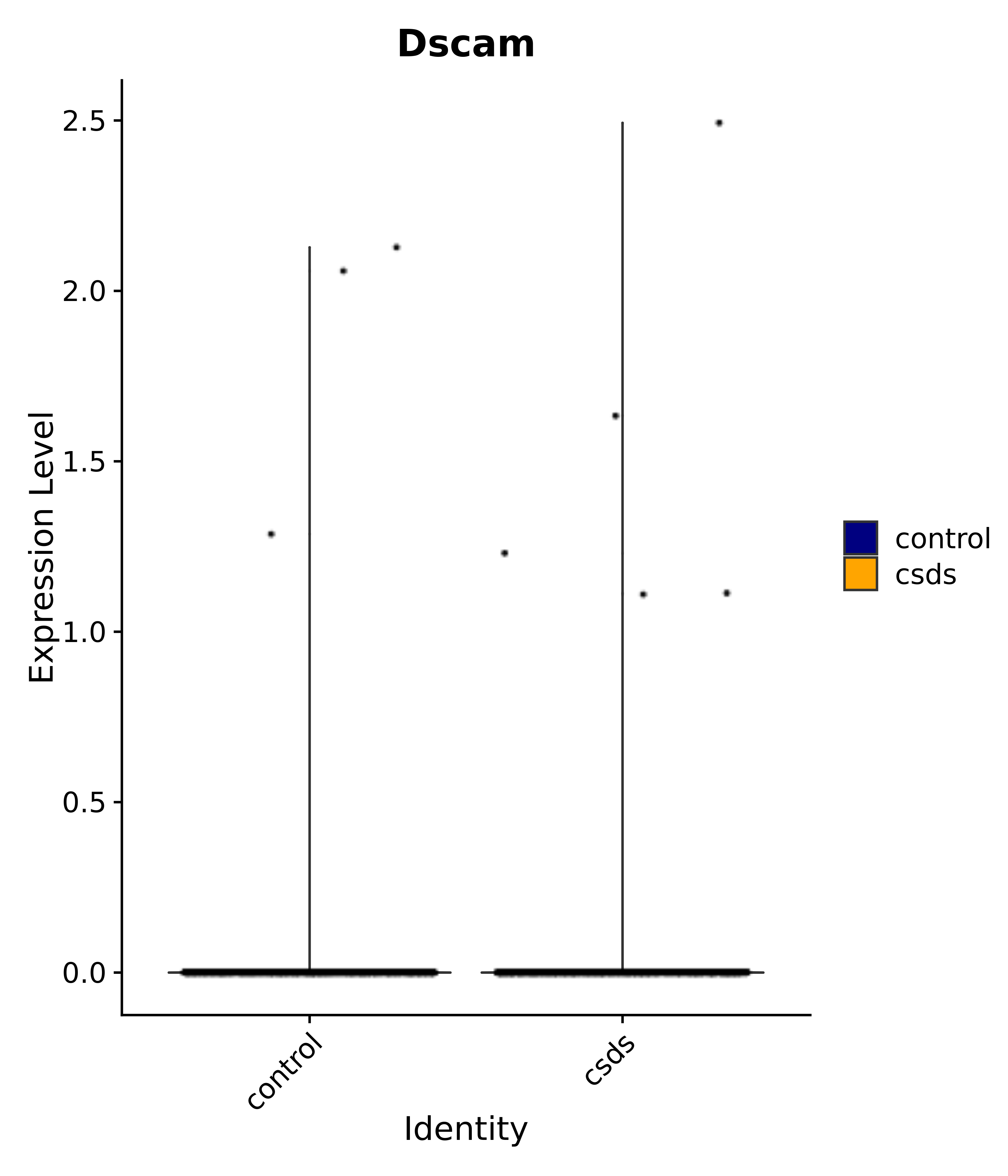
"Dscam",
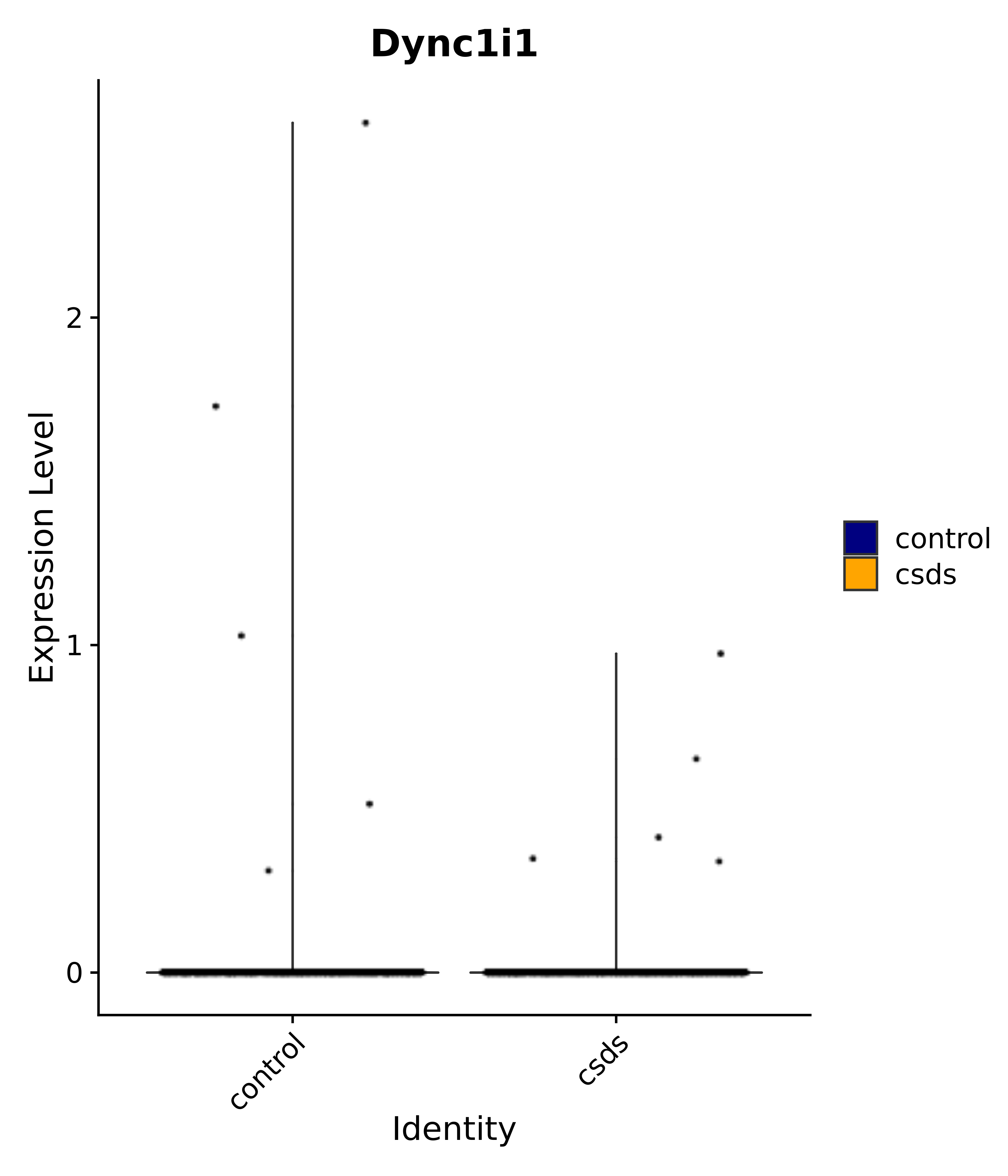
"Dync1i1",
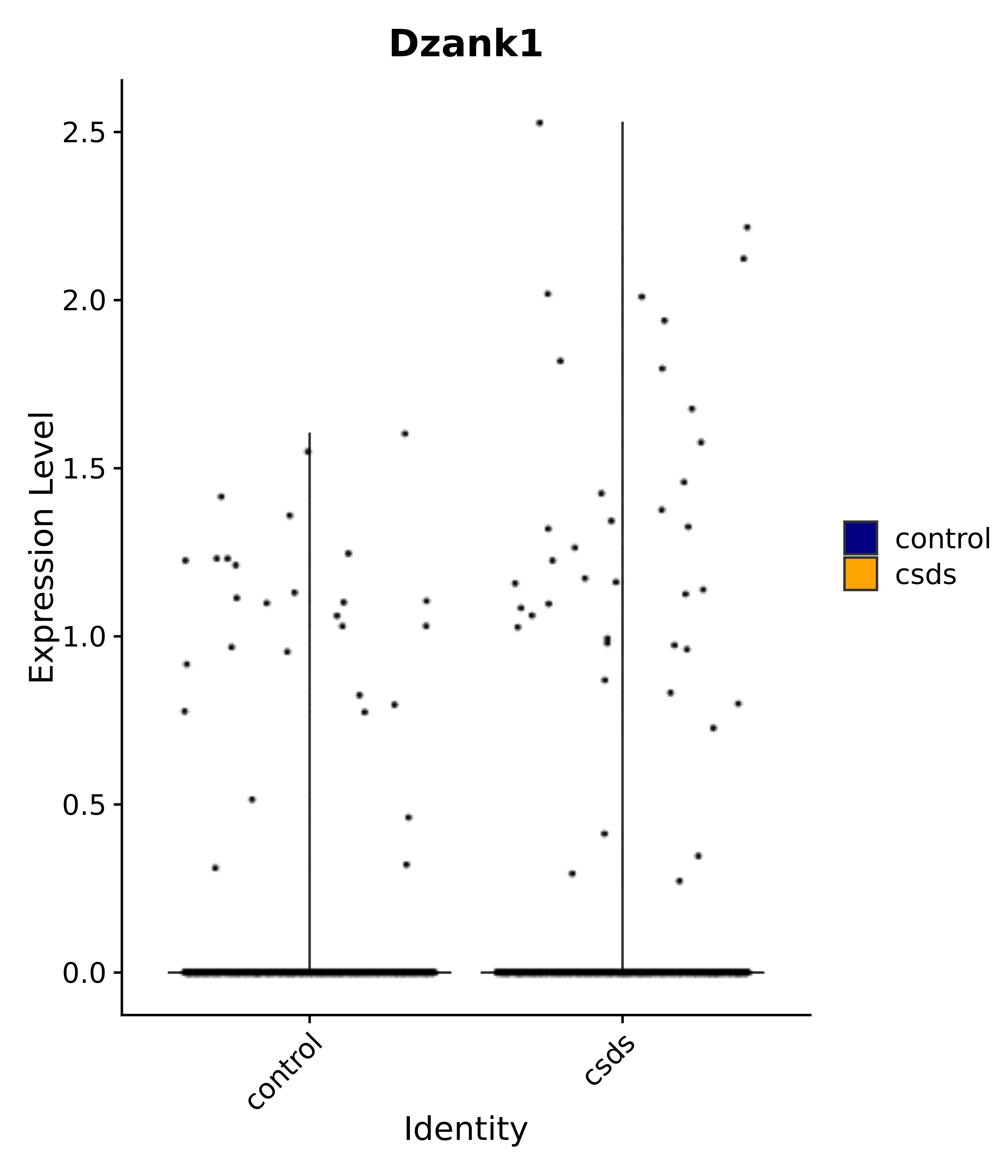
"Dzank1",
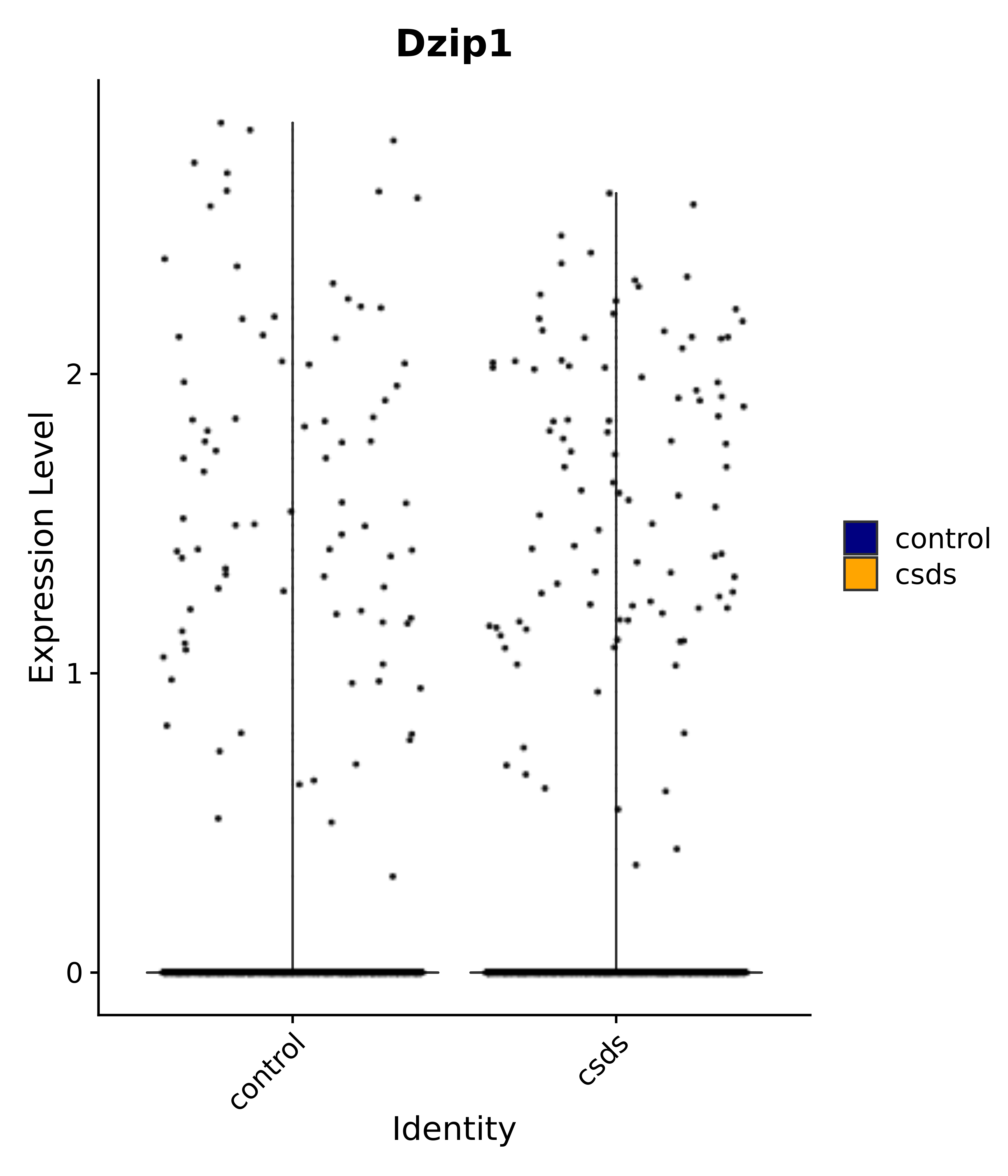
"Dzip1",
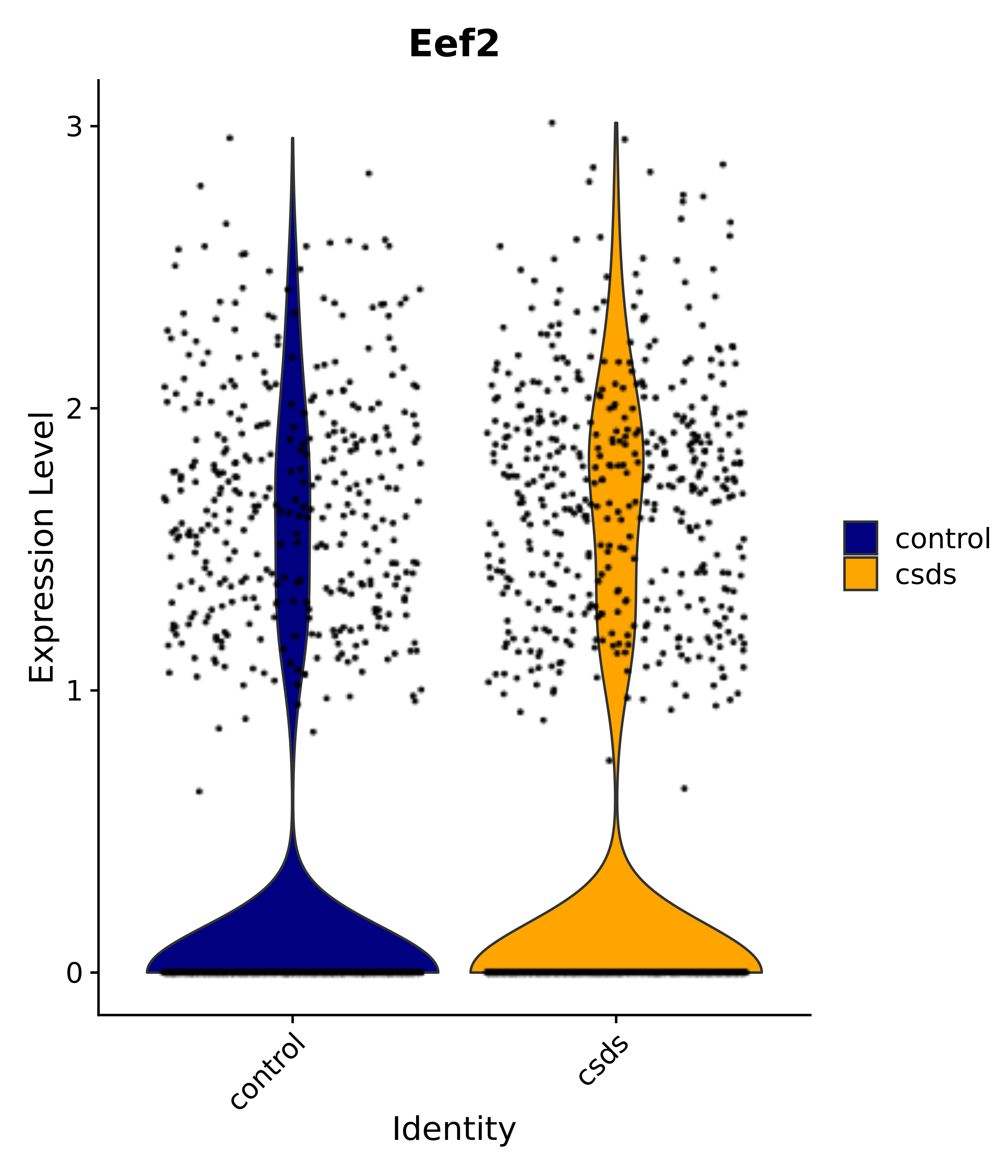
"Eef2",
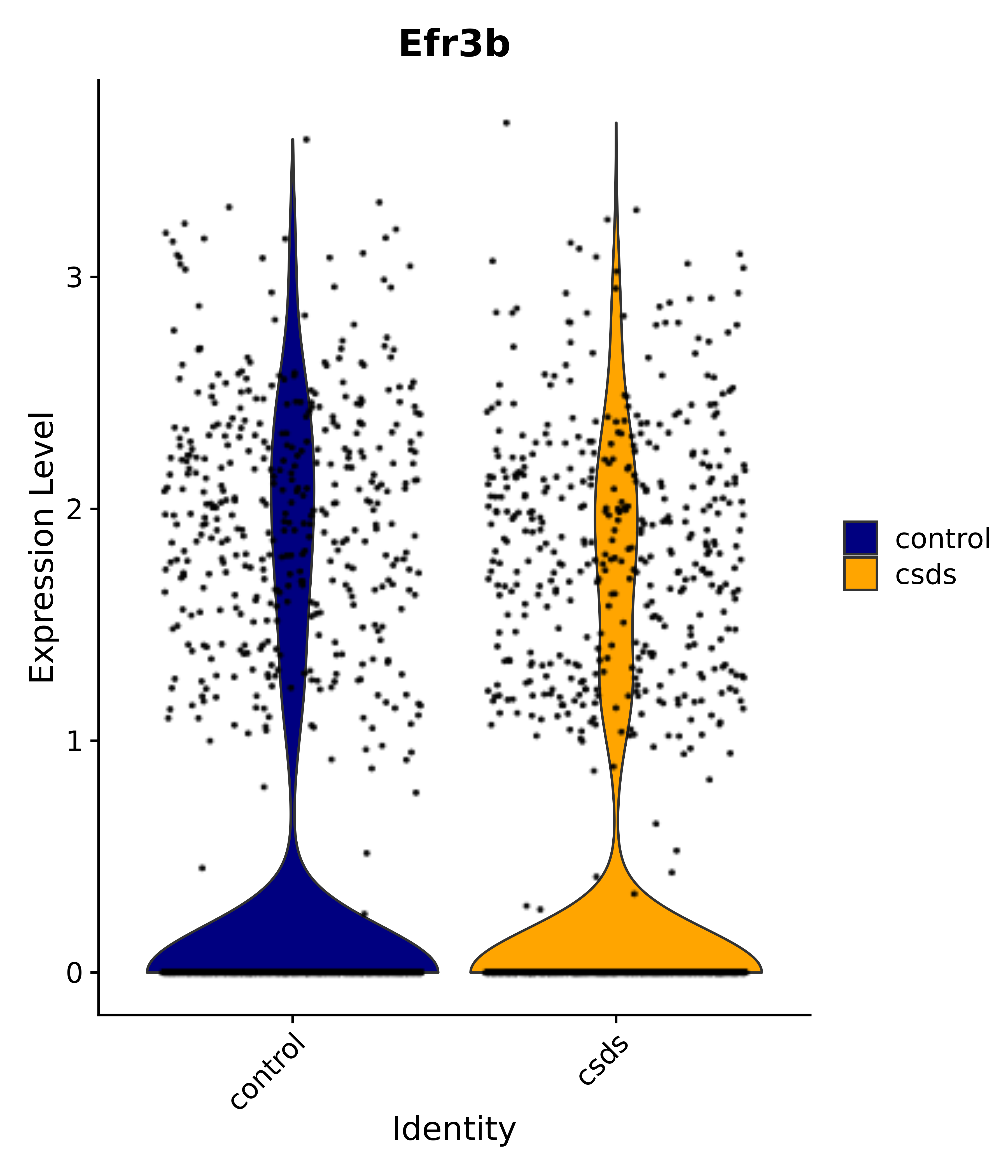
"Efr3b",
"Egfem1",
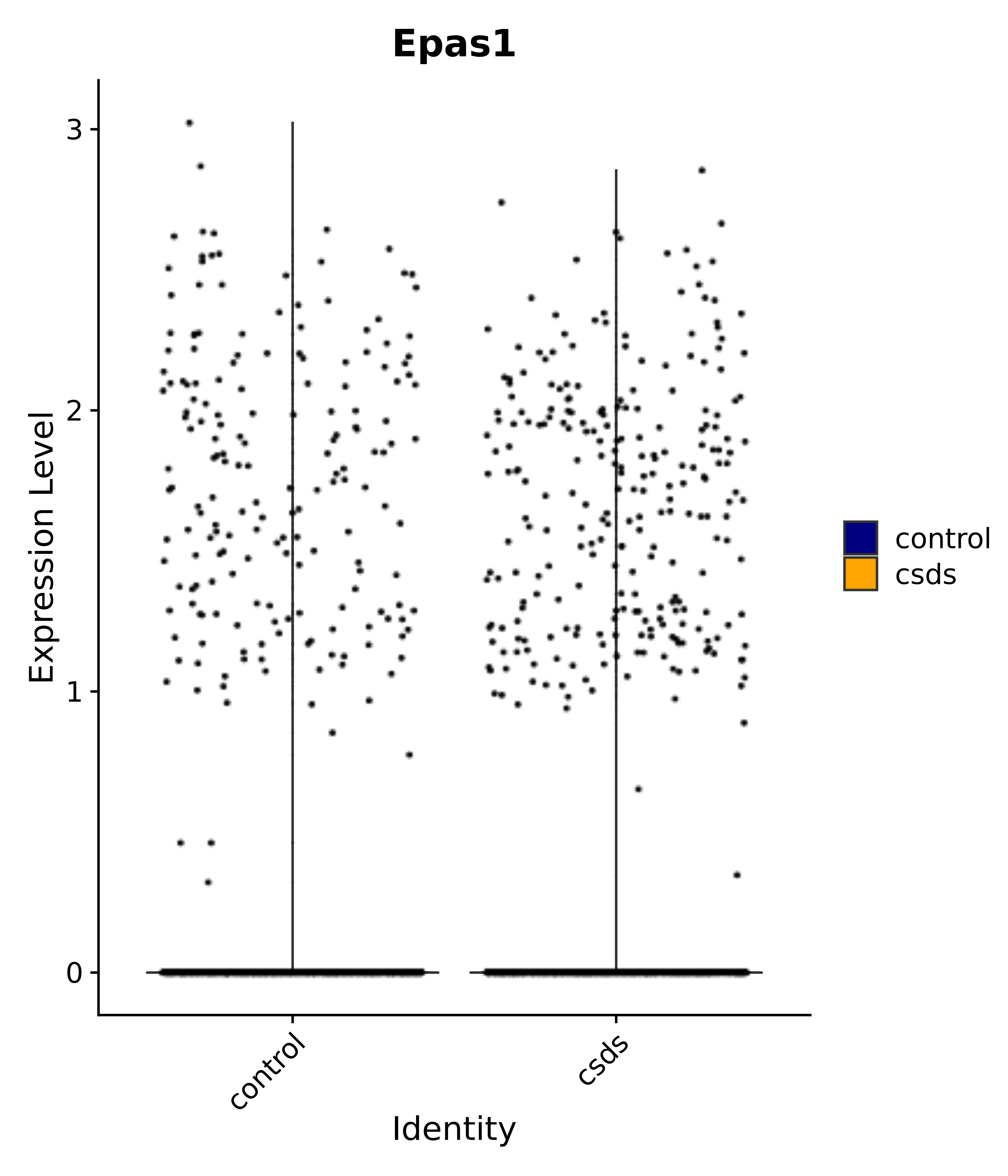
"Epas1",
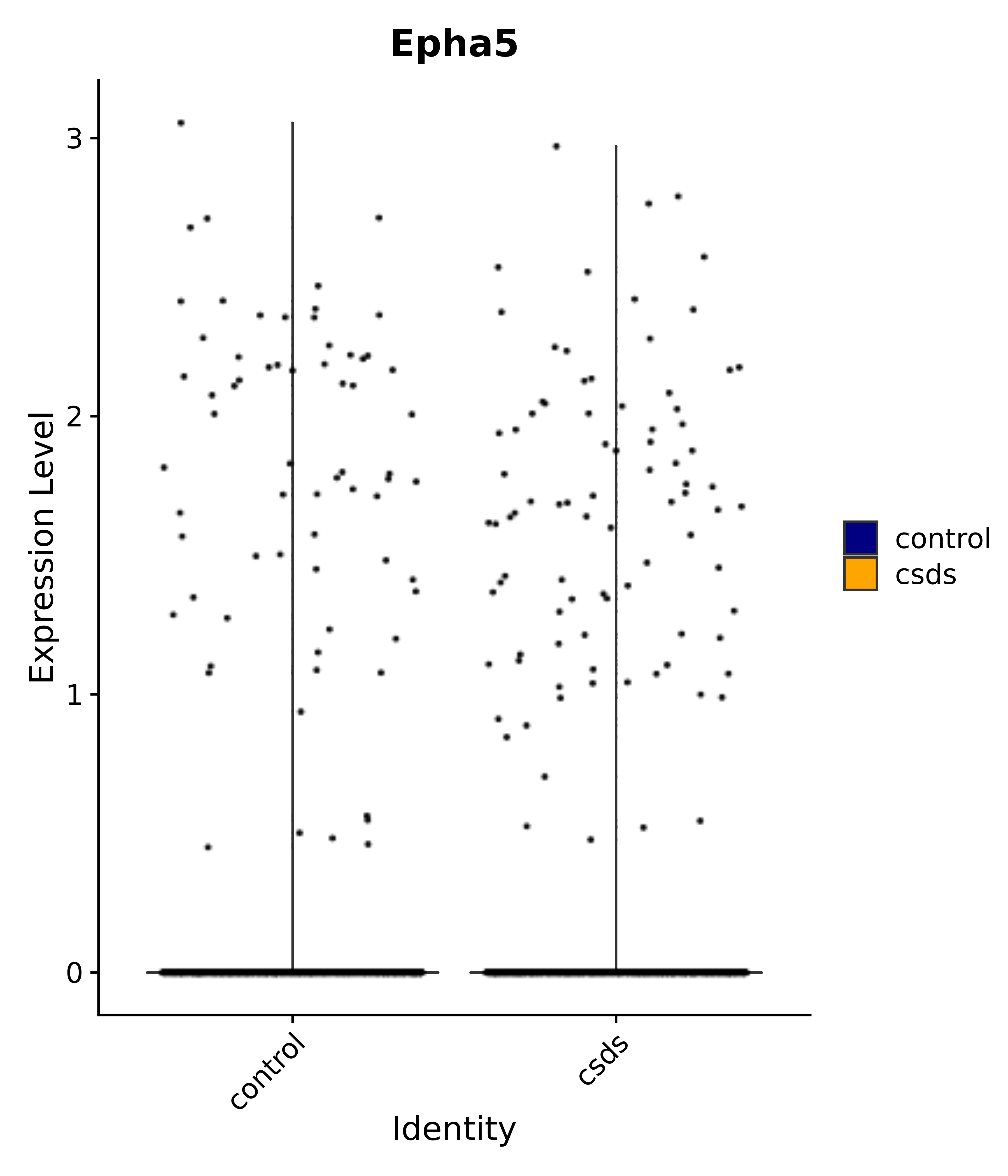
"Epha5",
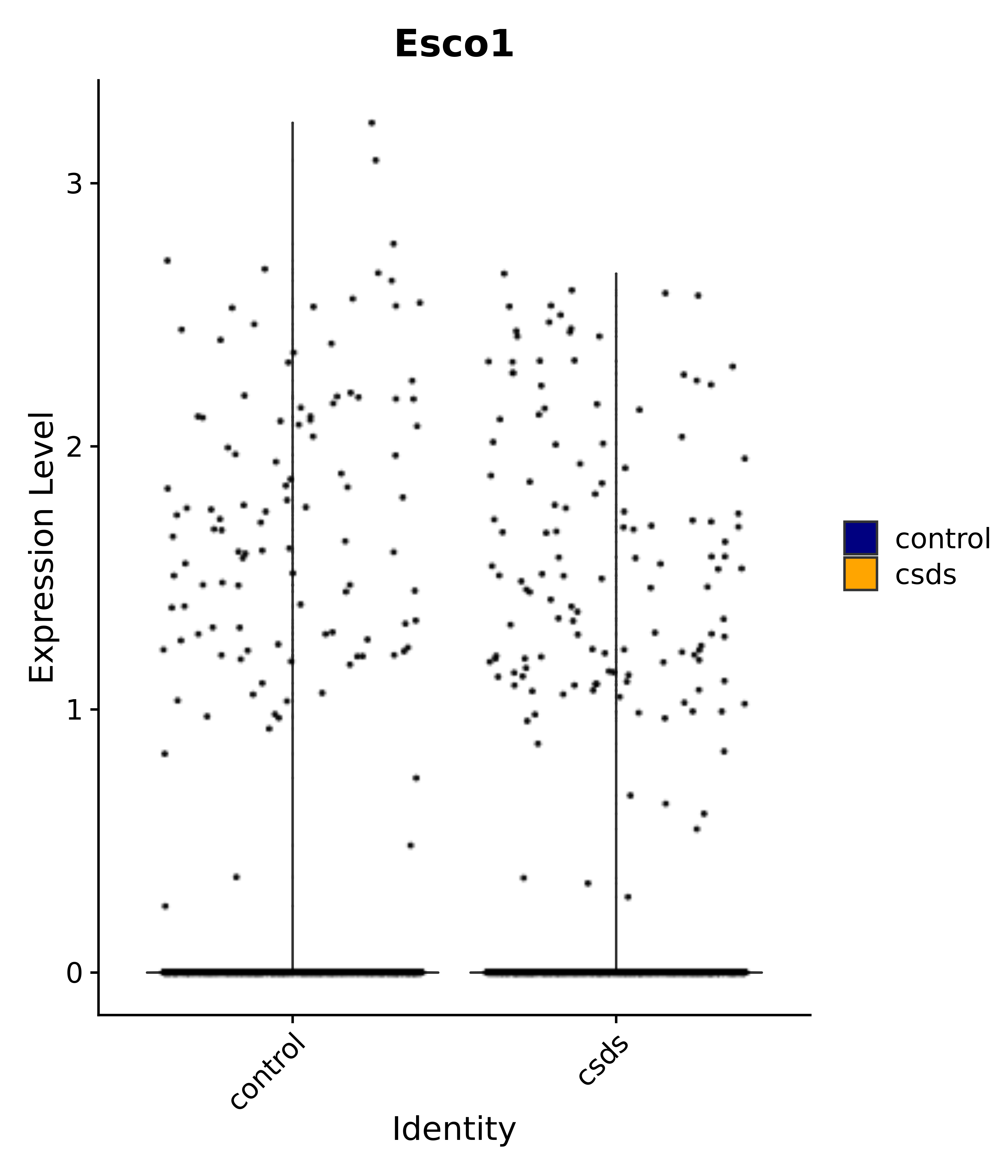
"Esco1",
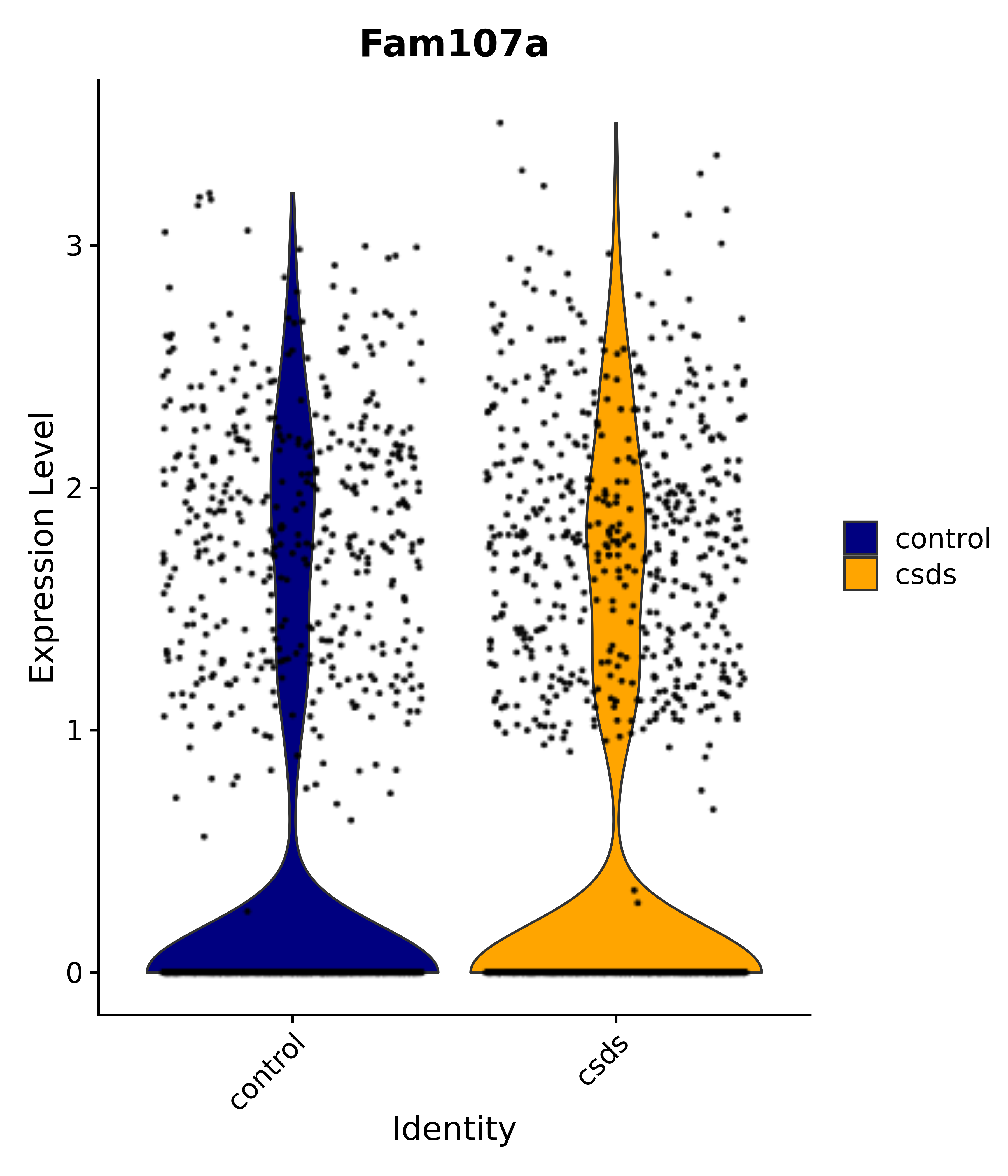
"Fam107a",
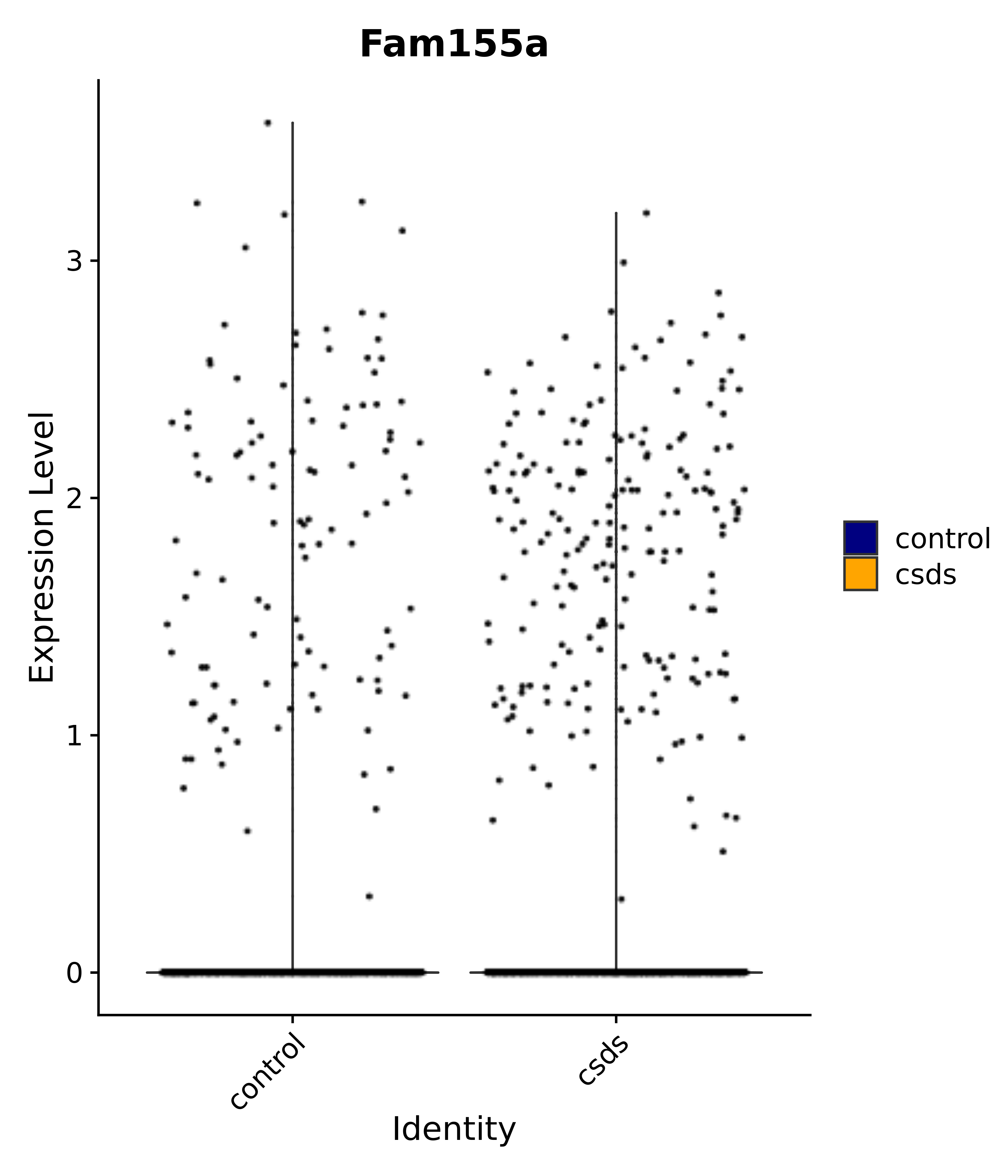
"Fam155a",
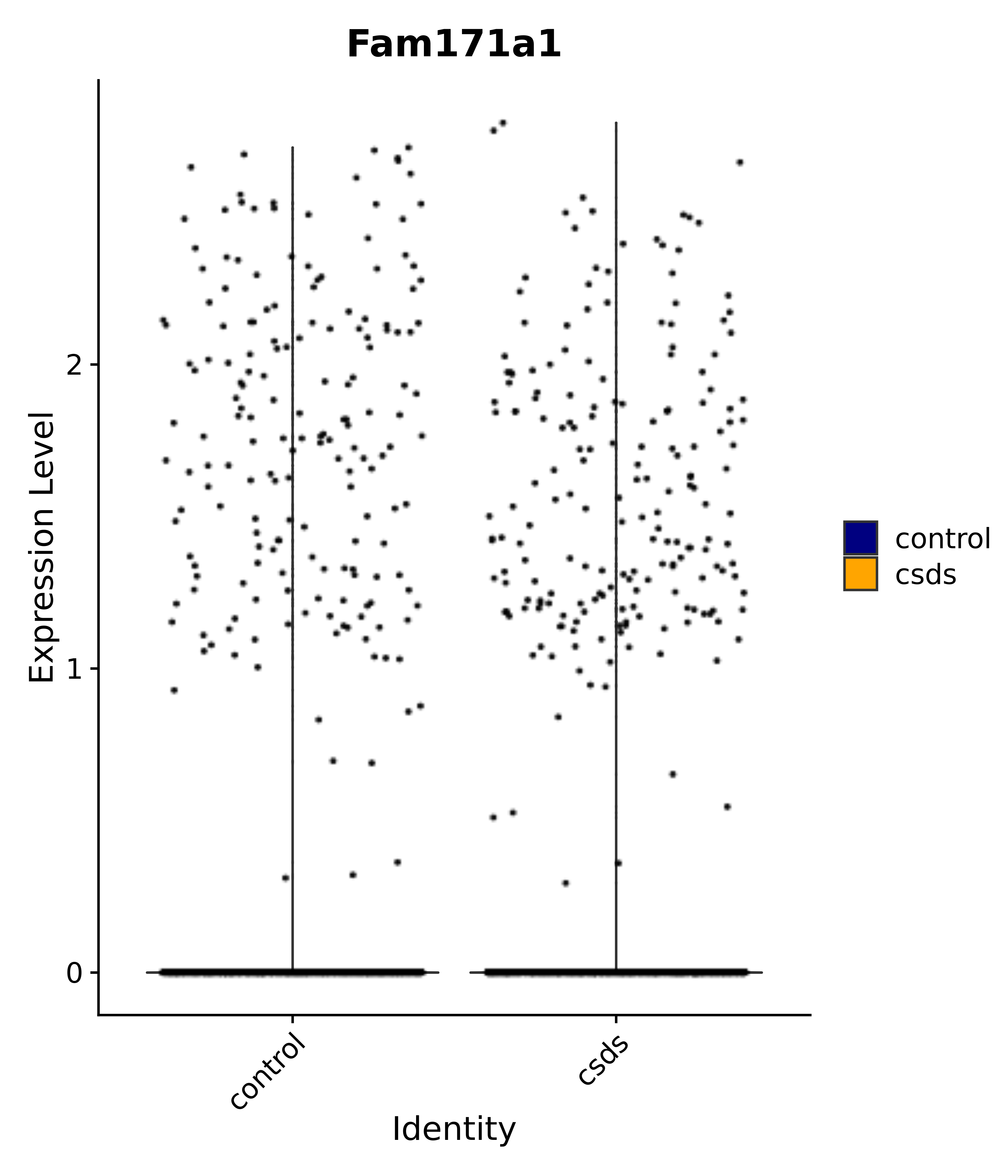
"Fam171a1",
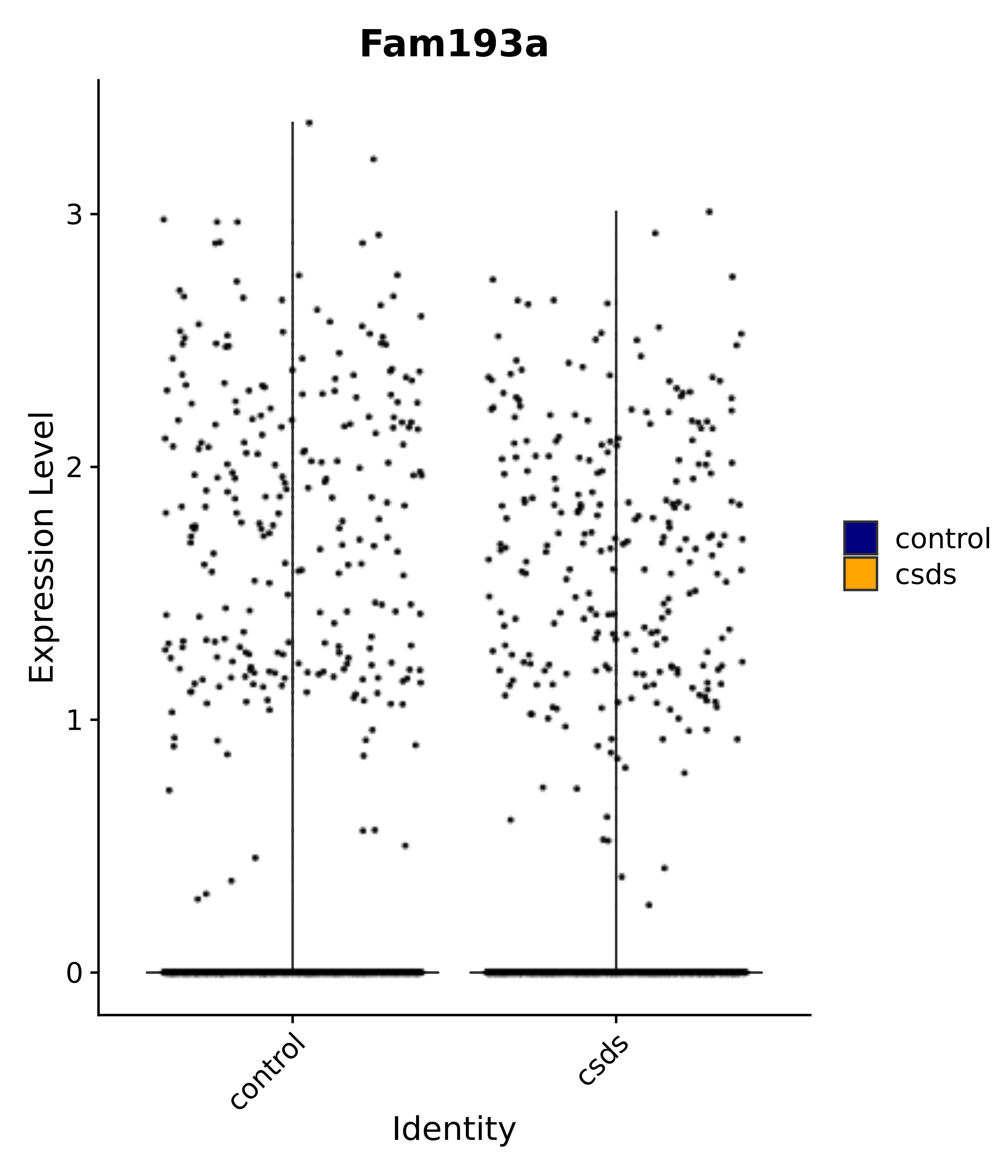
"Fam193a",
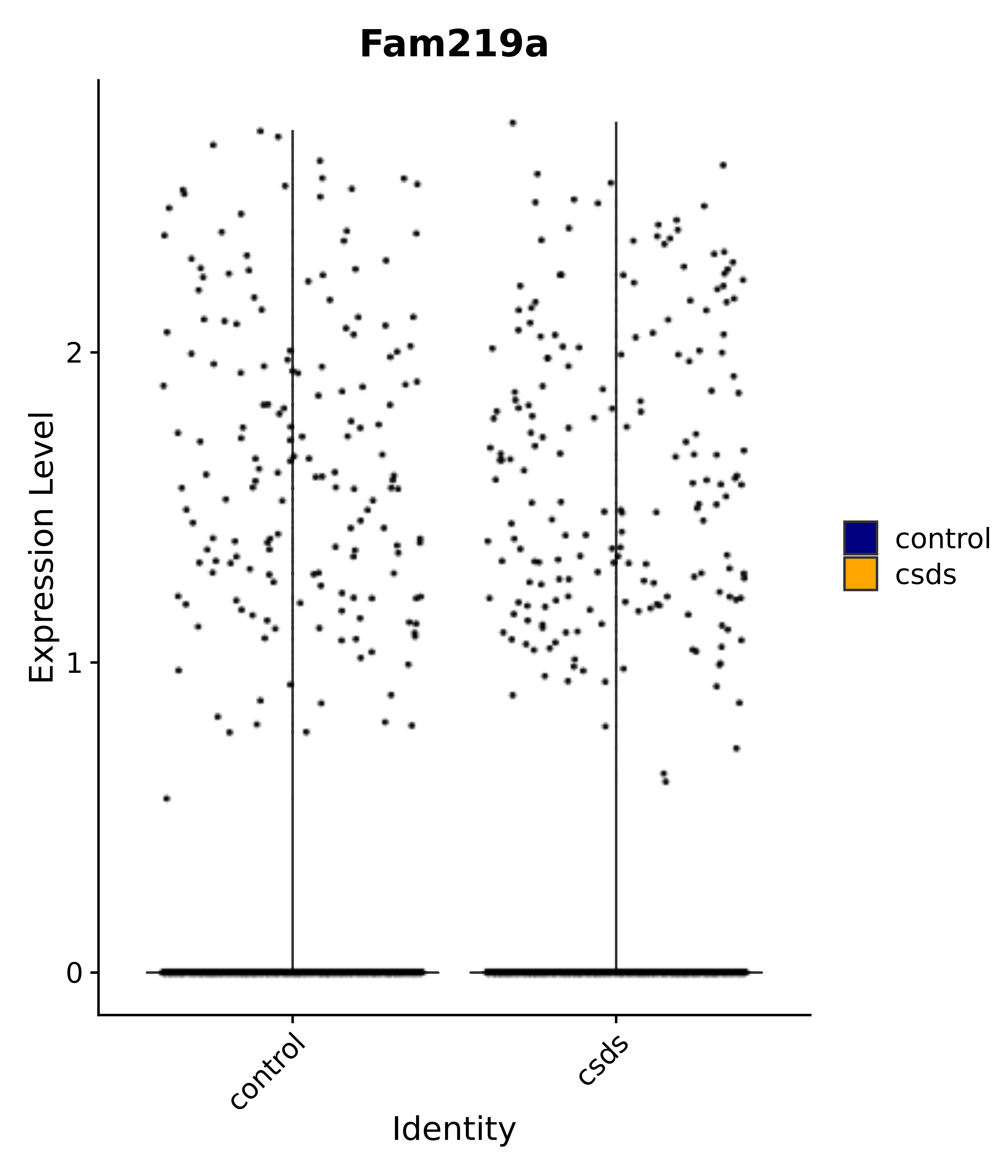
"Fam219a",
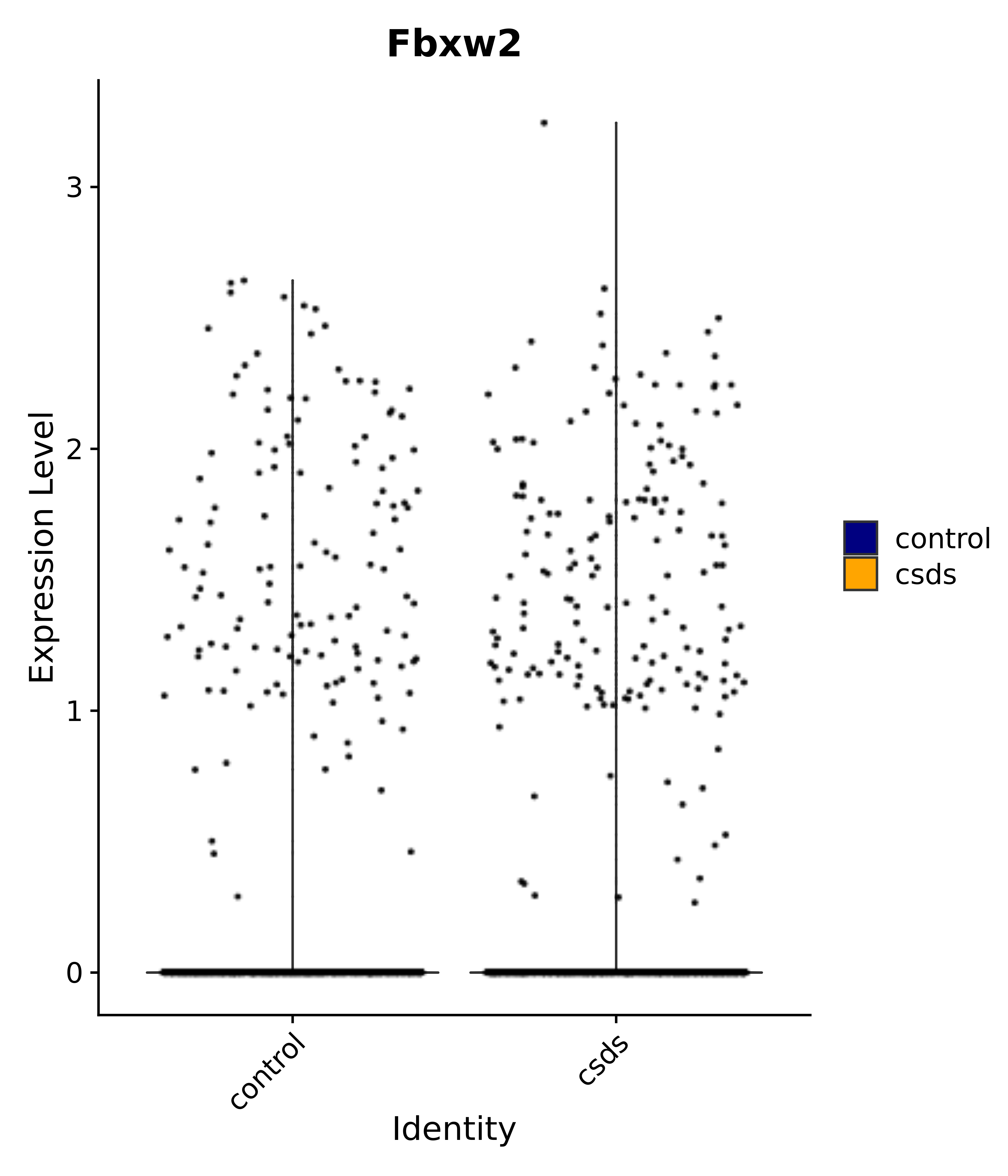
"Fbxw2",
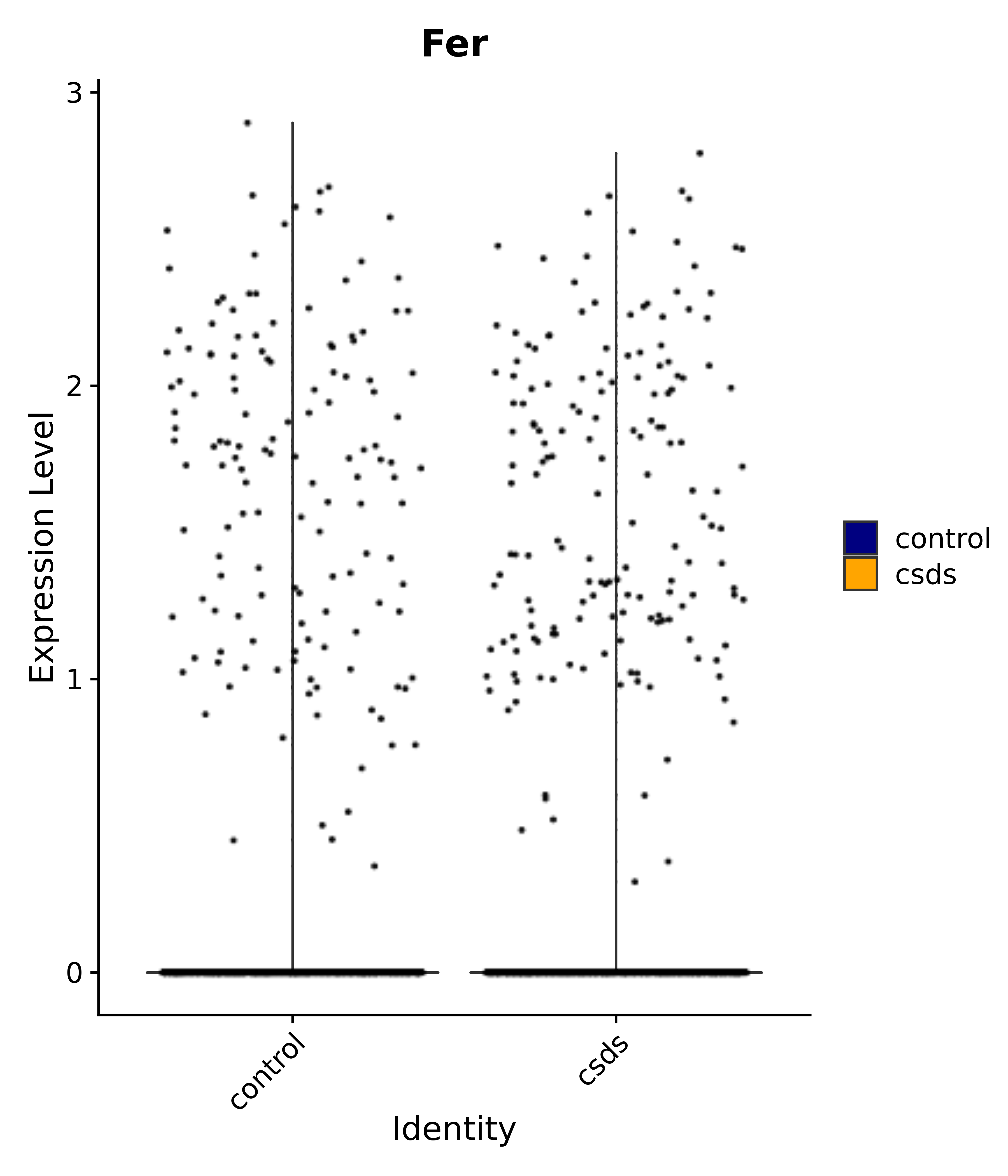
"Fer",
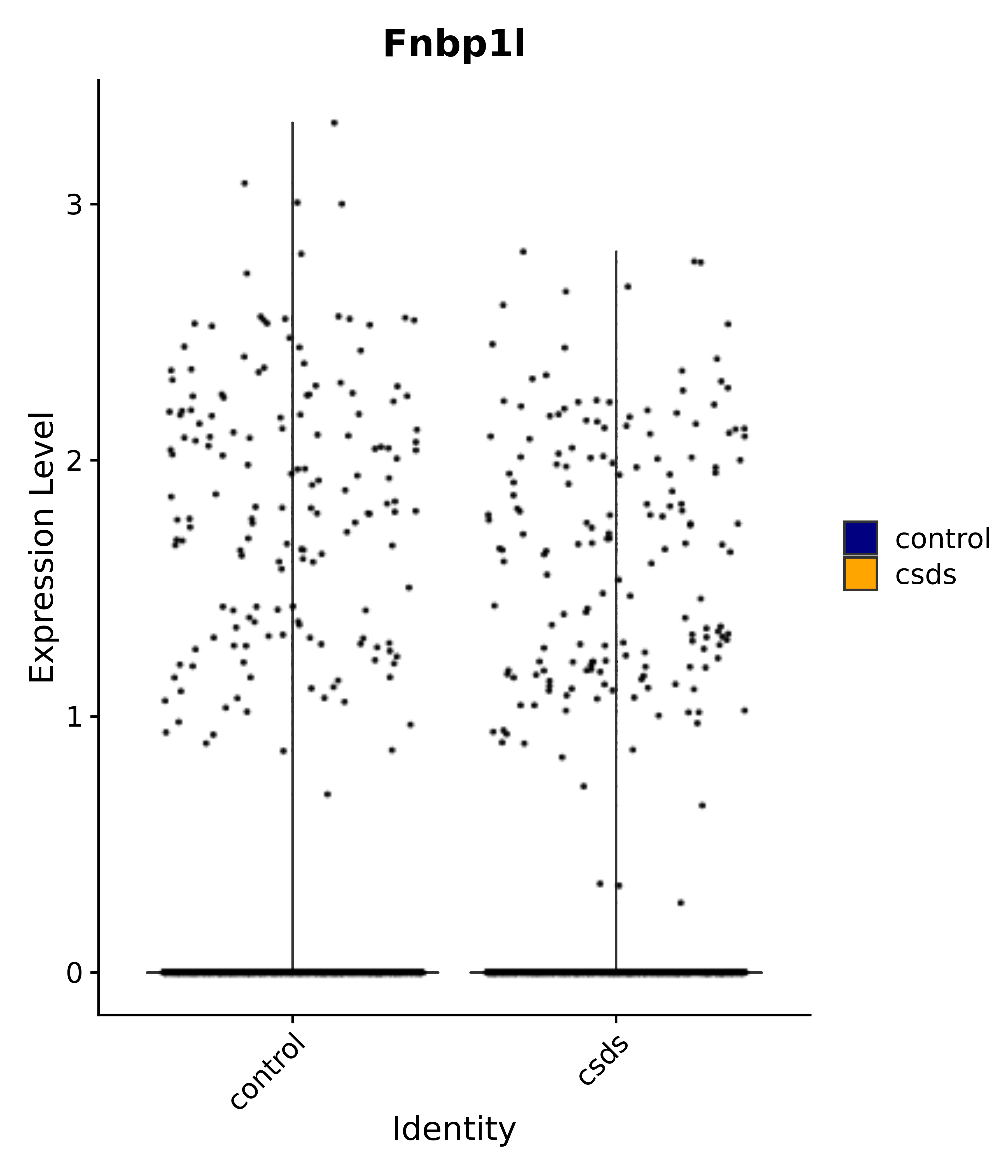
"Fnbp1l",
"Fos",
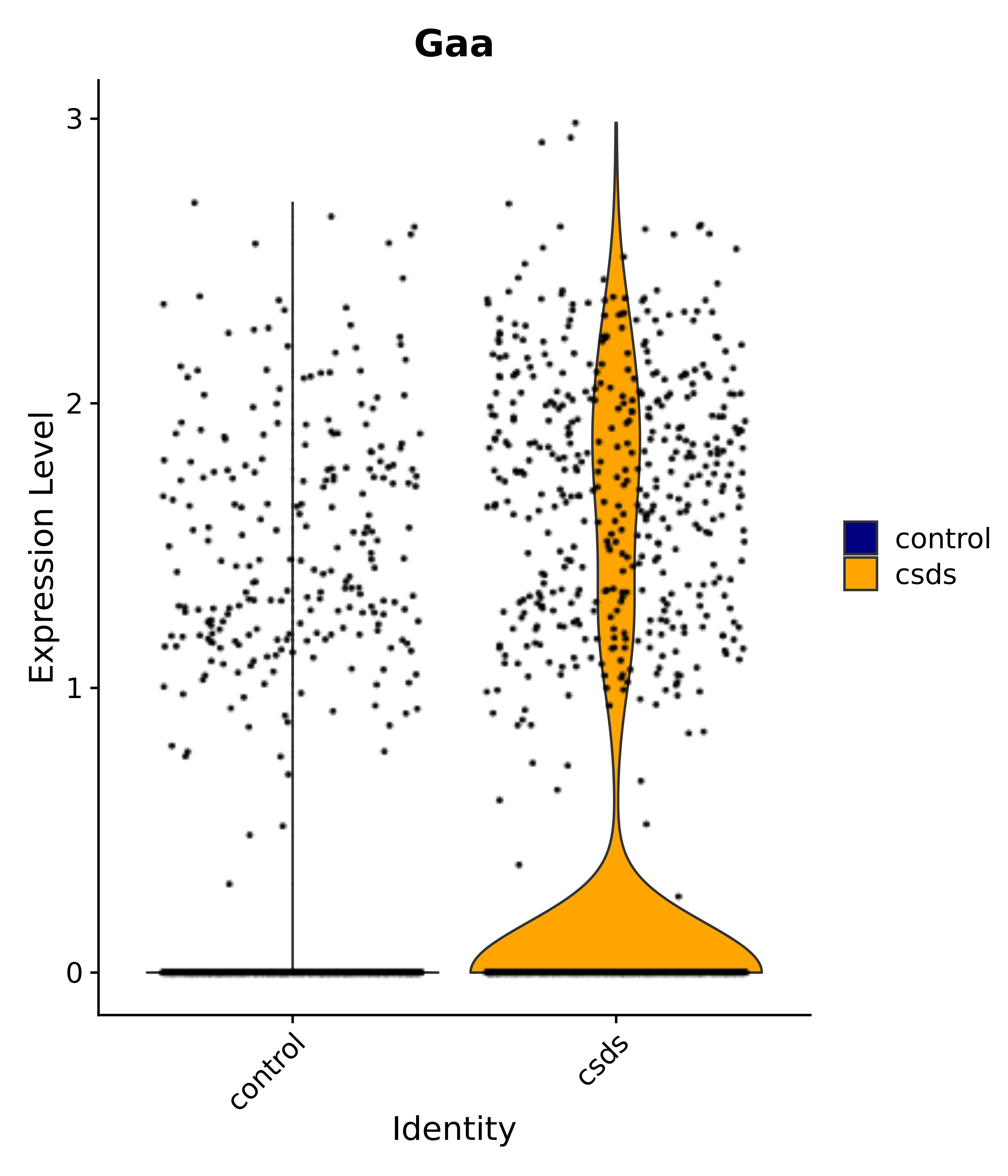
"Gaa",
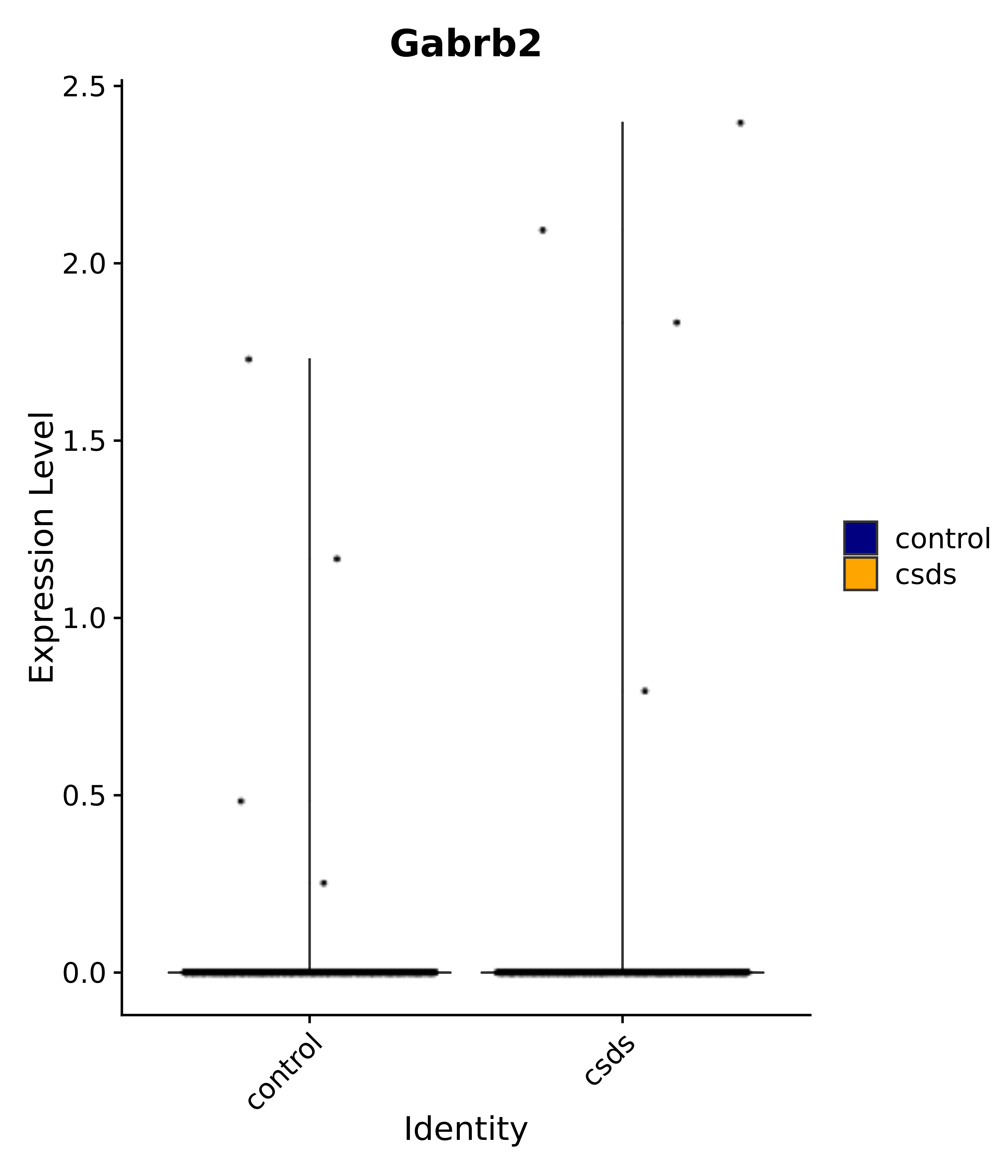
"Gabrb2",
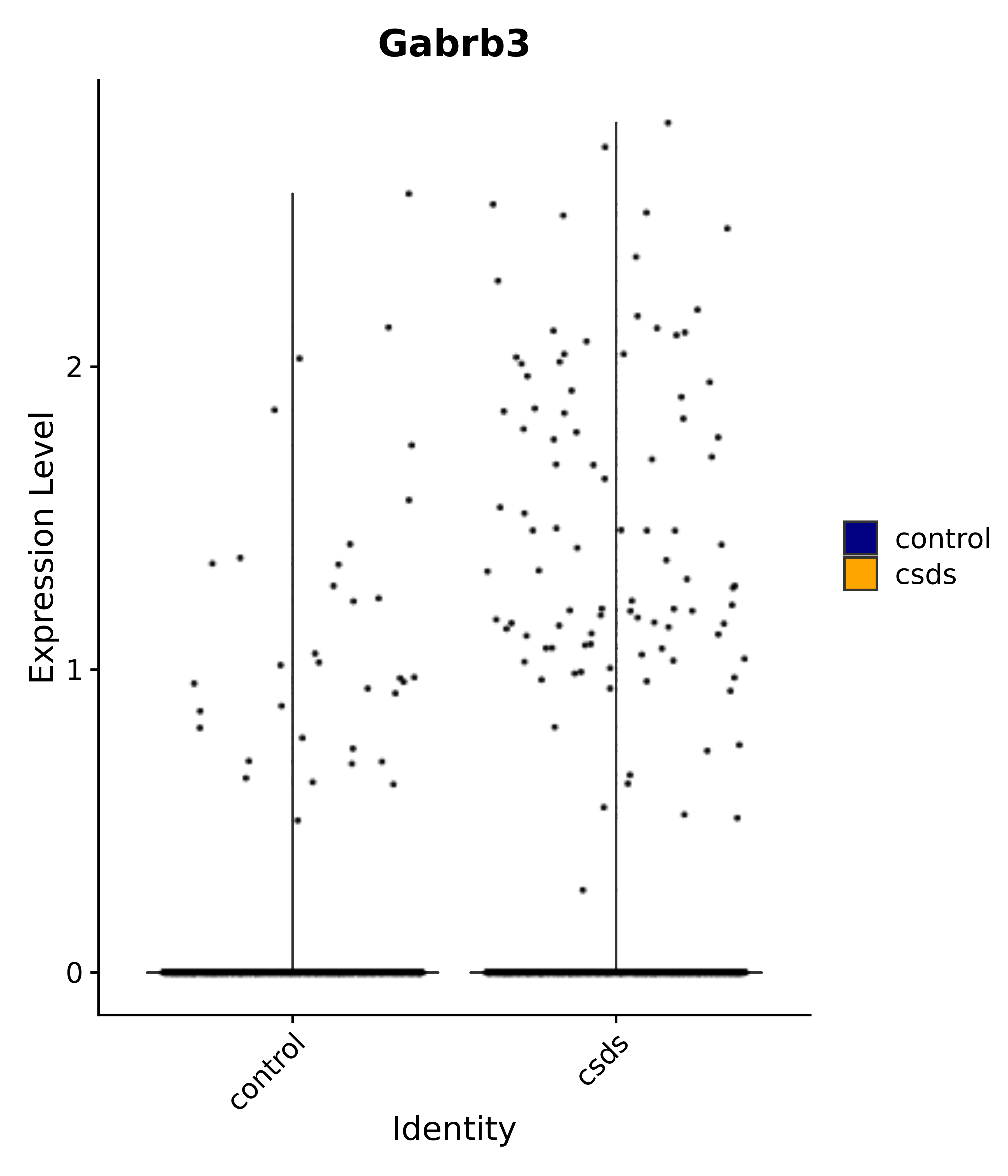
"Gabrb3",
"Gfap",
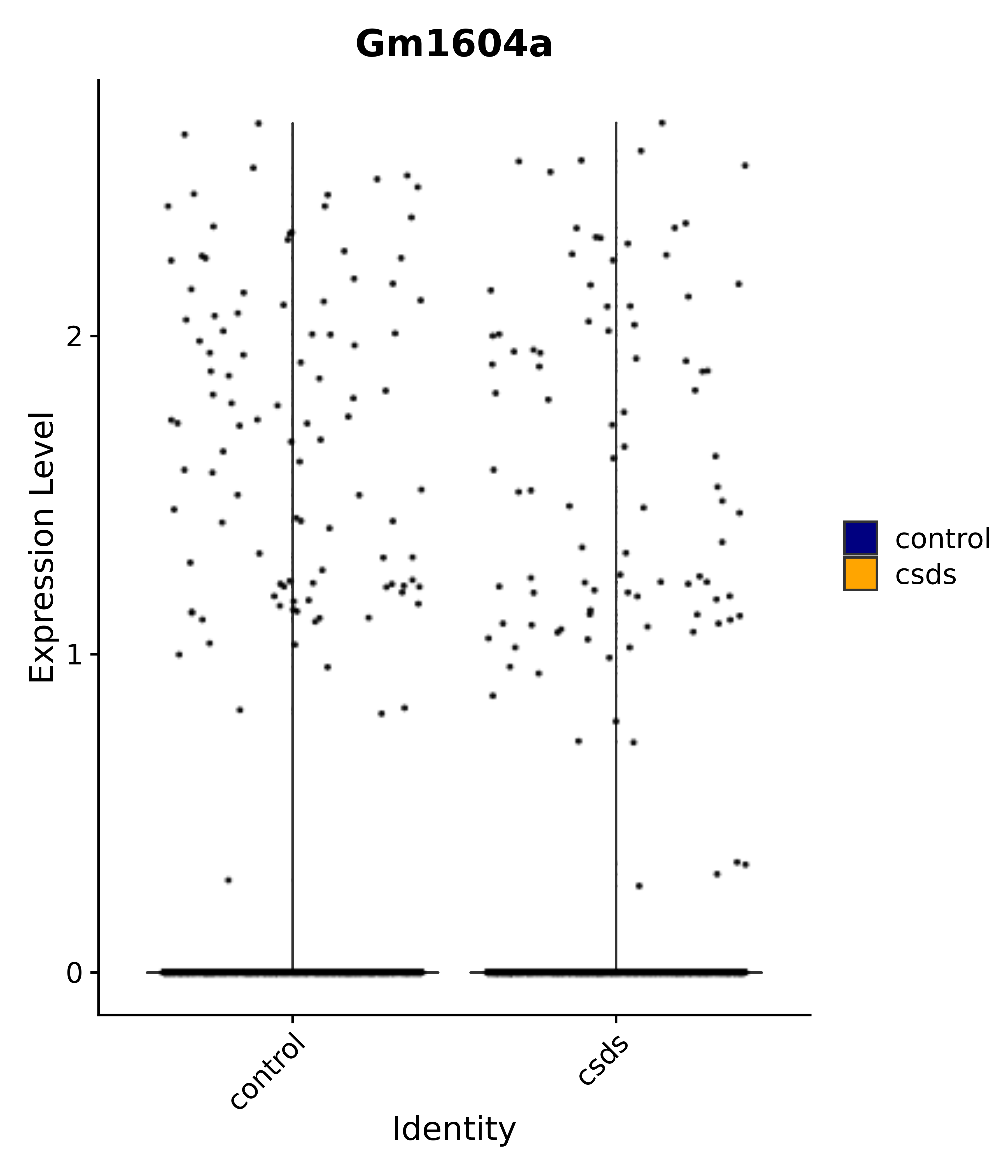
"Gm1604a",
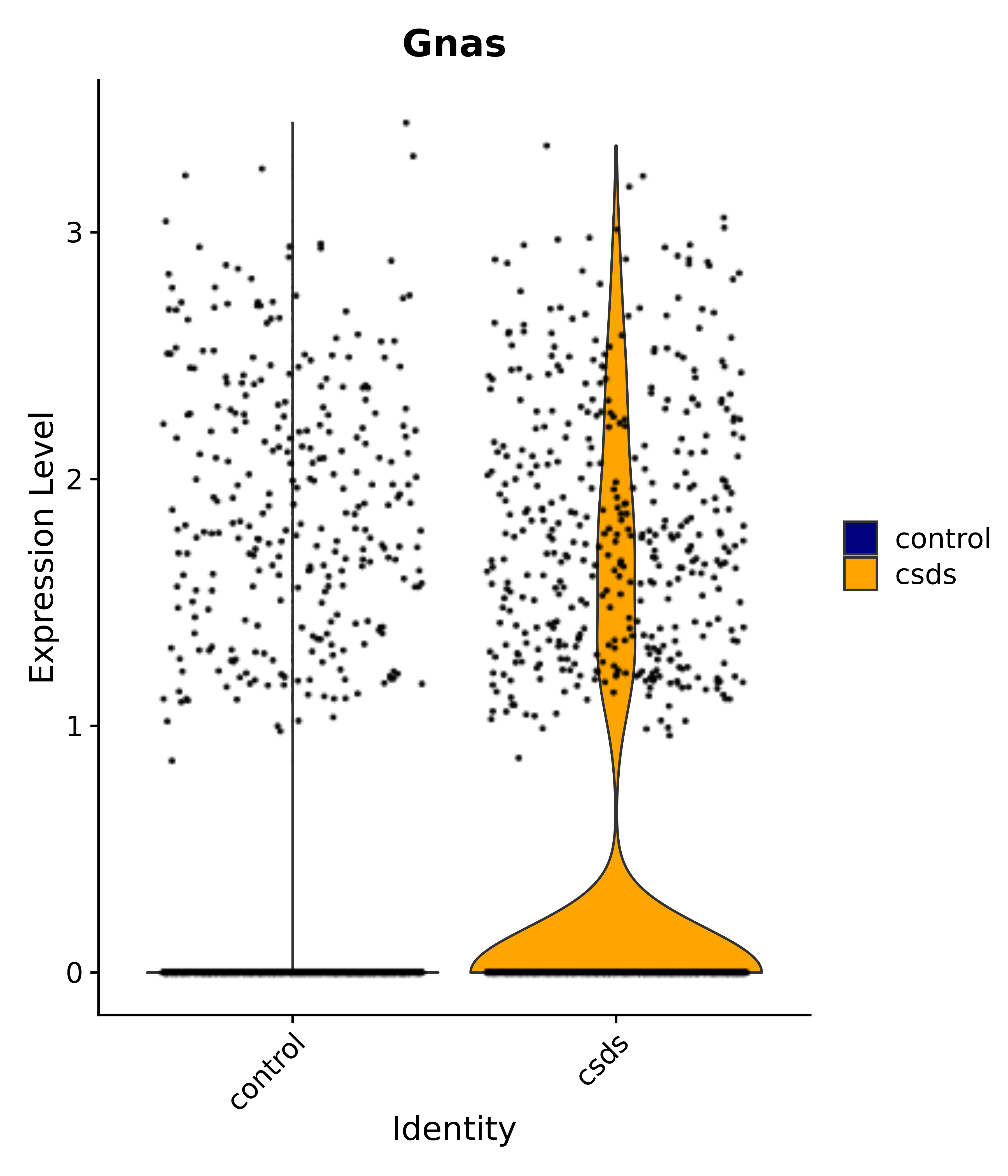
"Gnas",
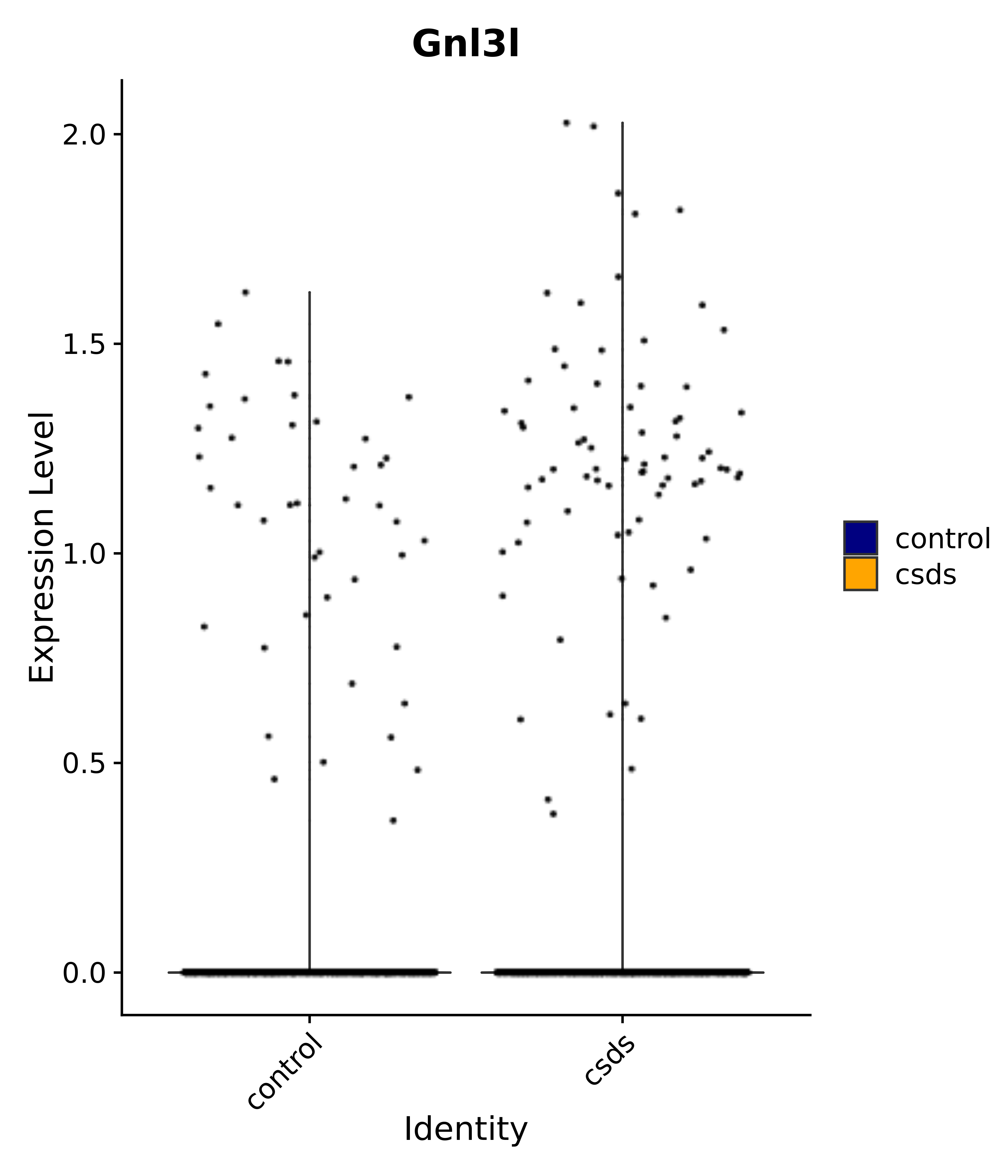
"Gnl3l",
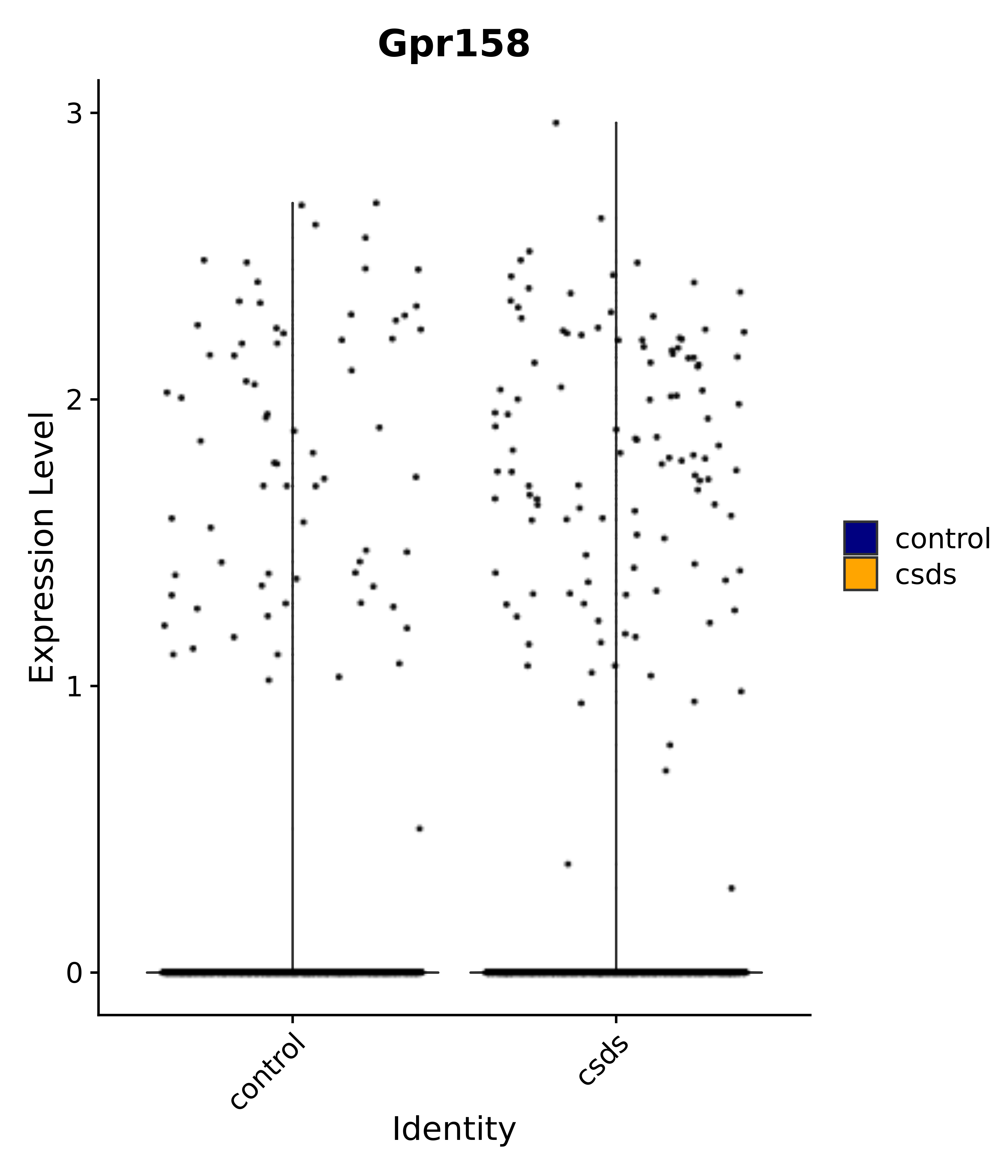
"Gpr158",
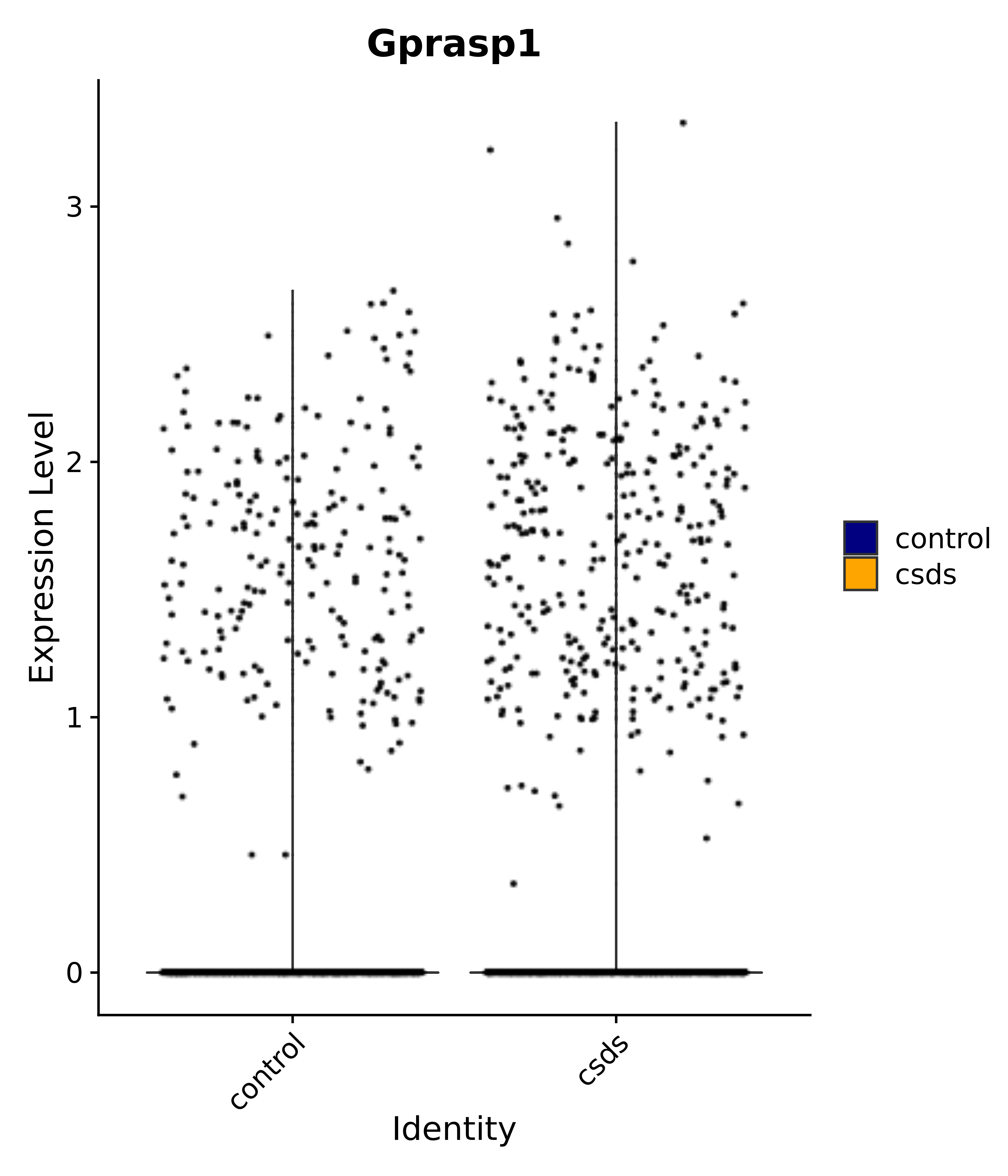
"Gprasp1",
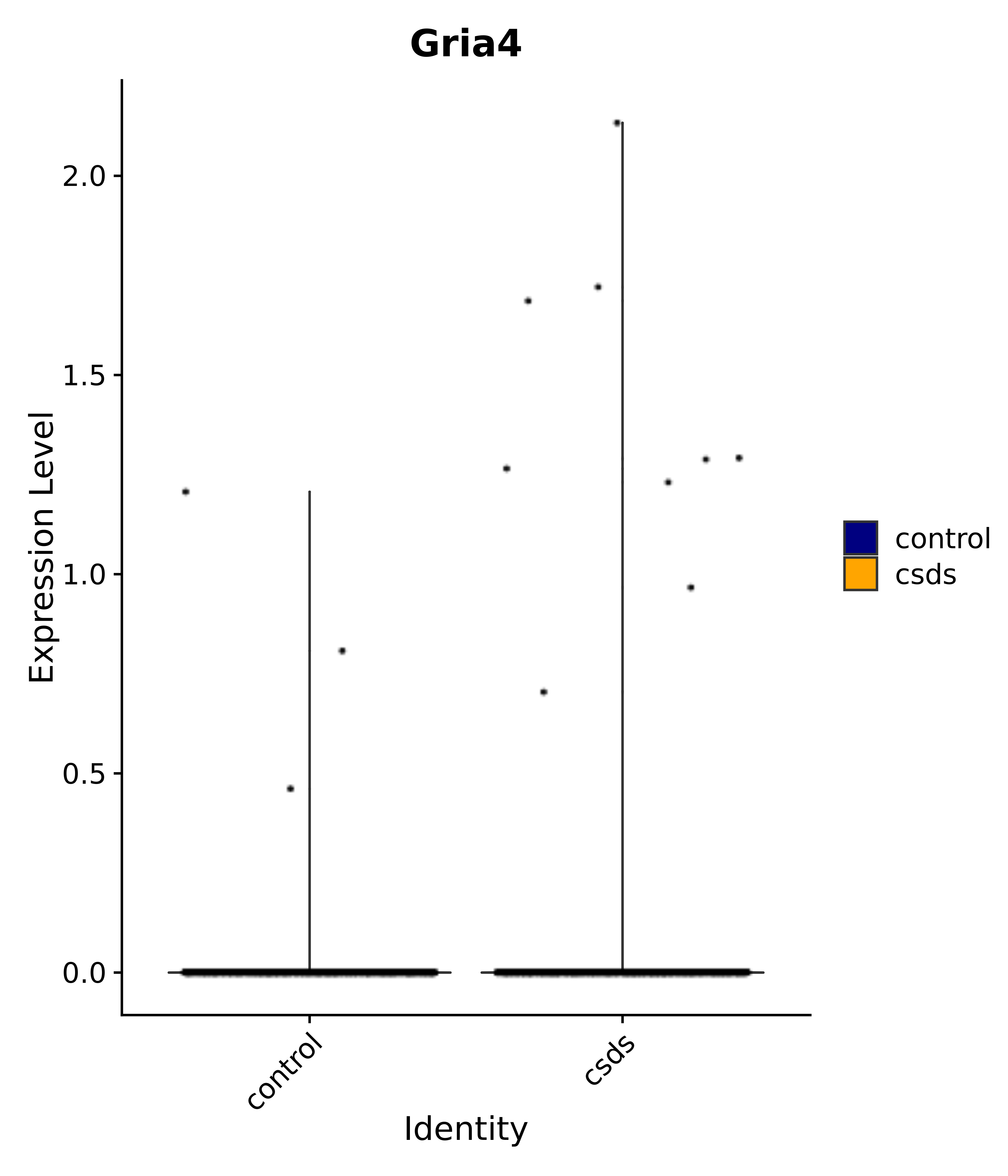
"Gria4",
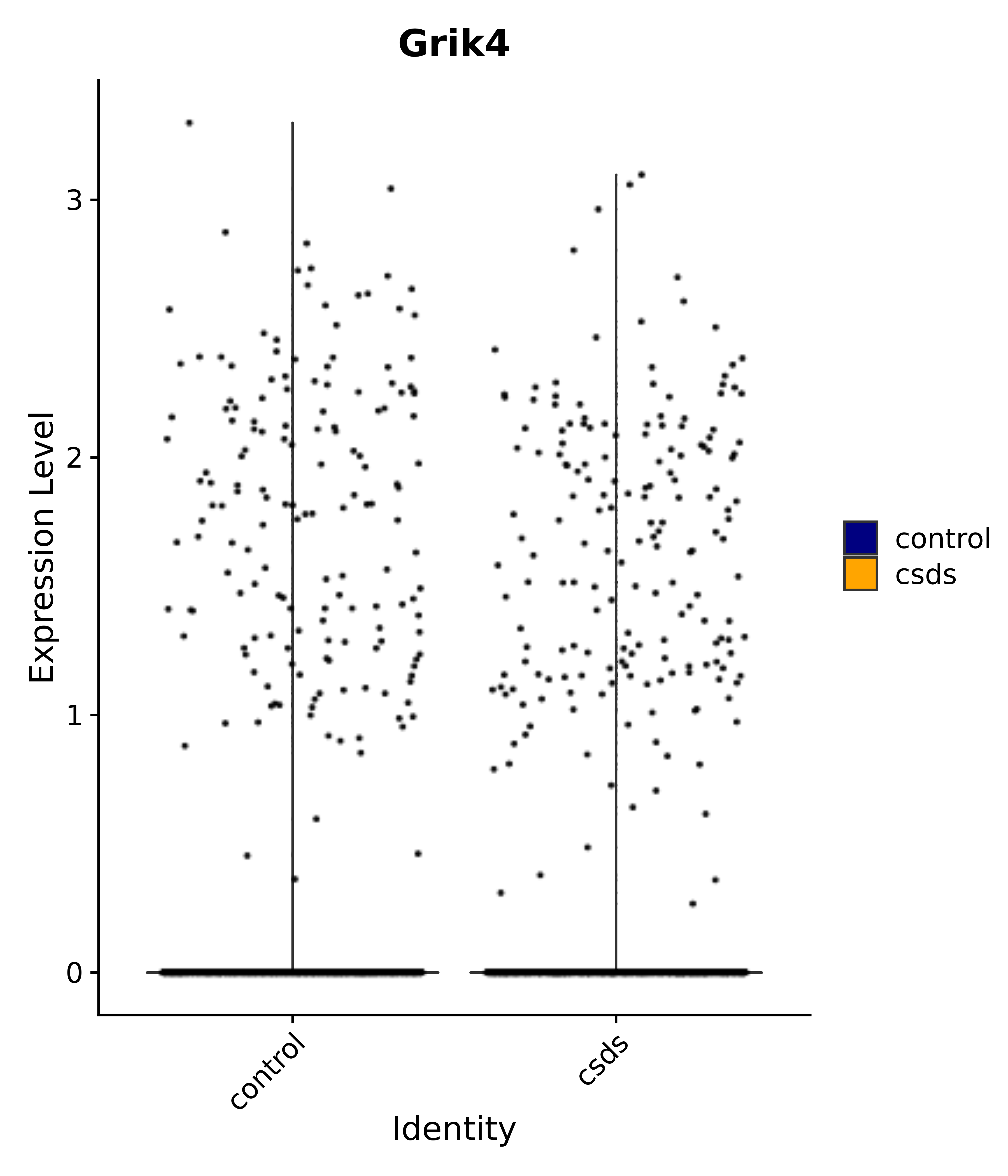
"Grik4",
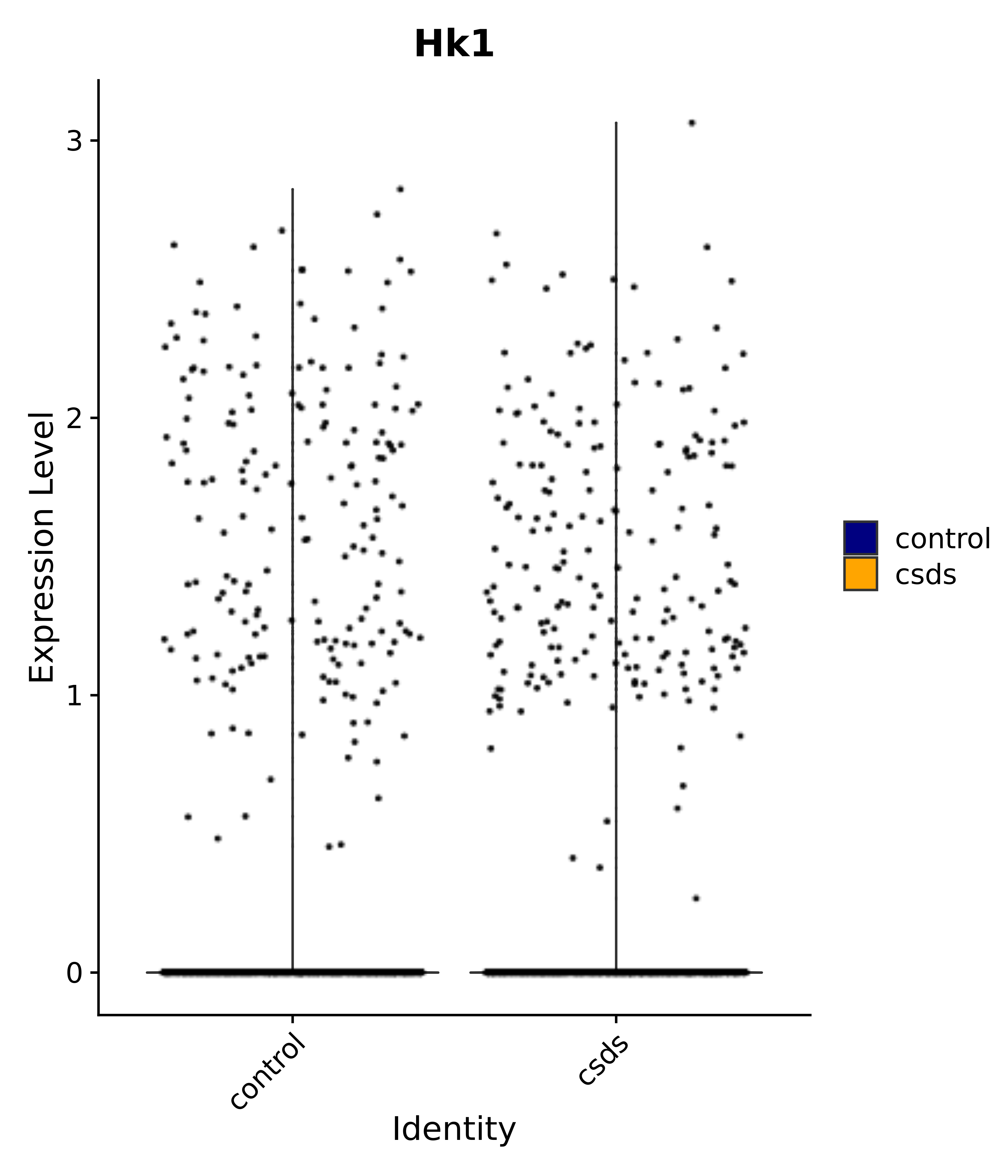
"Hk1",
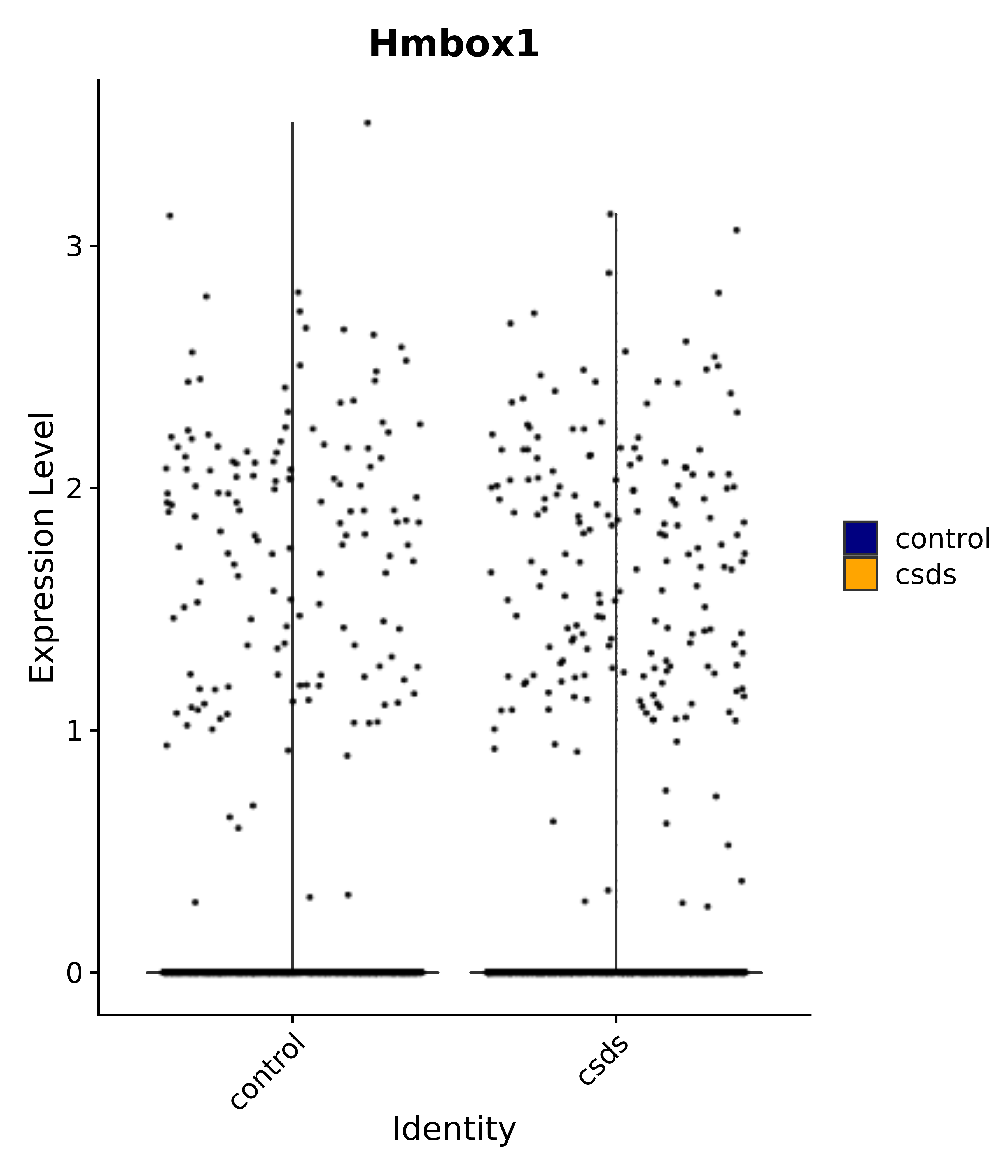
"Hmbox1",
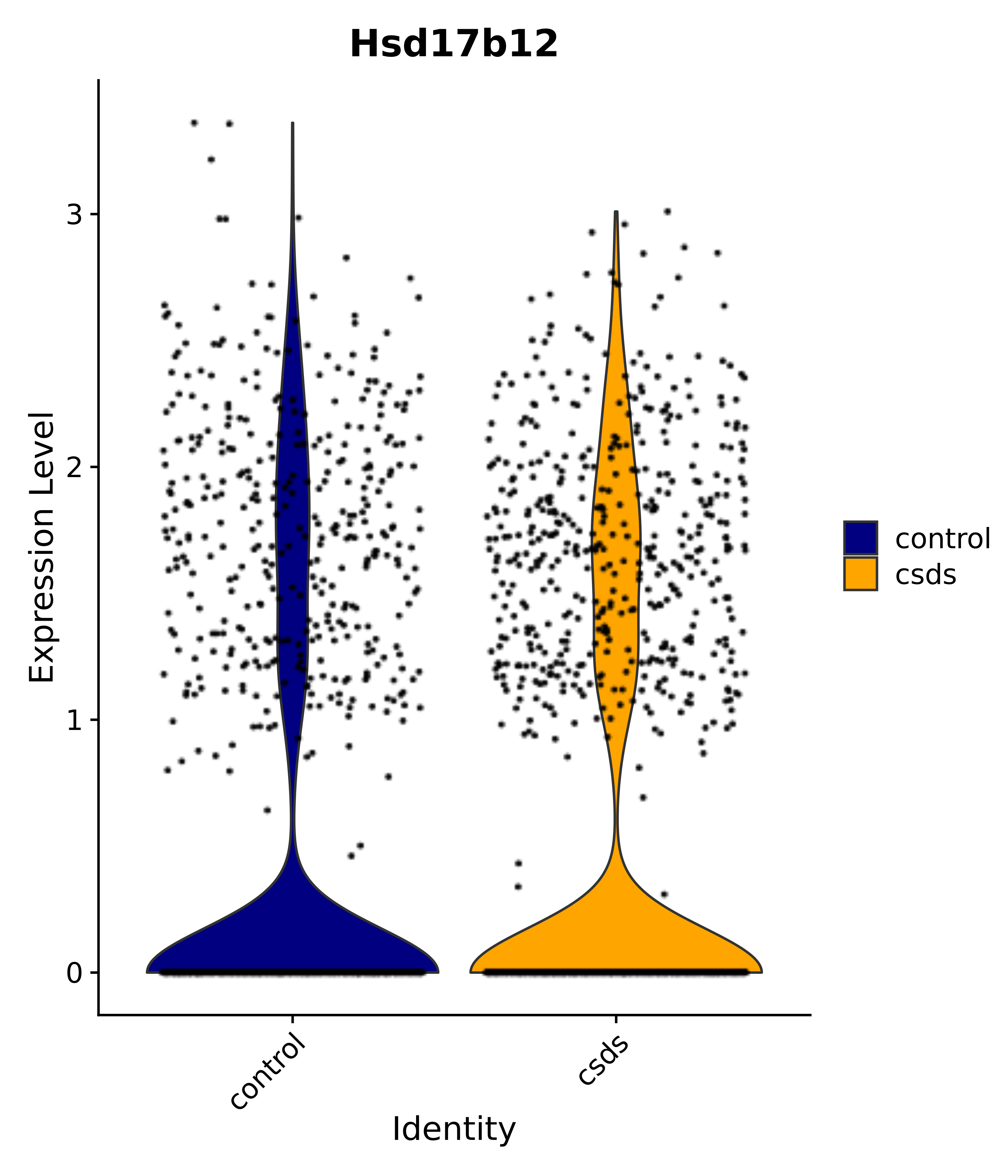
"Hsd17b12",
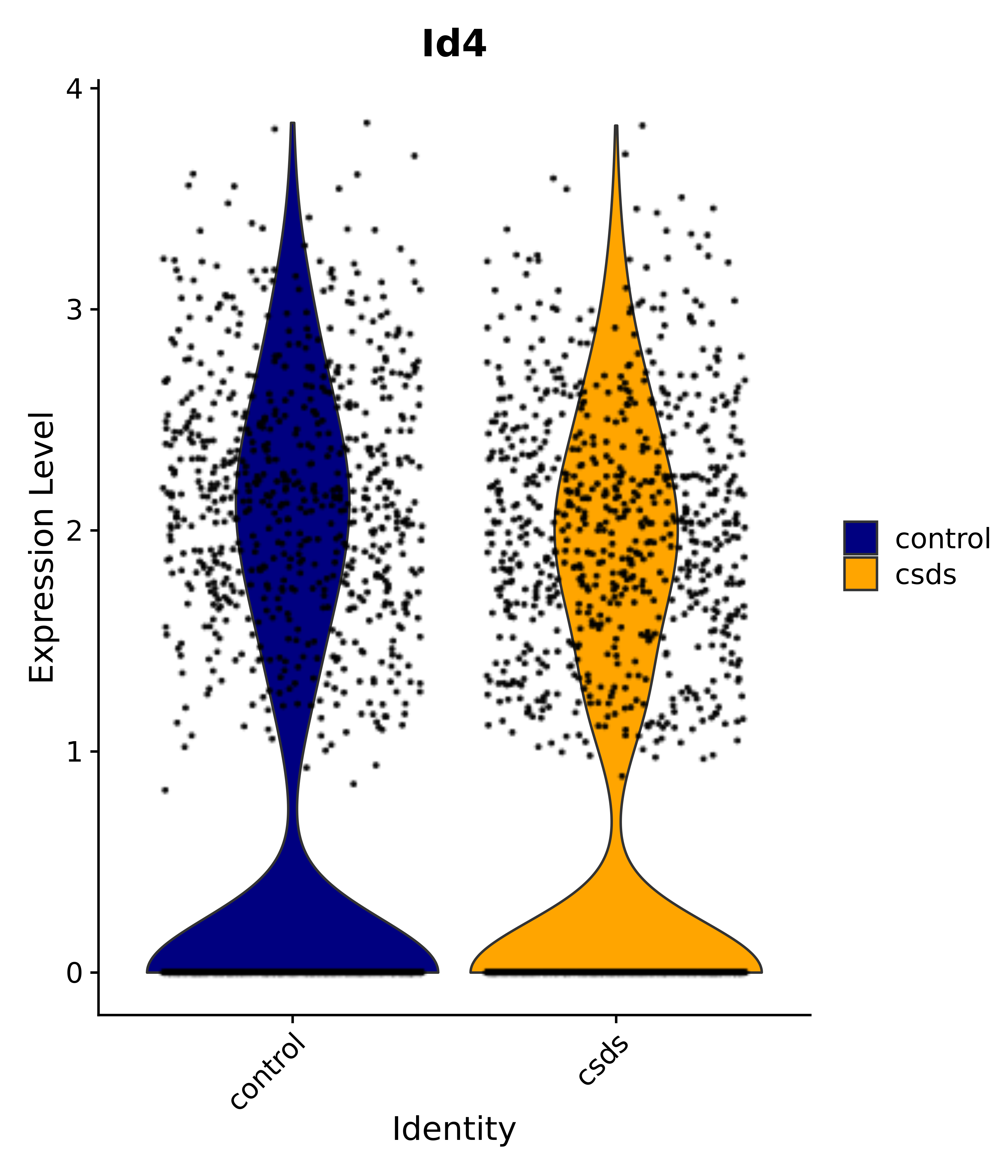
"Id4",
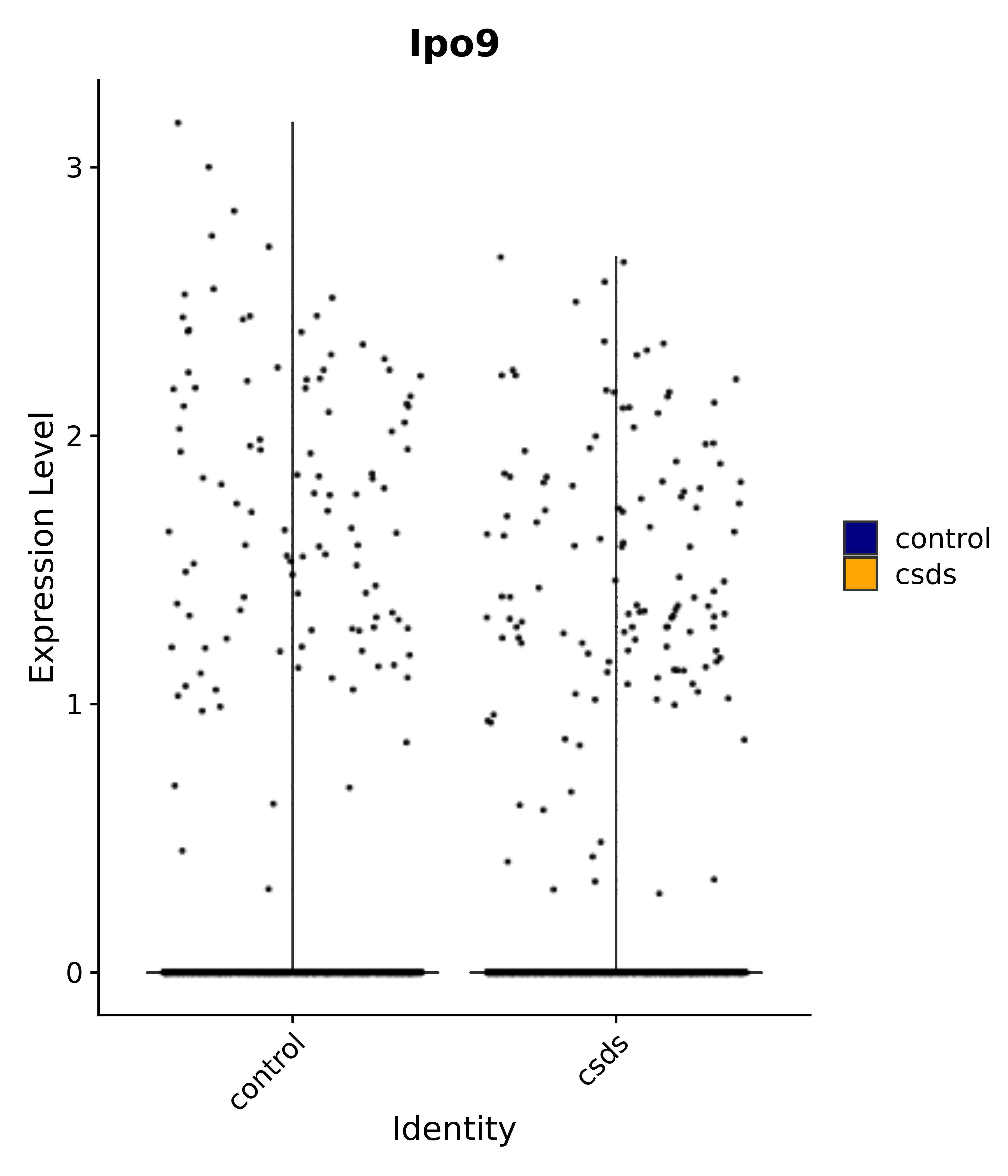
"Ipo9",
"Jun",
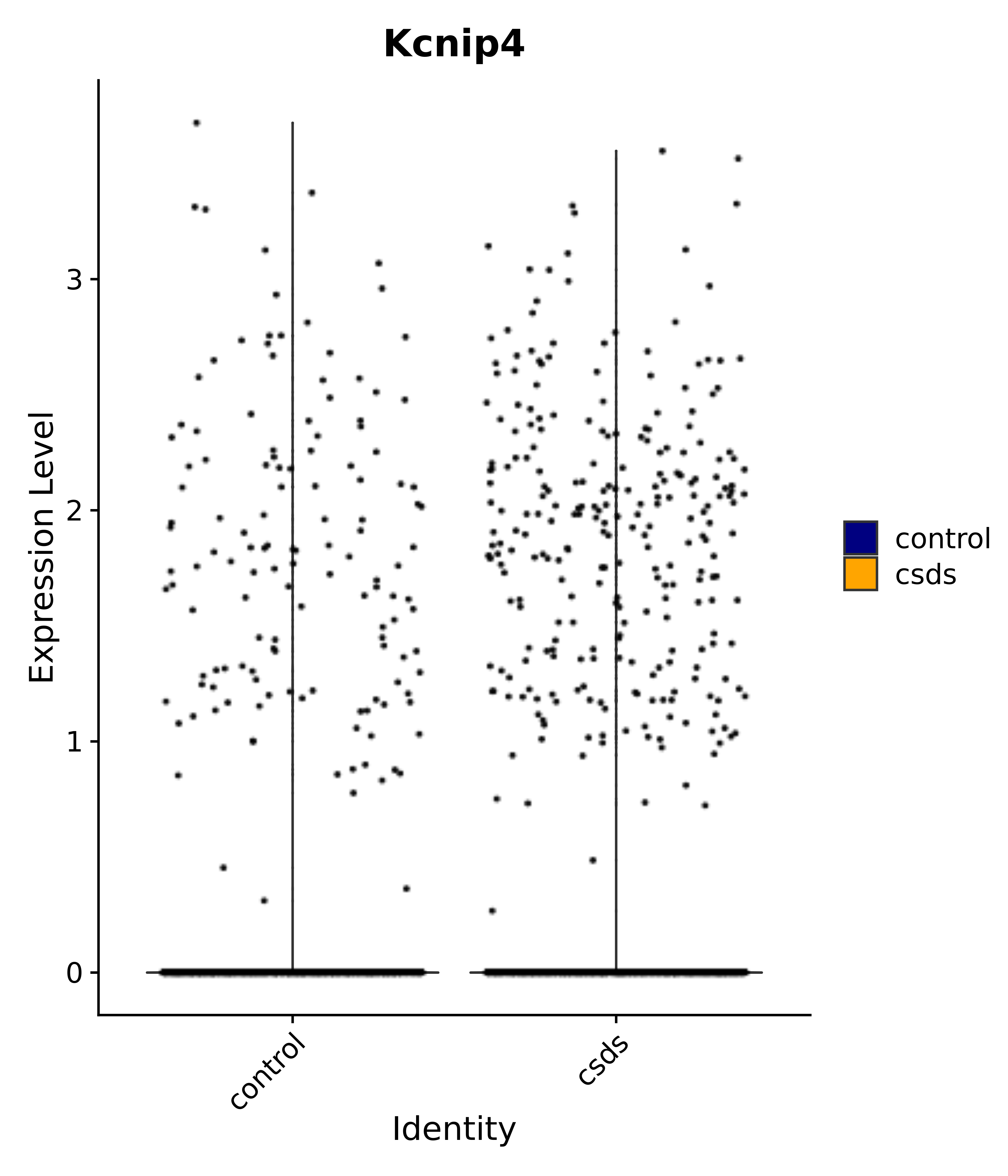
"Kcnip4",
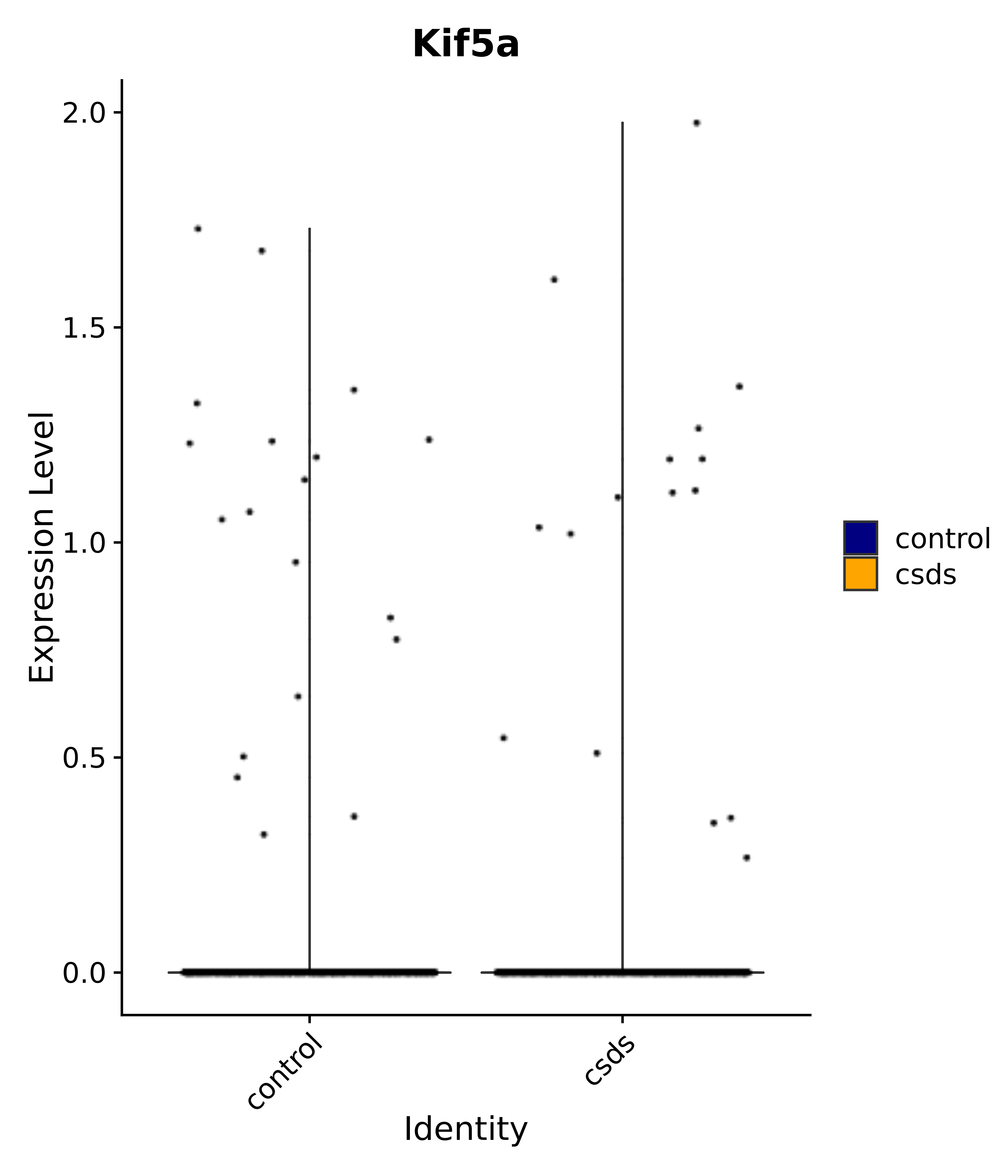
"Kif5a",
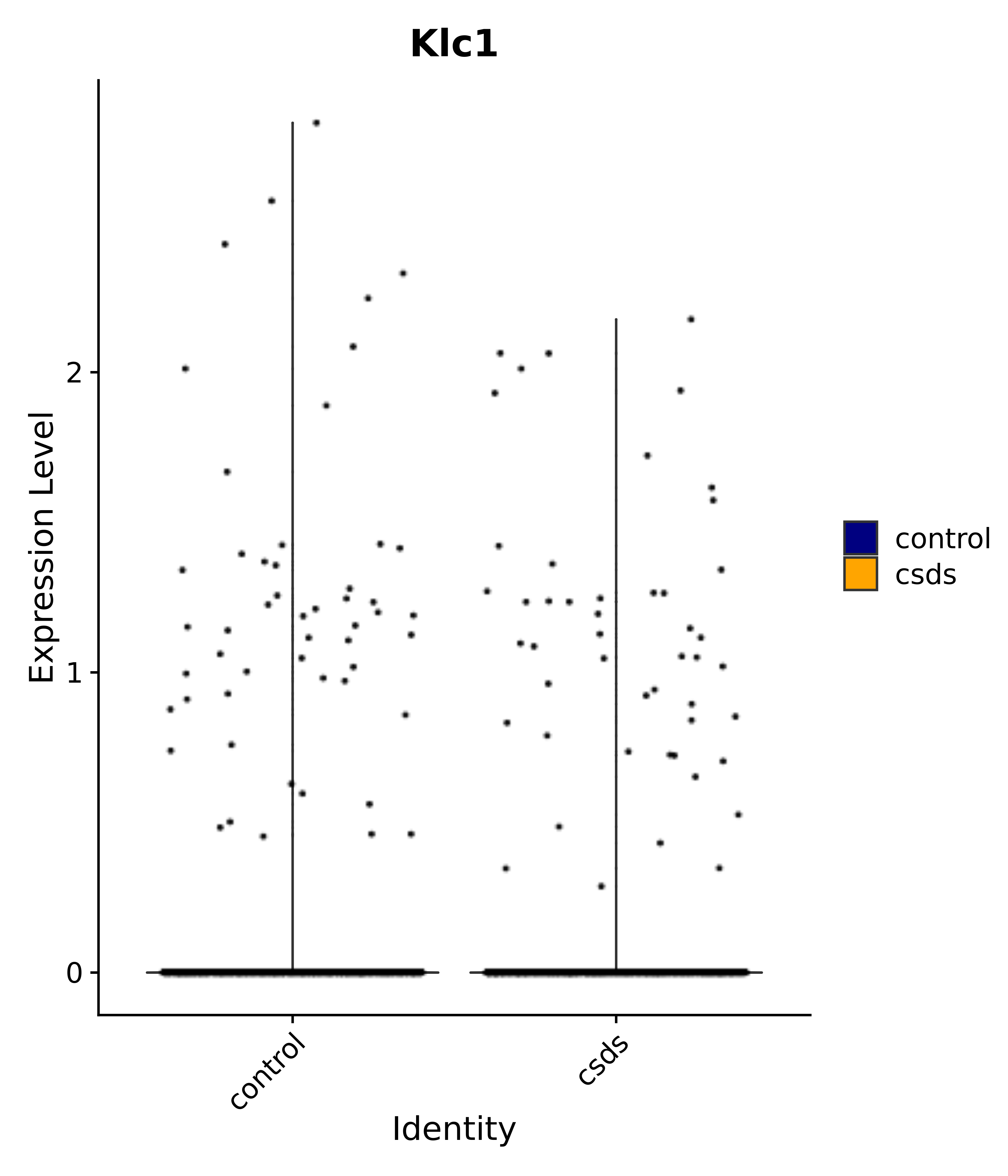
"Klc1",
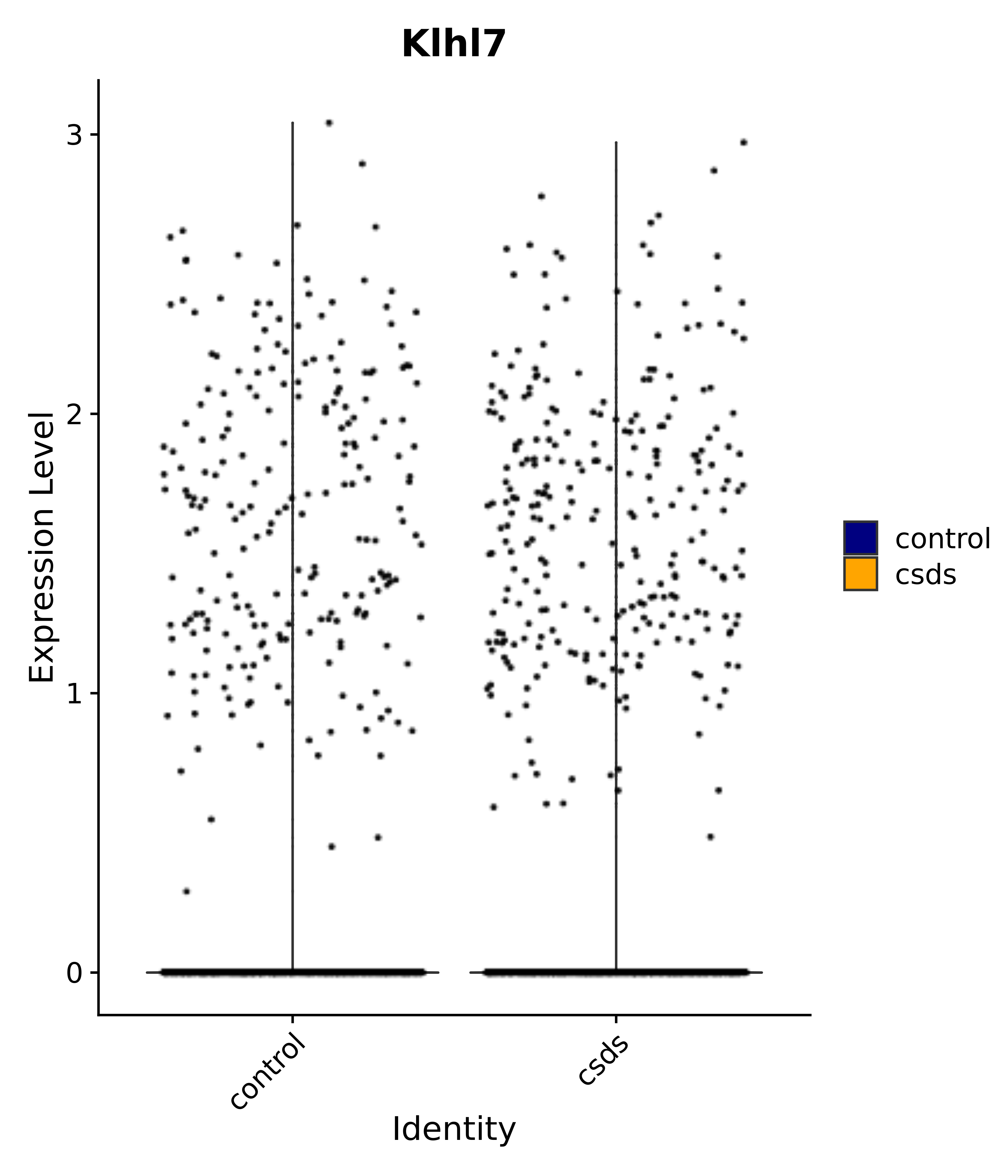
"Klhl7",
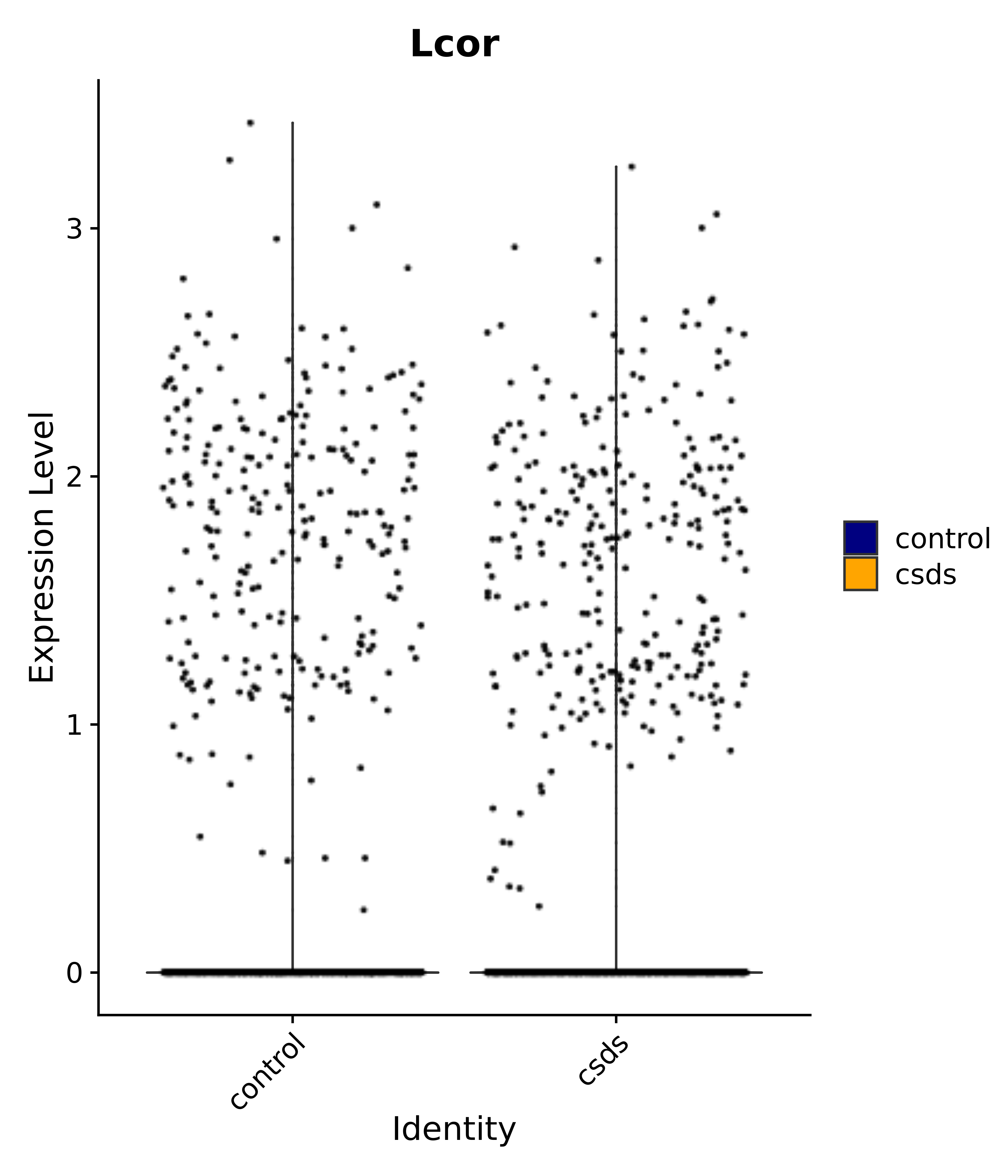
"Lcor",
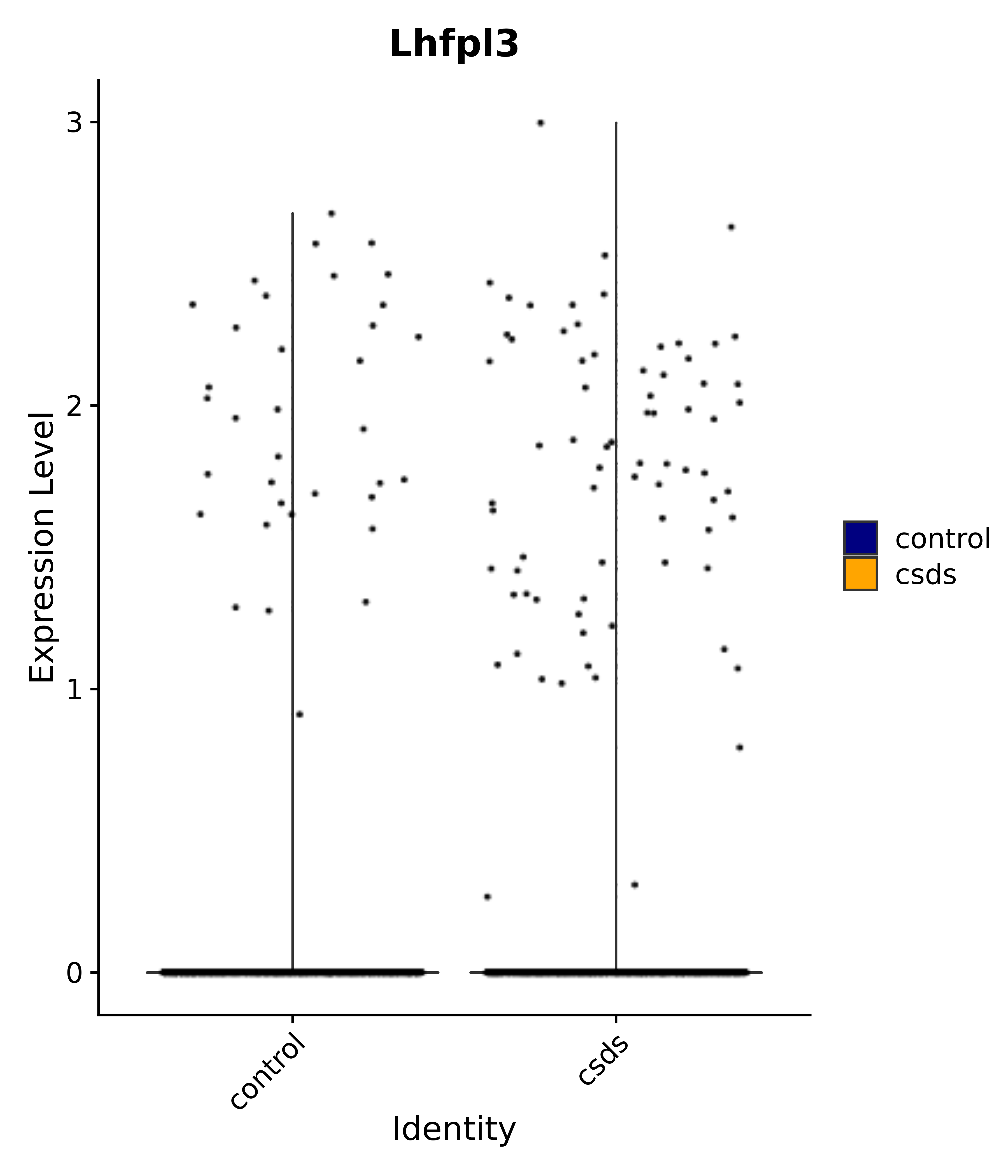
"Lhfpl3",
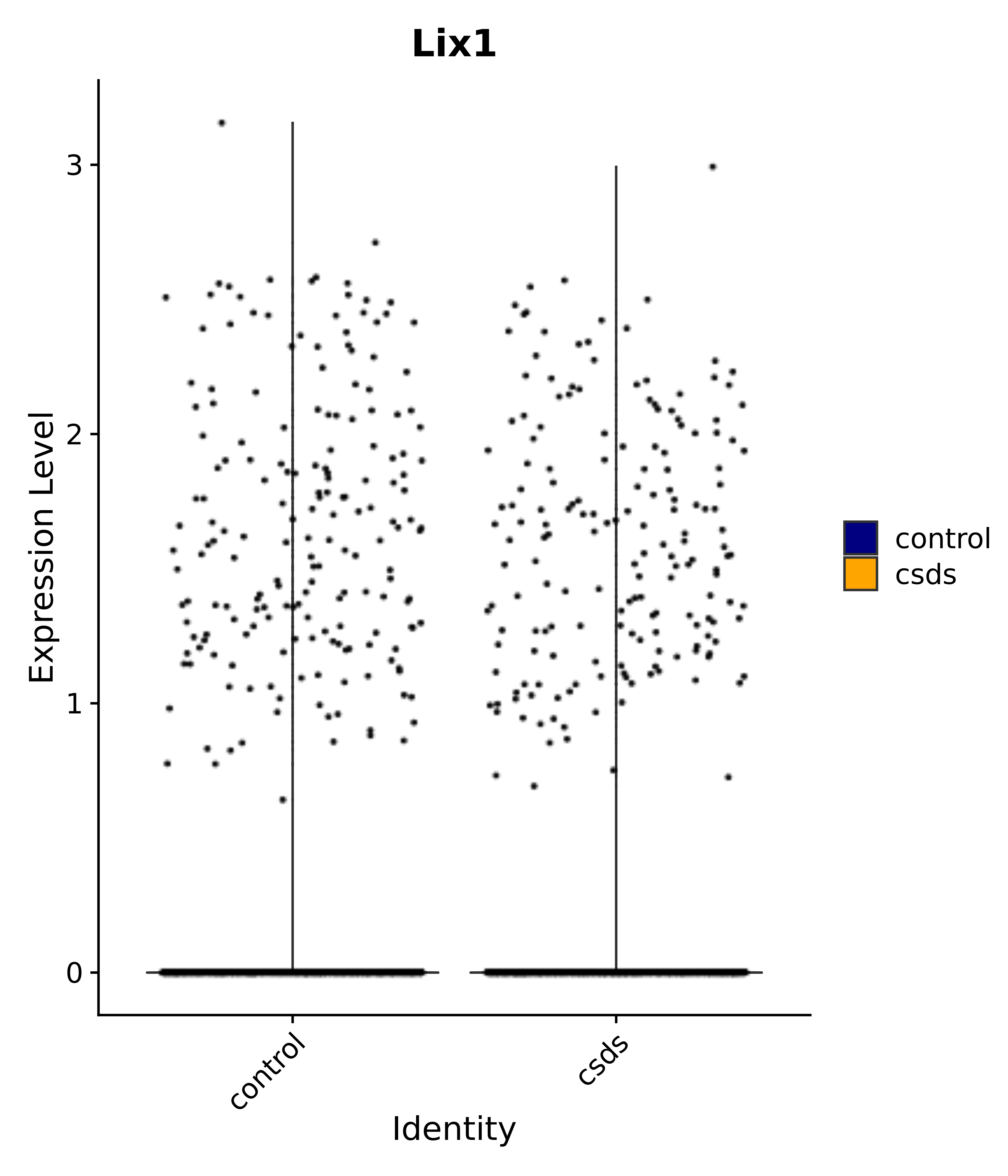
"Lix1",
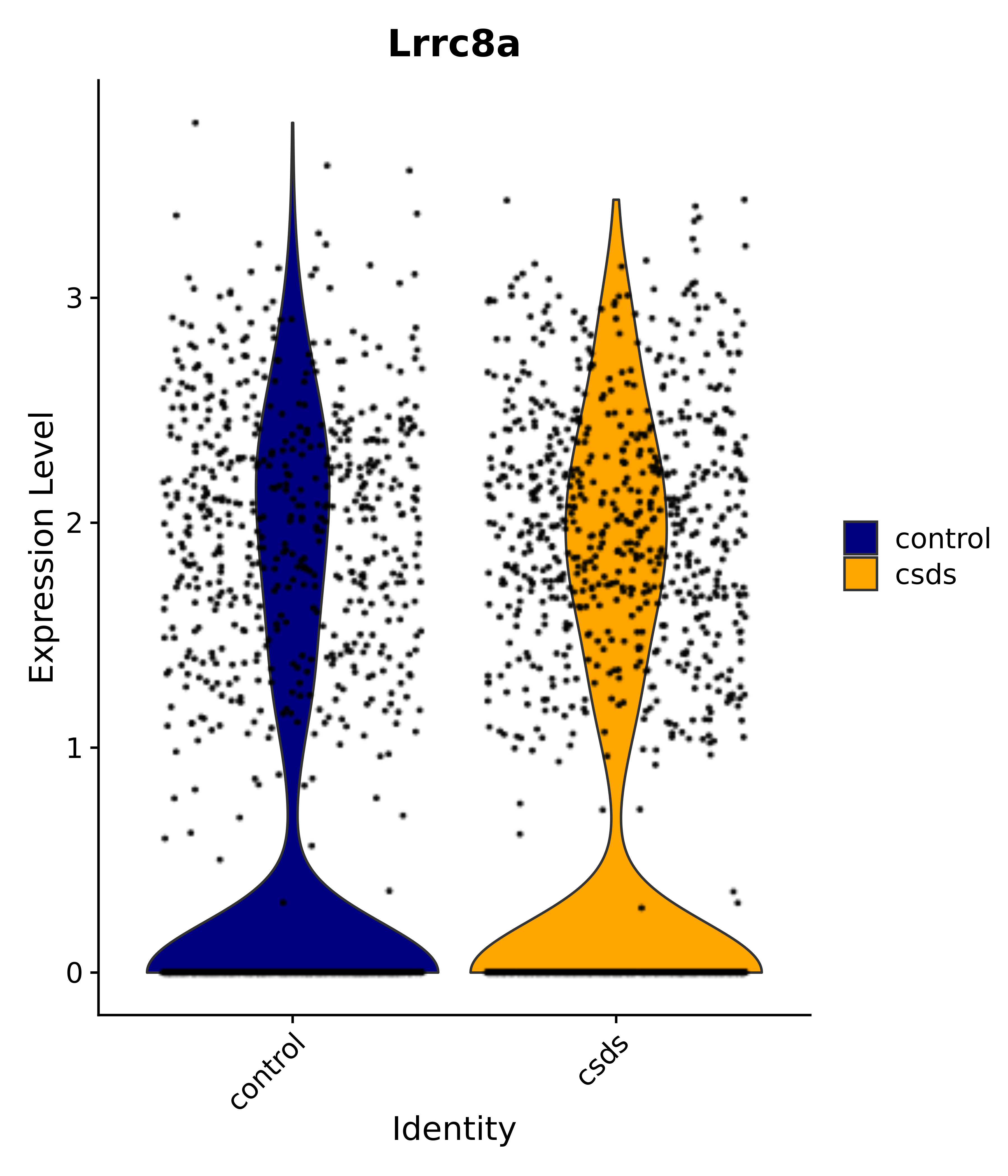
"Lrrc8a",
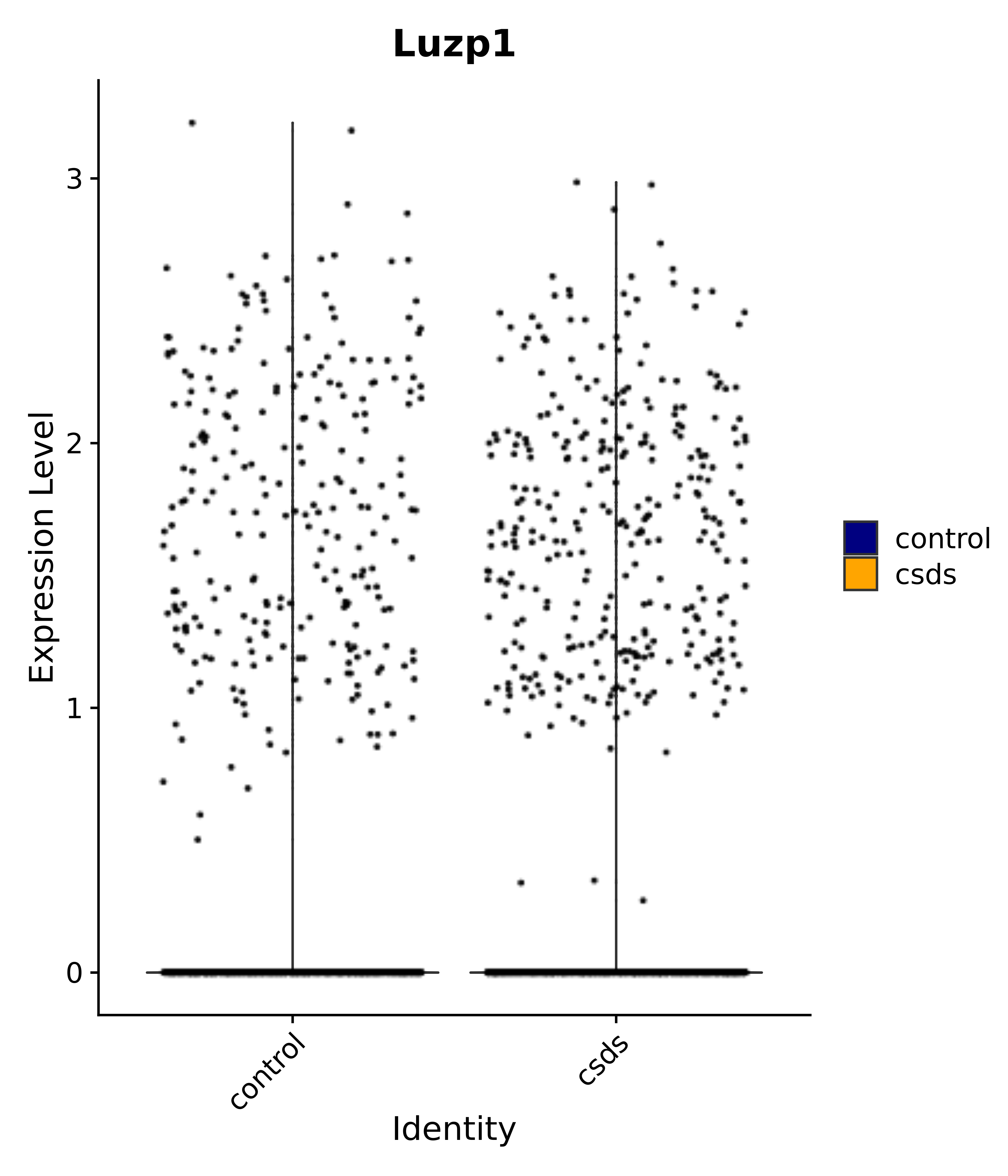
"Luzp1",
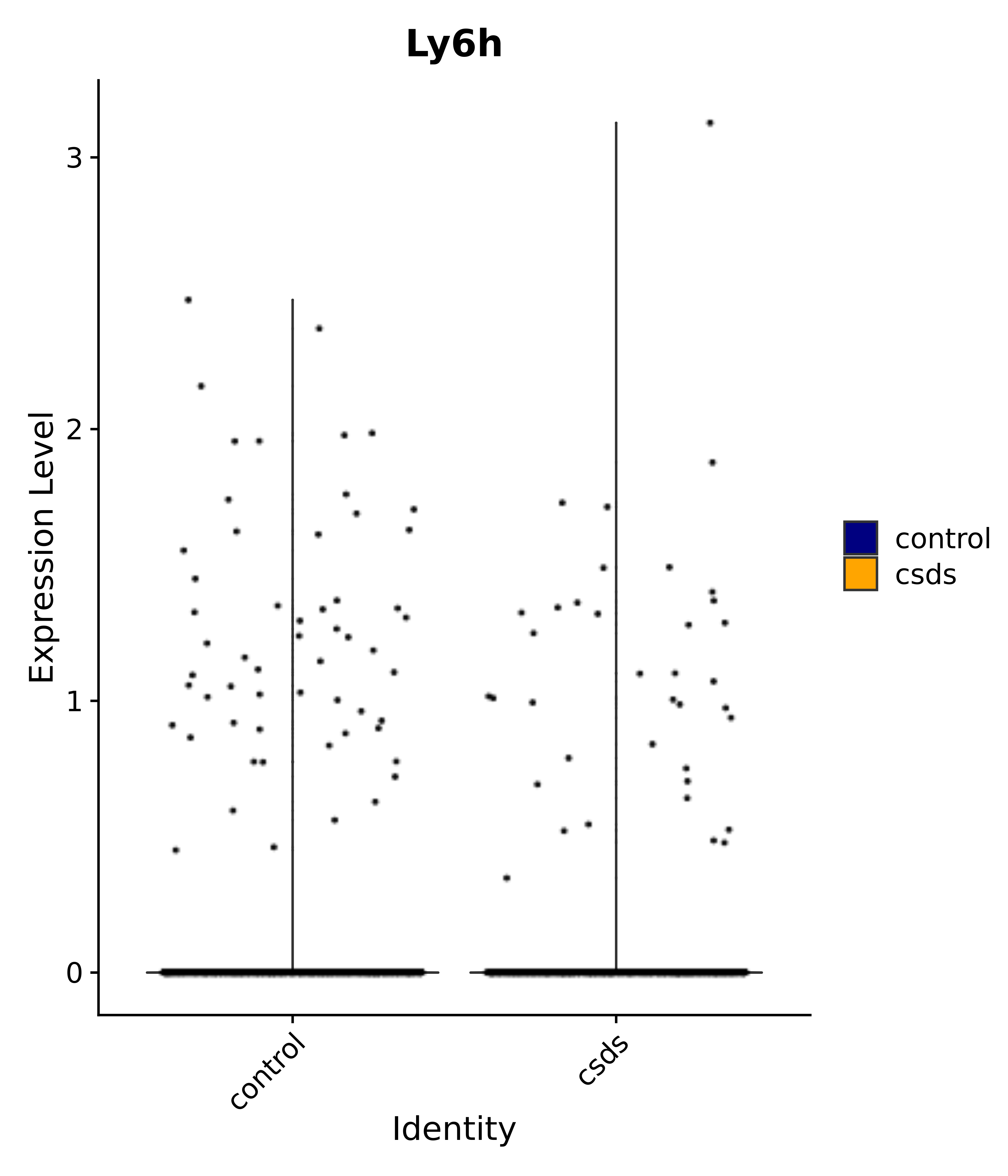
"Ly6h",
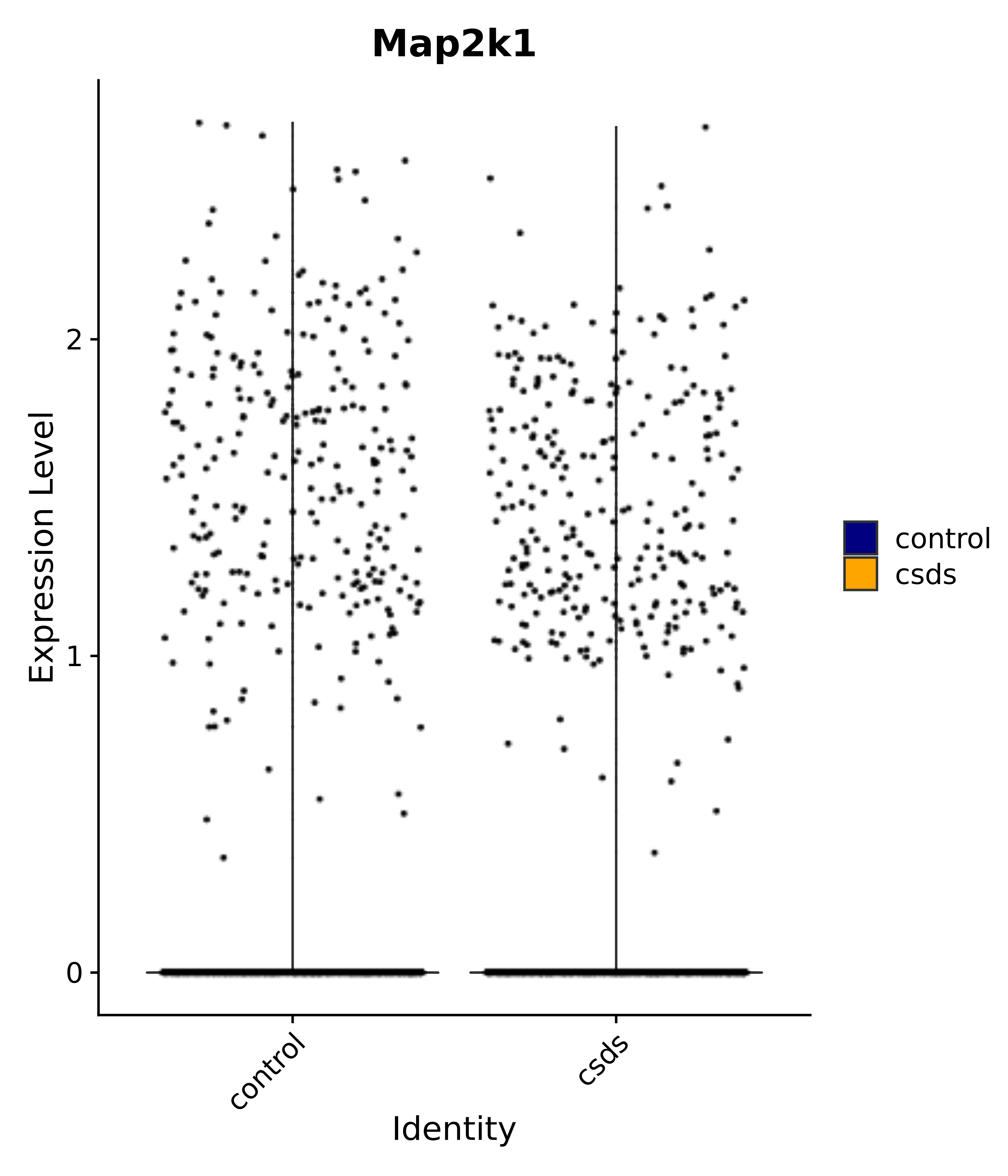
"Map2k1",
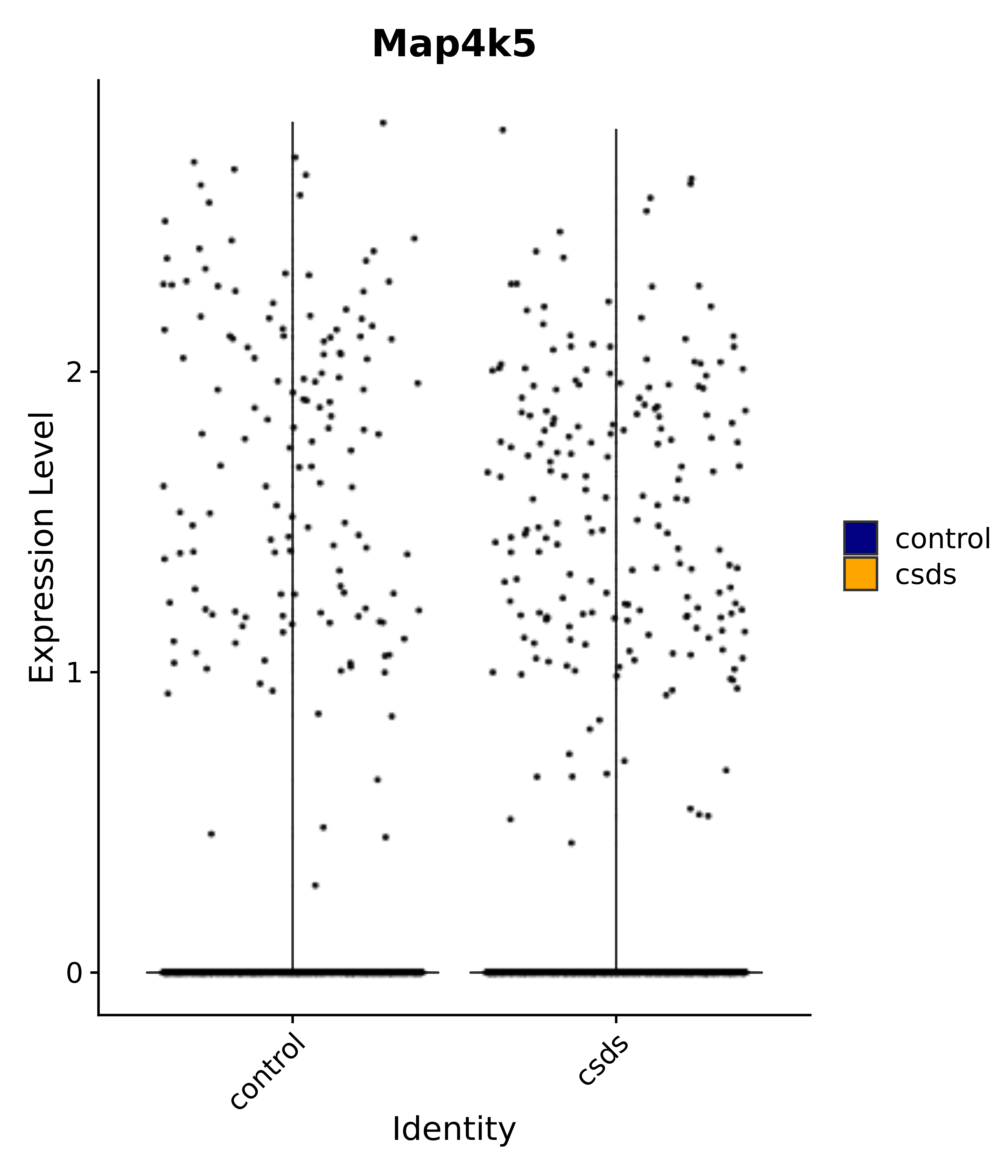
"Map4k5",
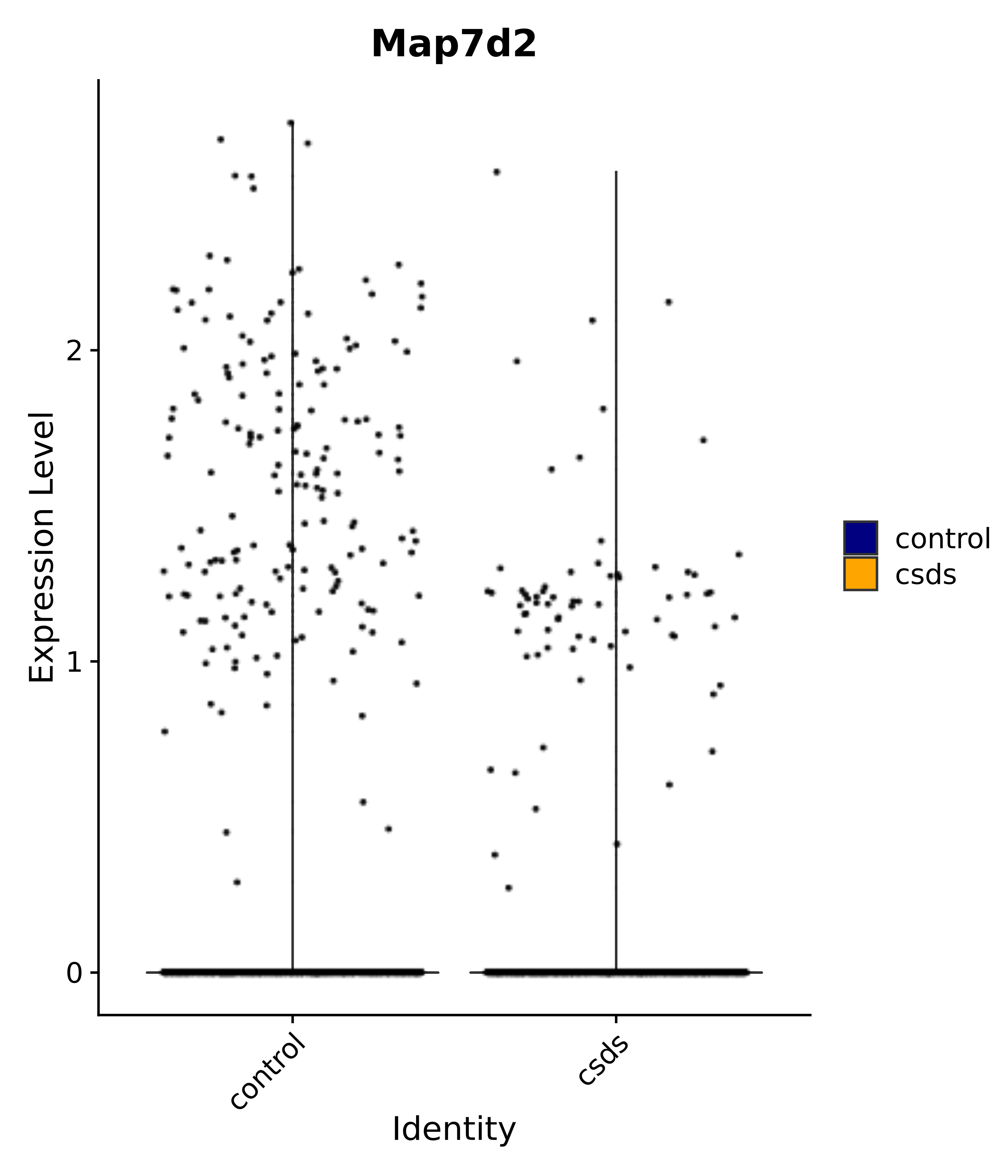
"Map7d2",
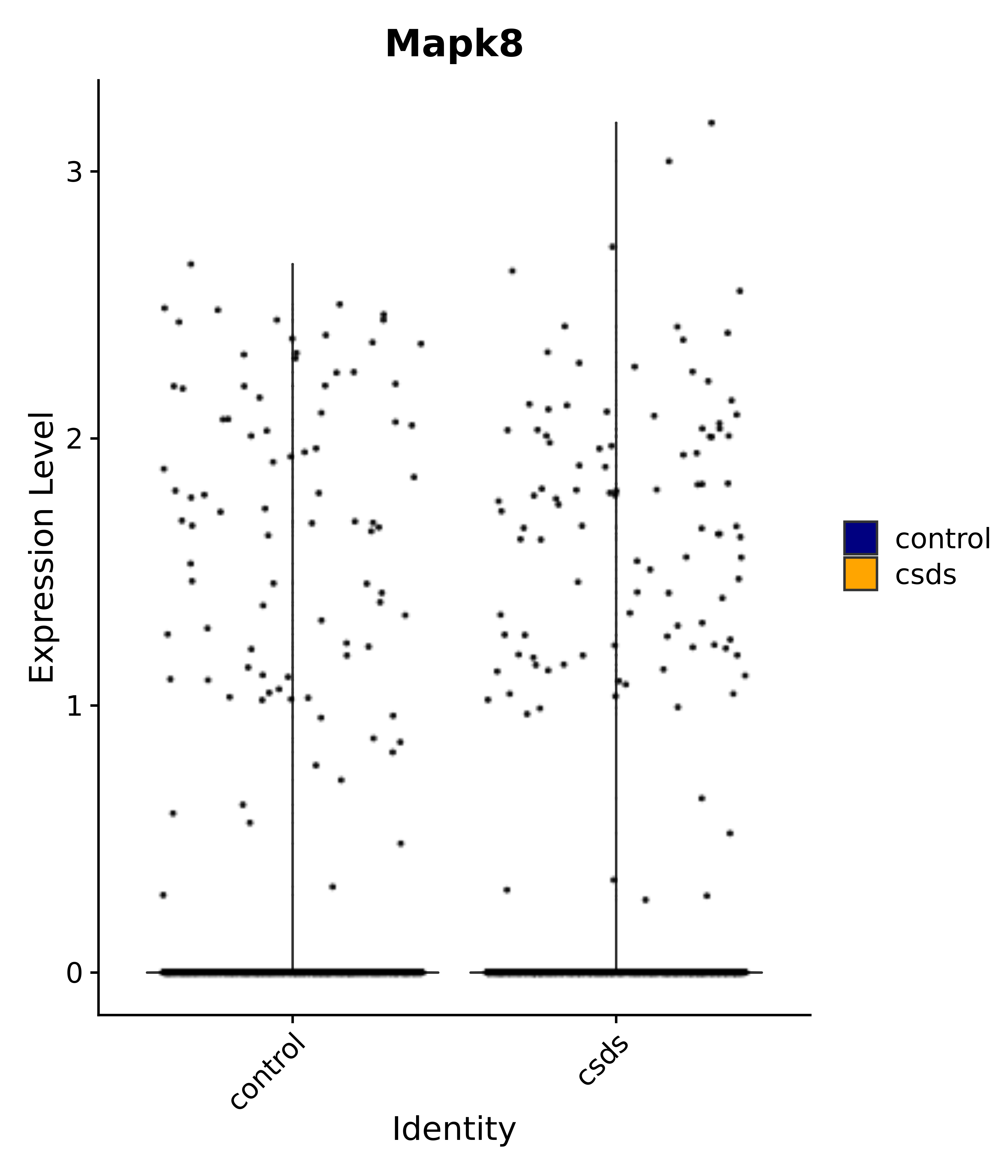
"Mapk8",
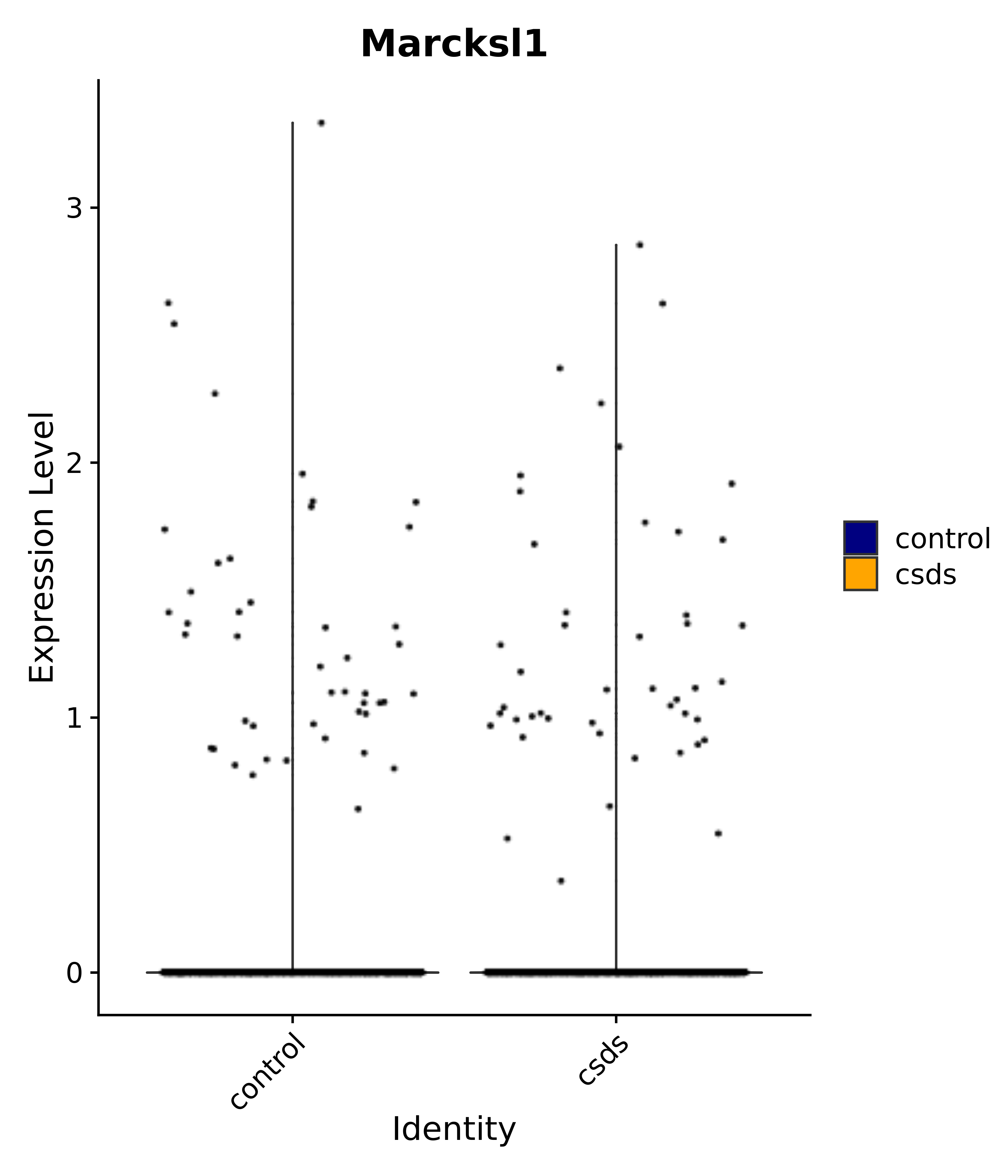
"Marcksl1",
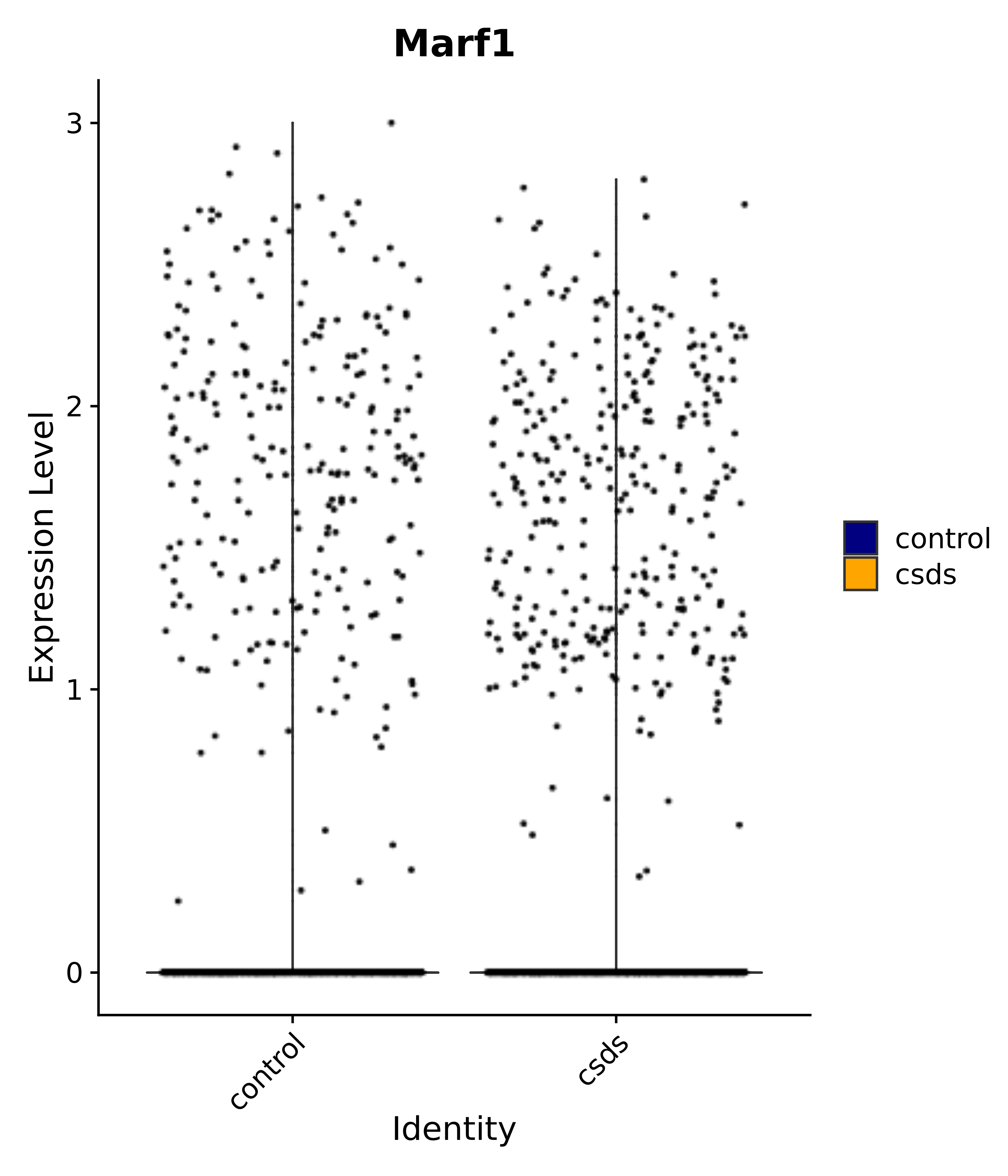
"Marf1",
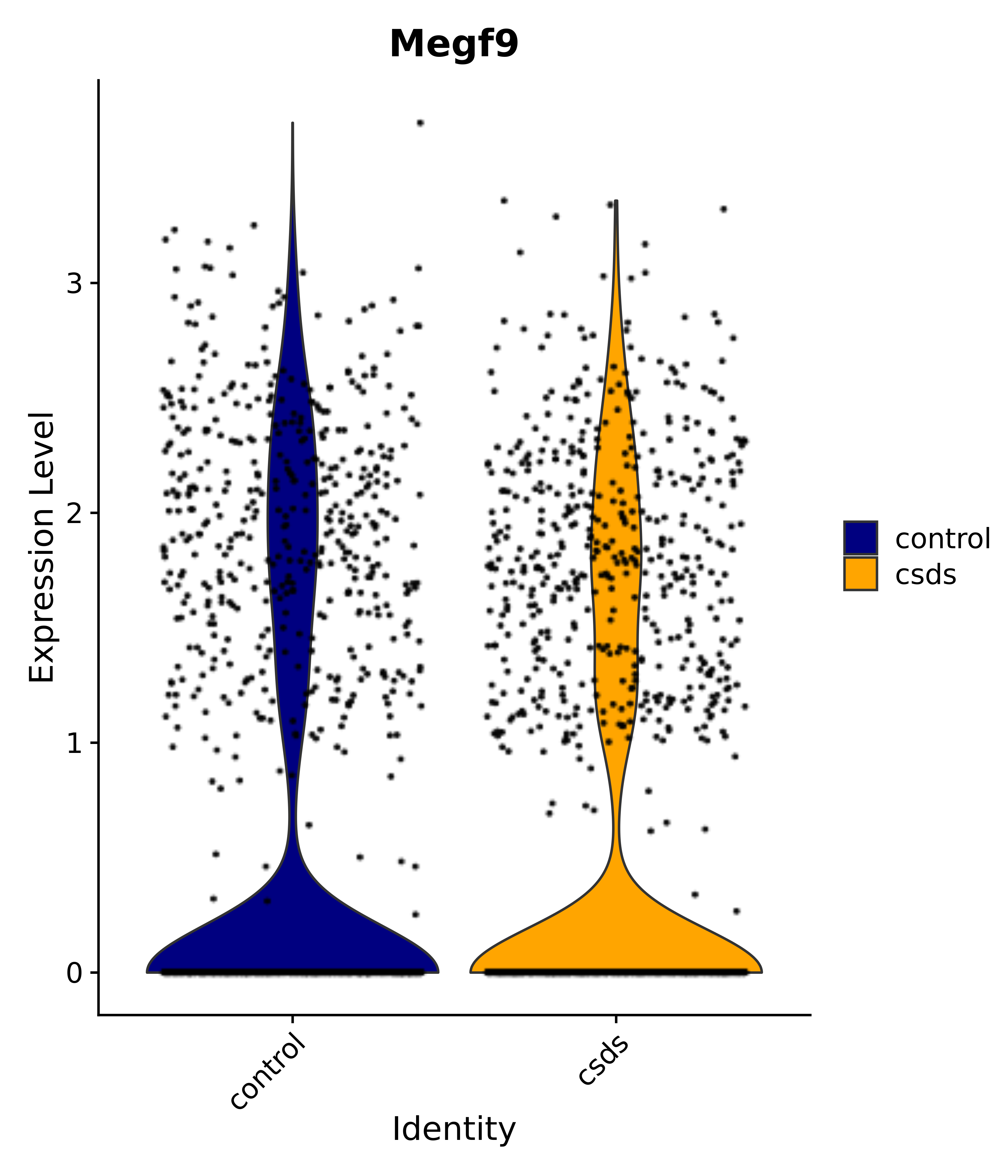
"Megf9",
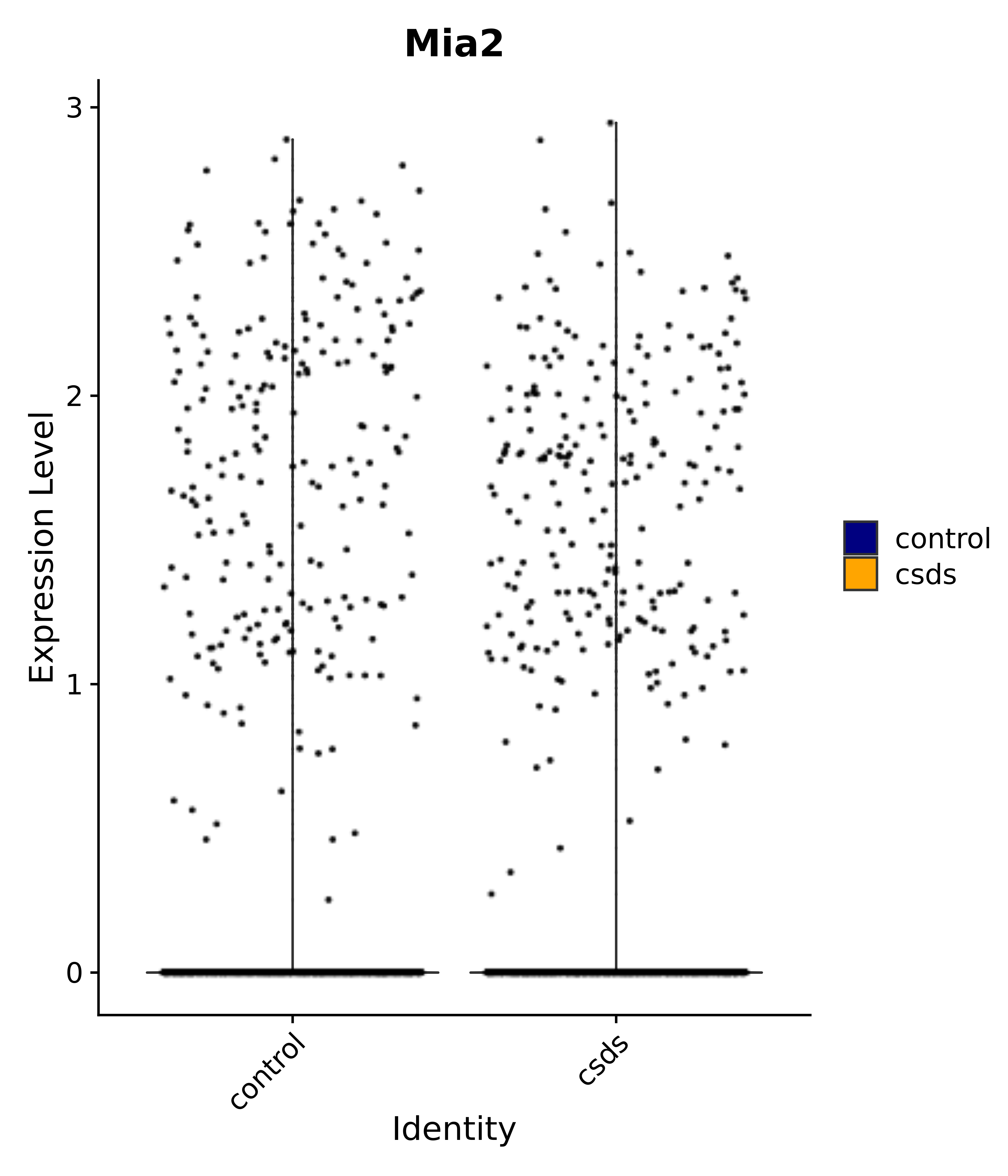
"Mia2",
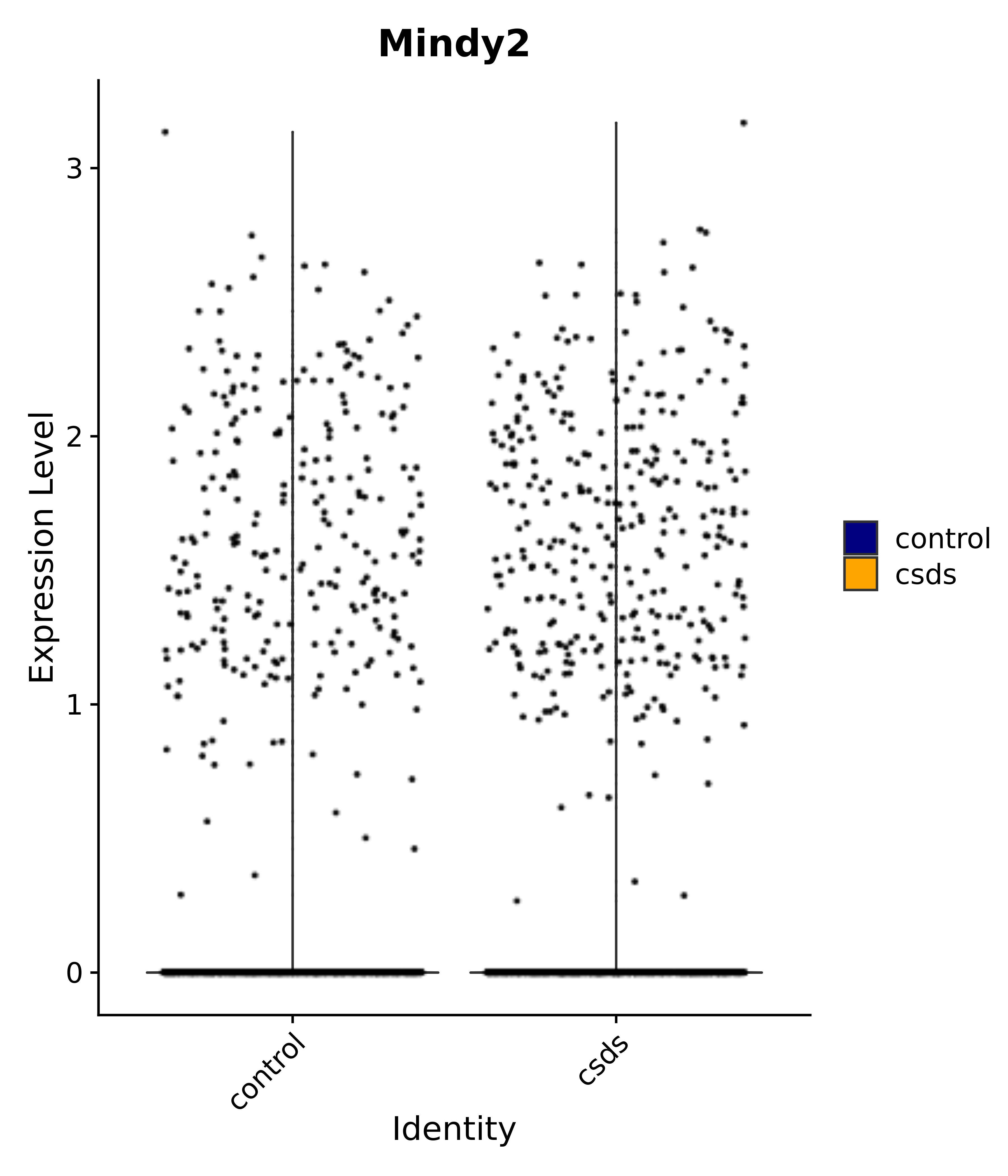
"Mindy2",
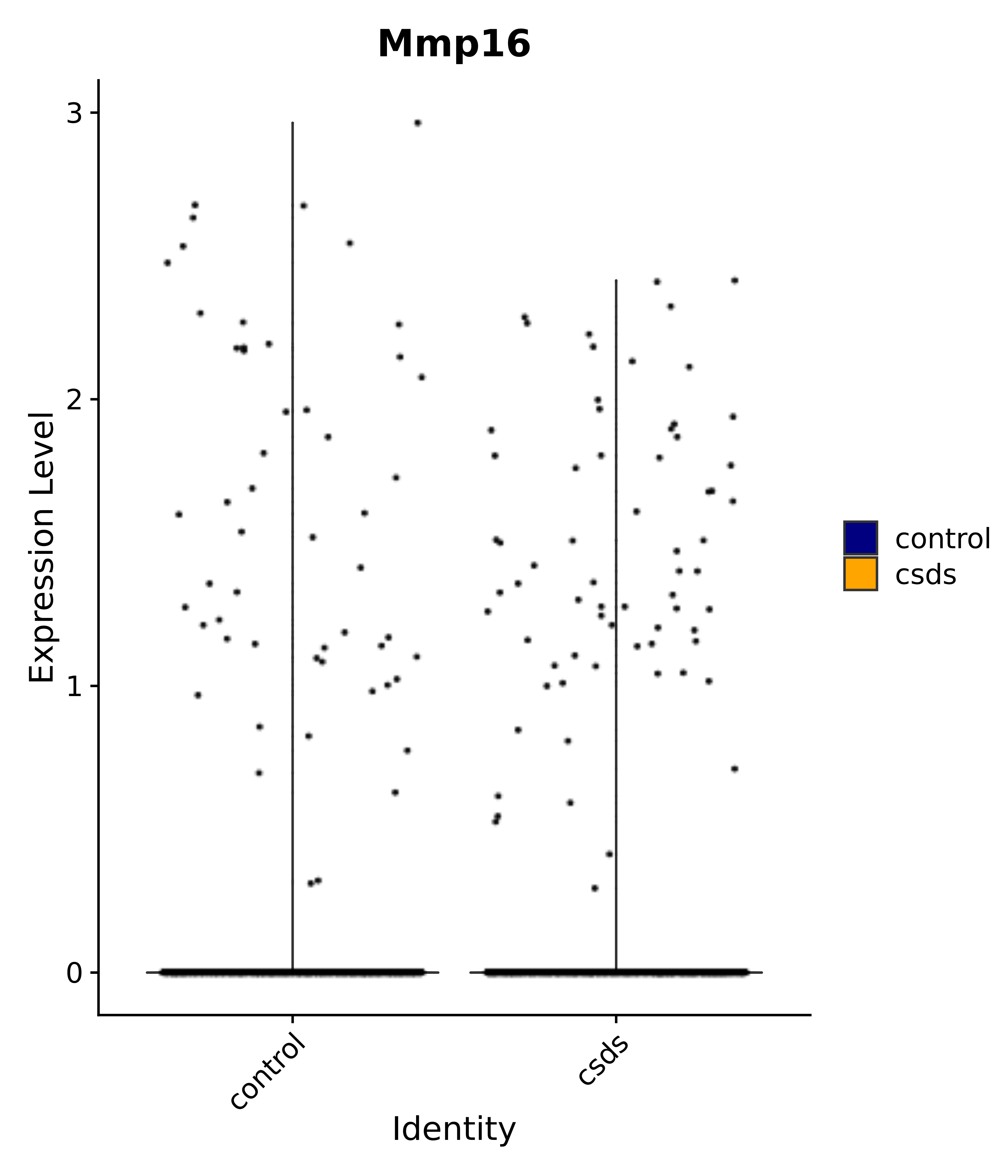
"Mmp16",
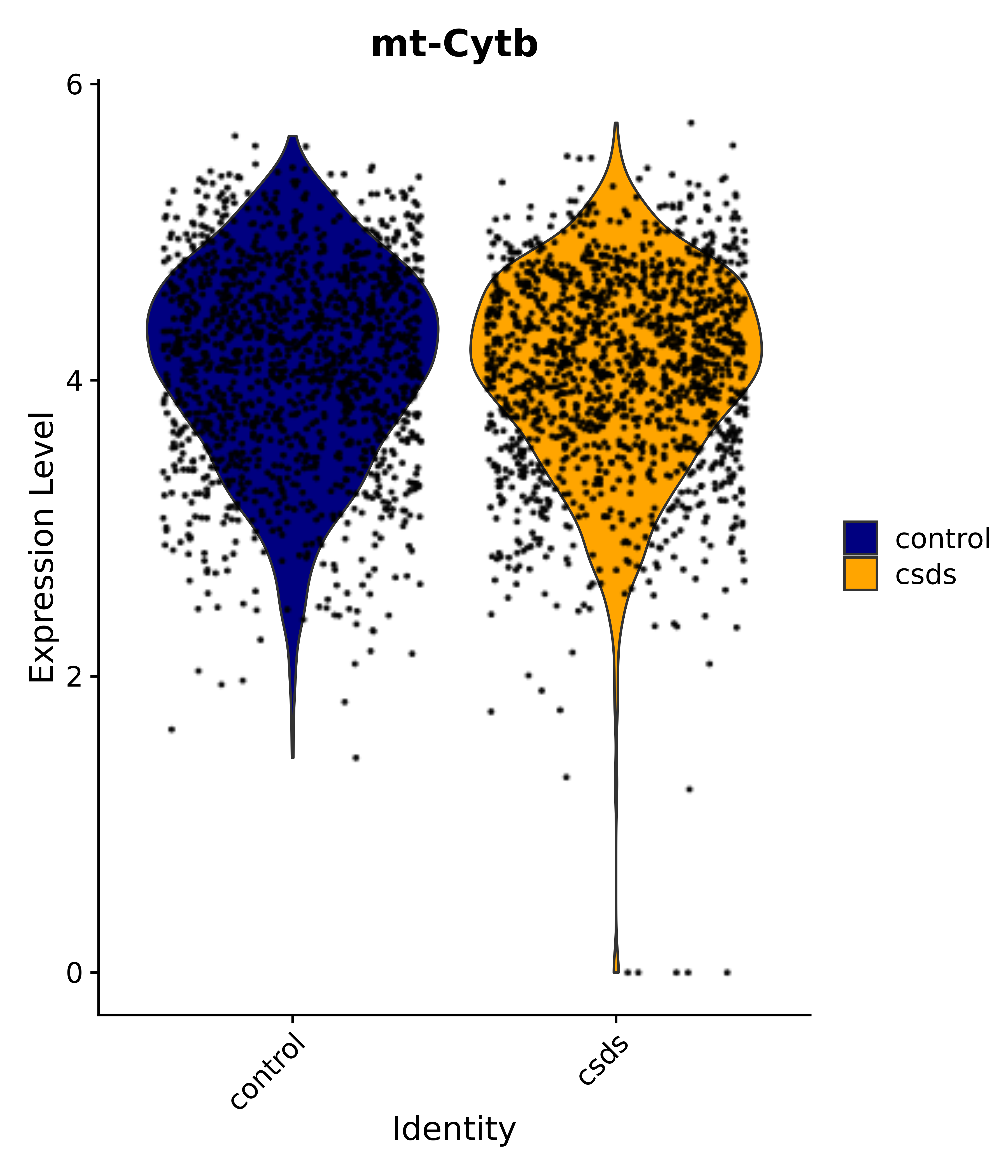
"mt-Cytb",
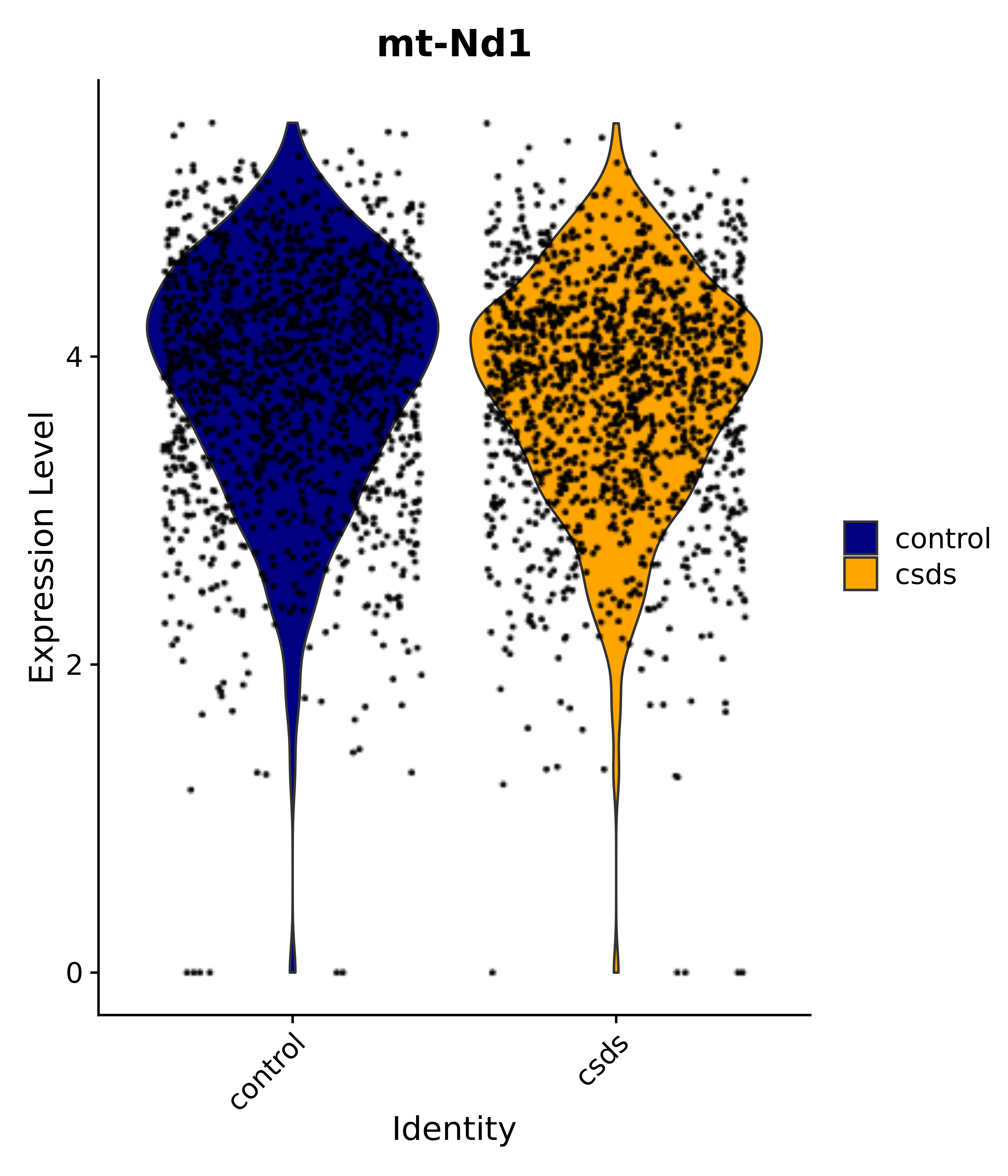
"mt-Nd1",
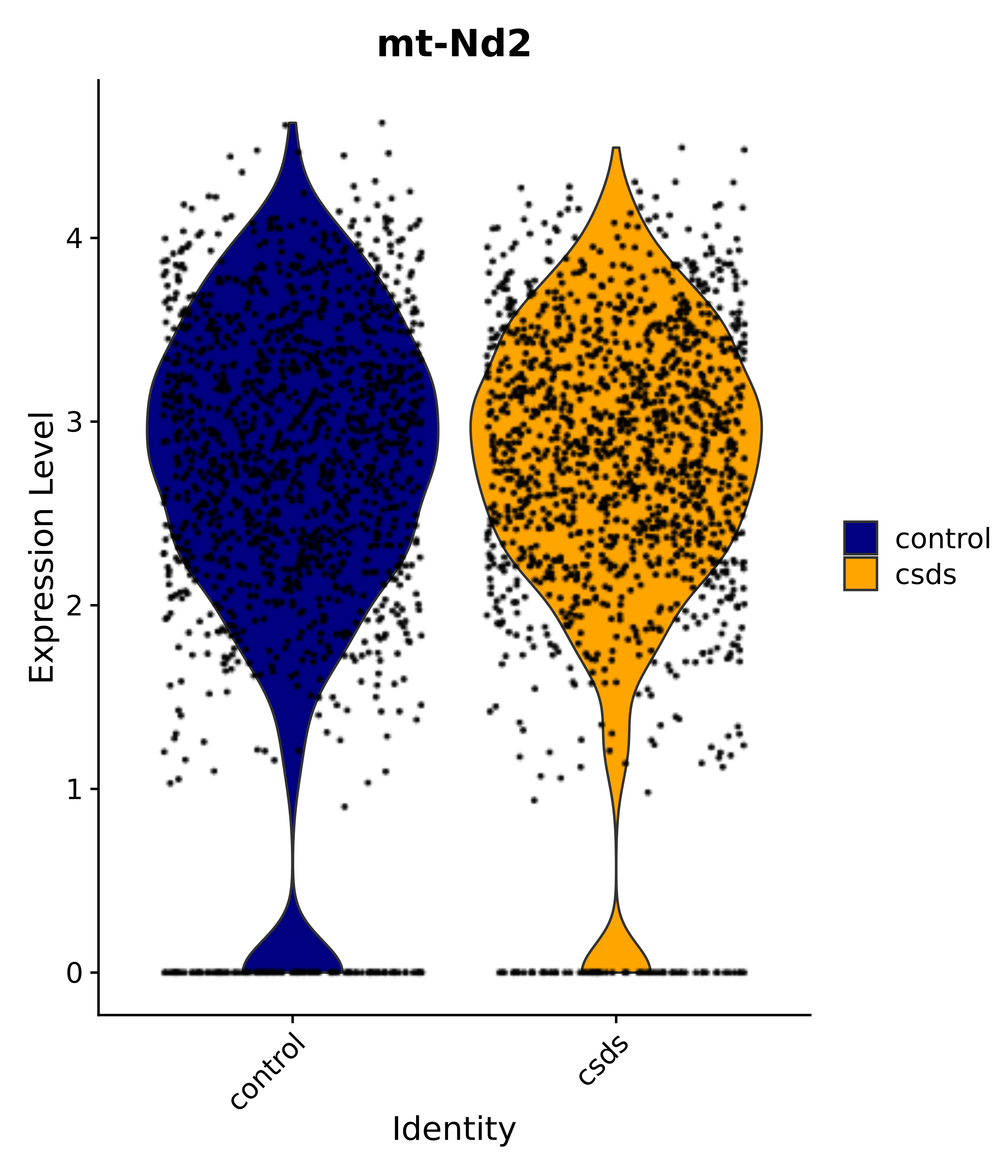
"mt-Nd2",
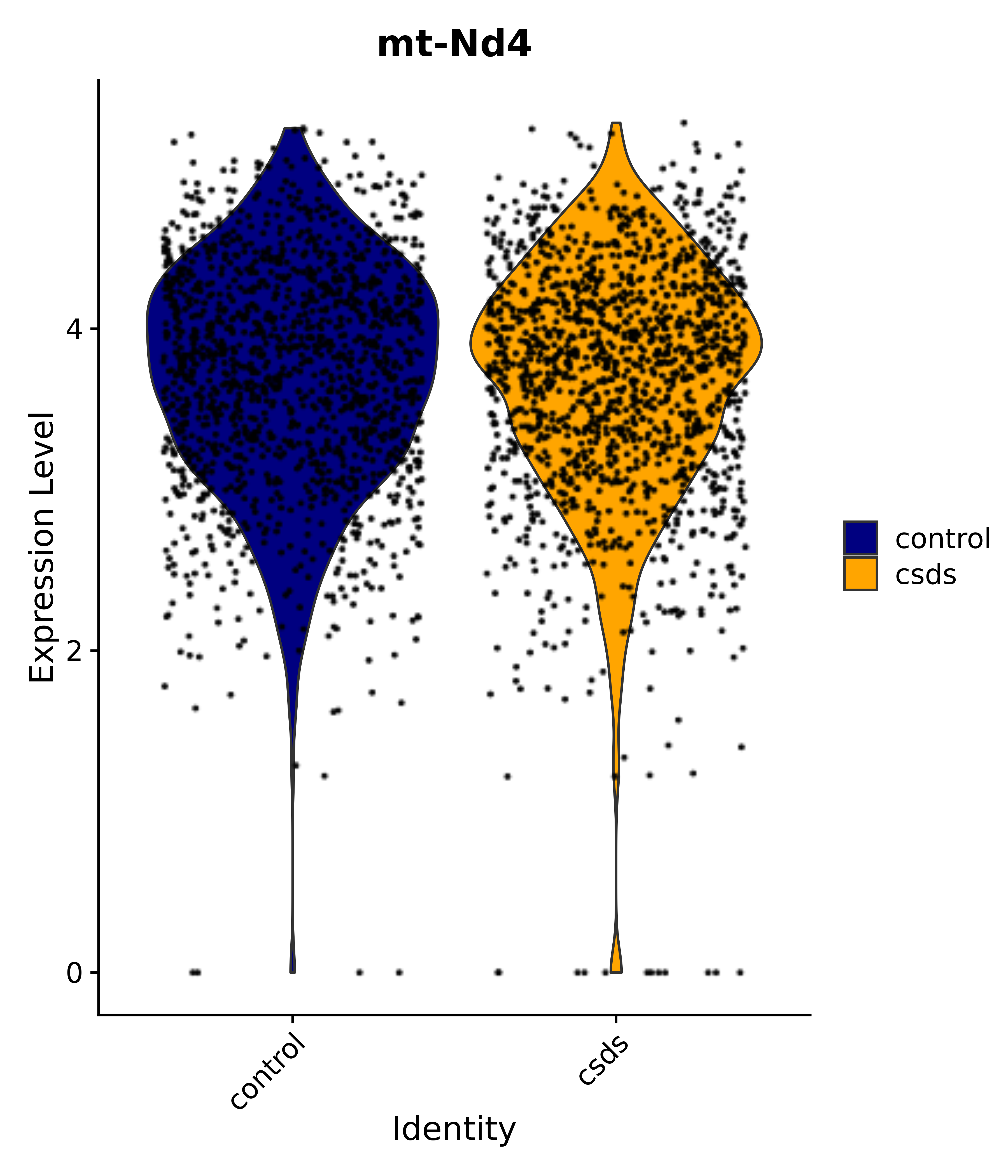
"mt-Nd4",
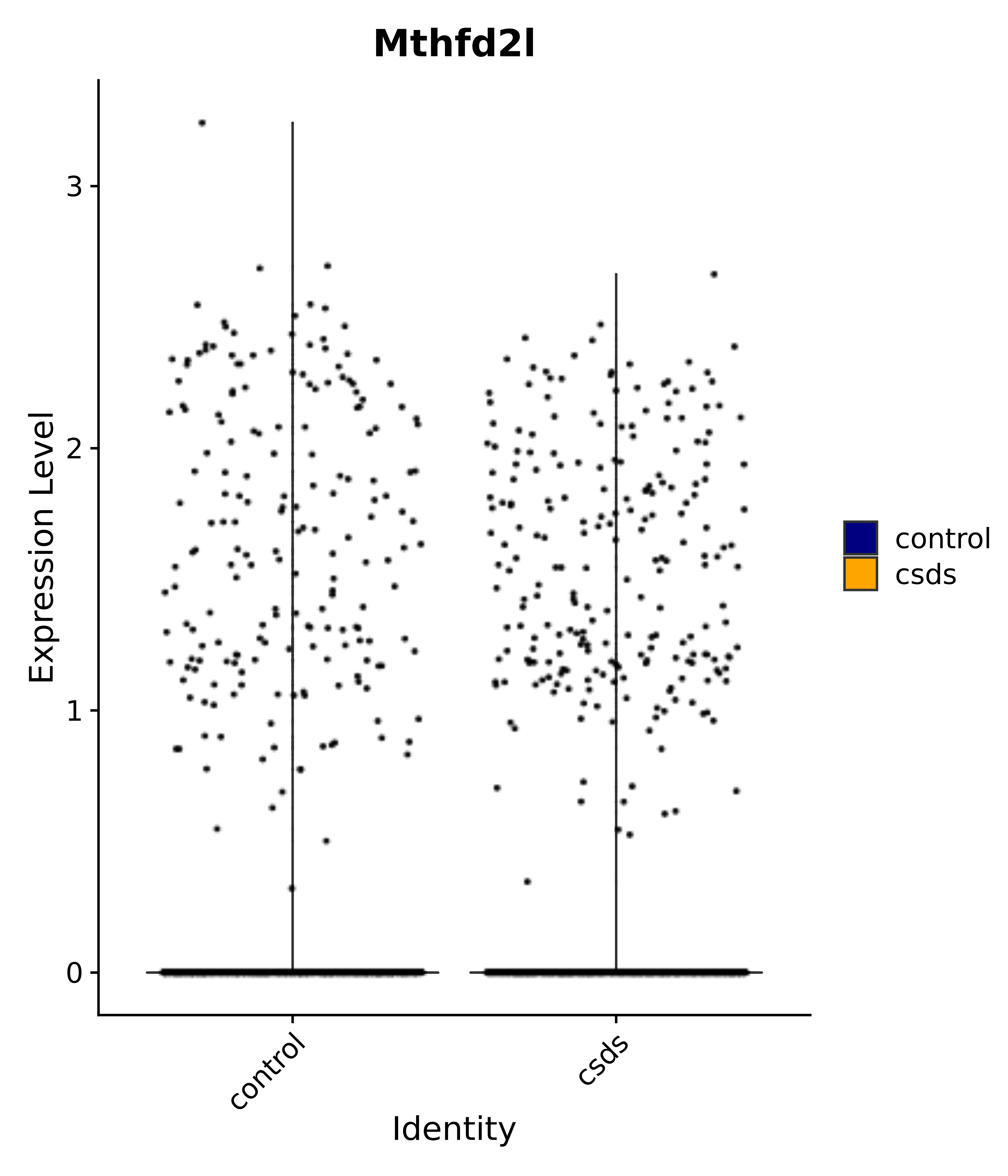
"Mthfd2l",
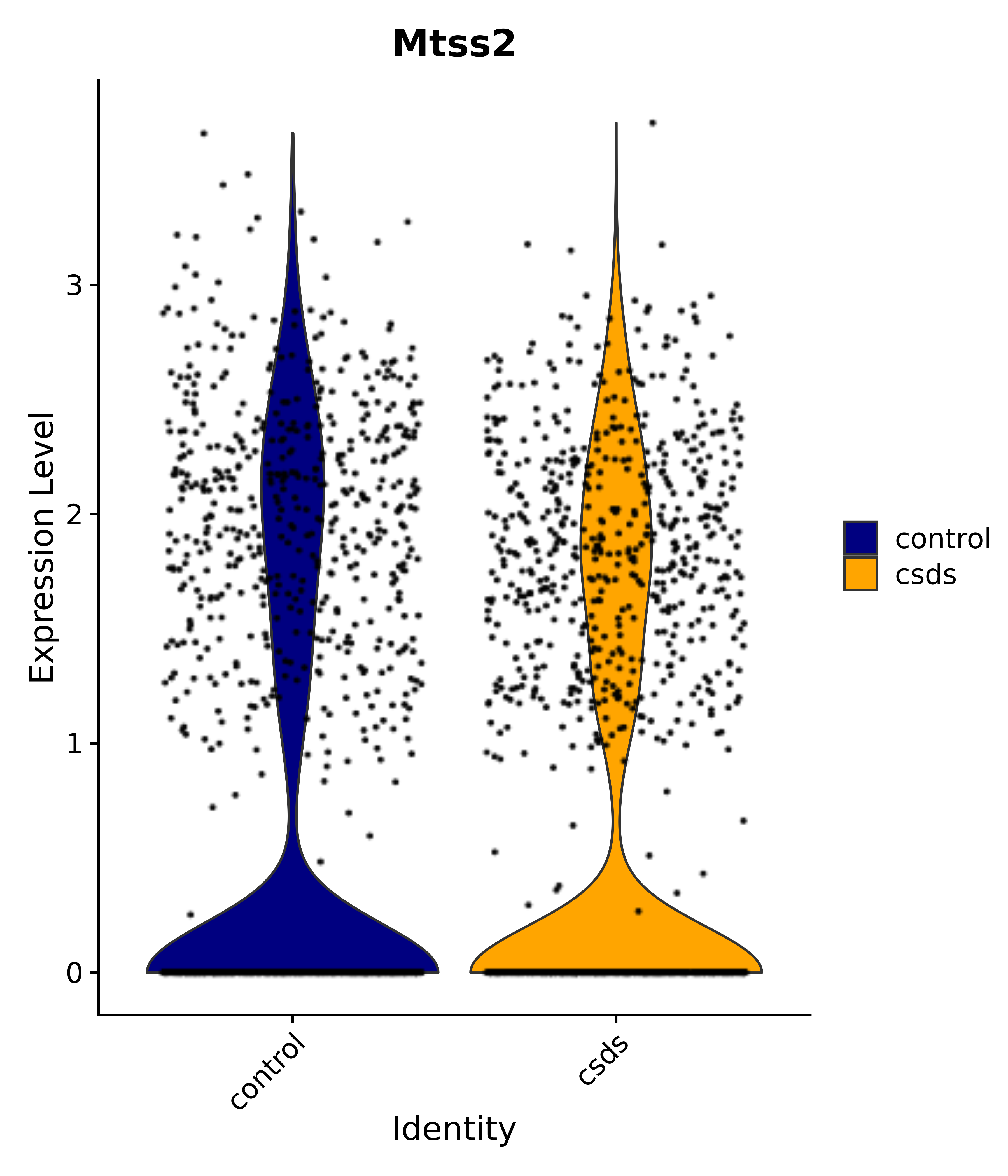
"Mtss2",
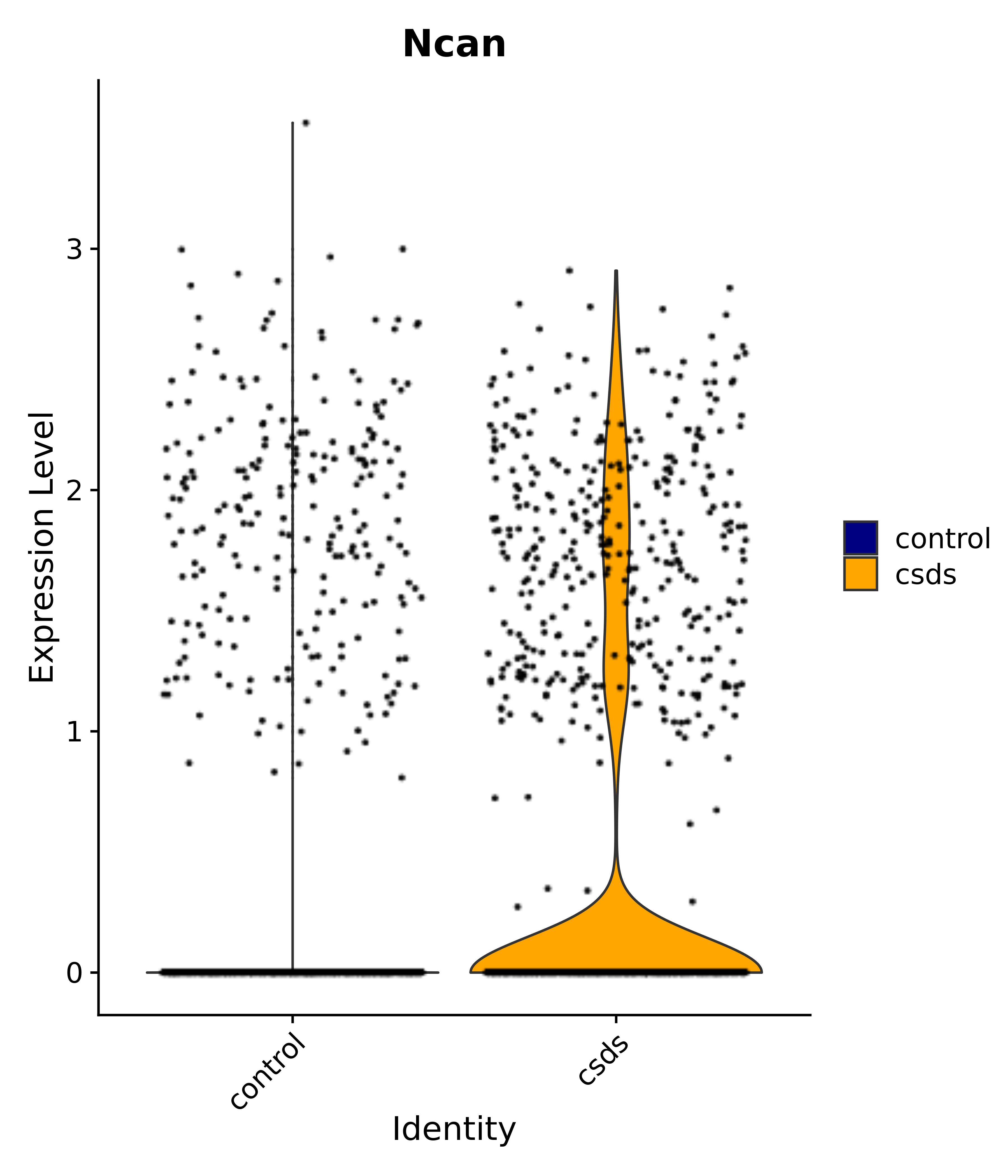
"Ncan",
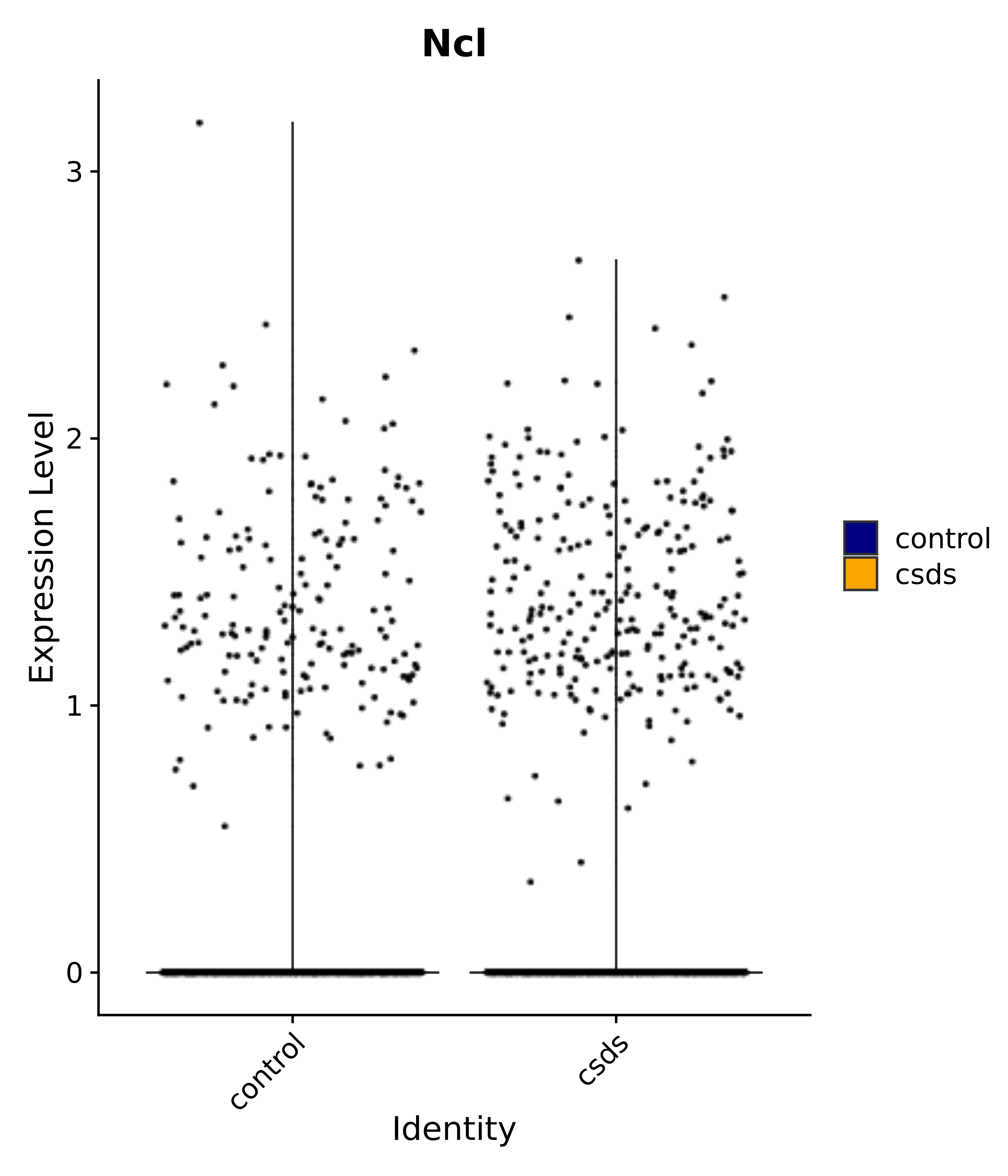
"Ncl",
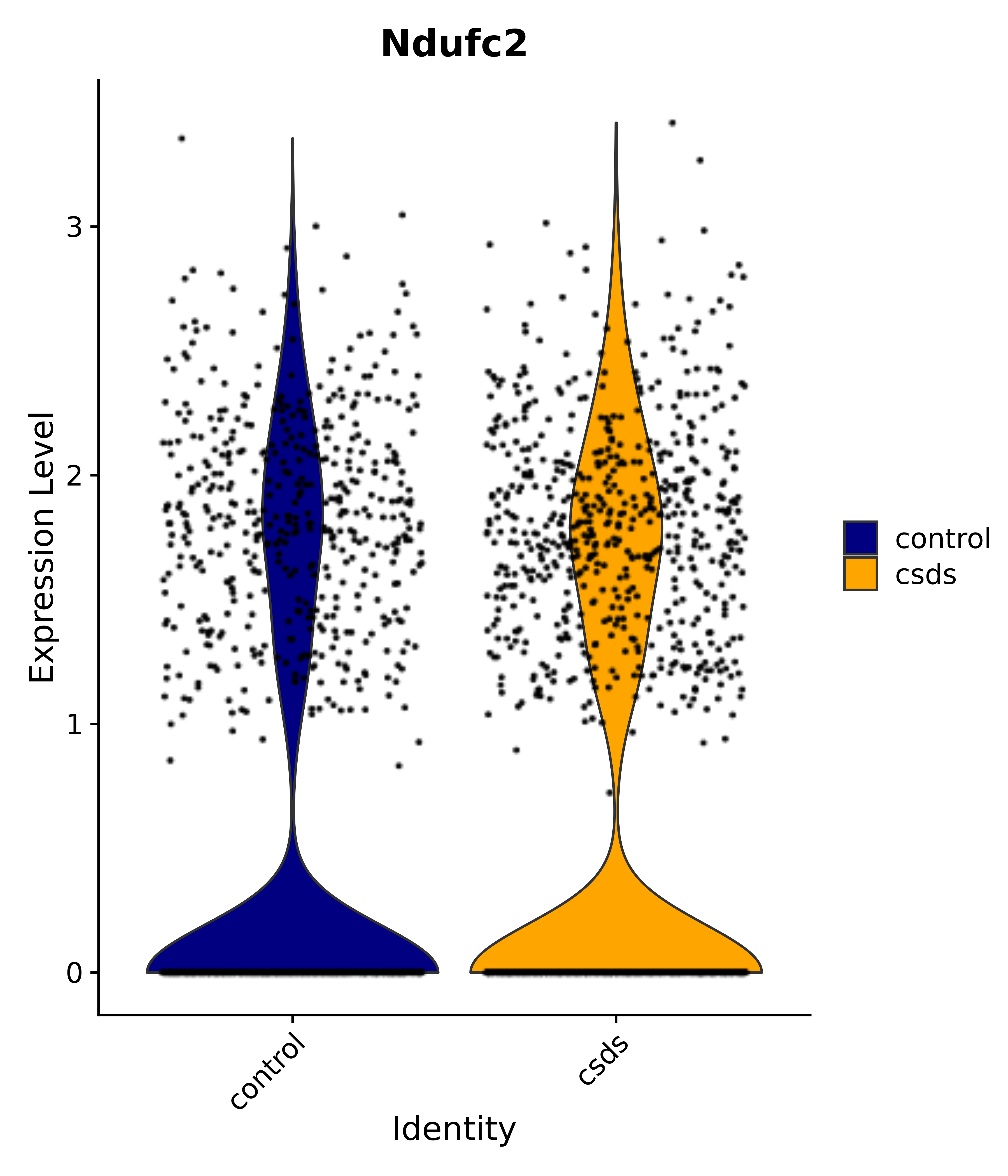
"Ndufc2",
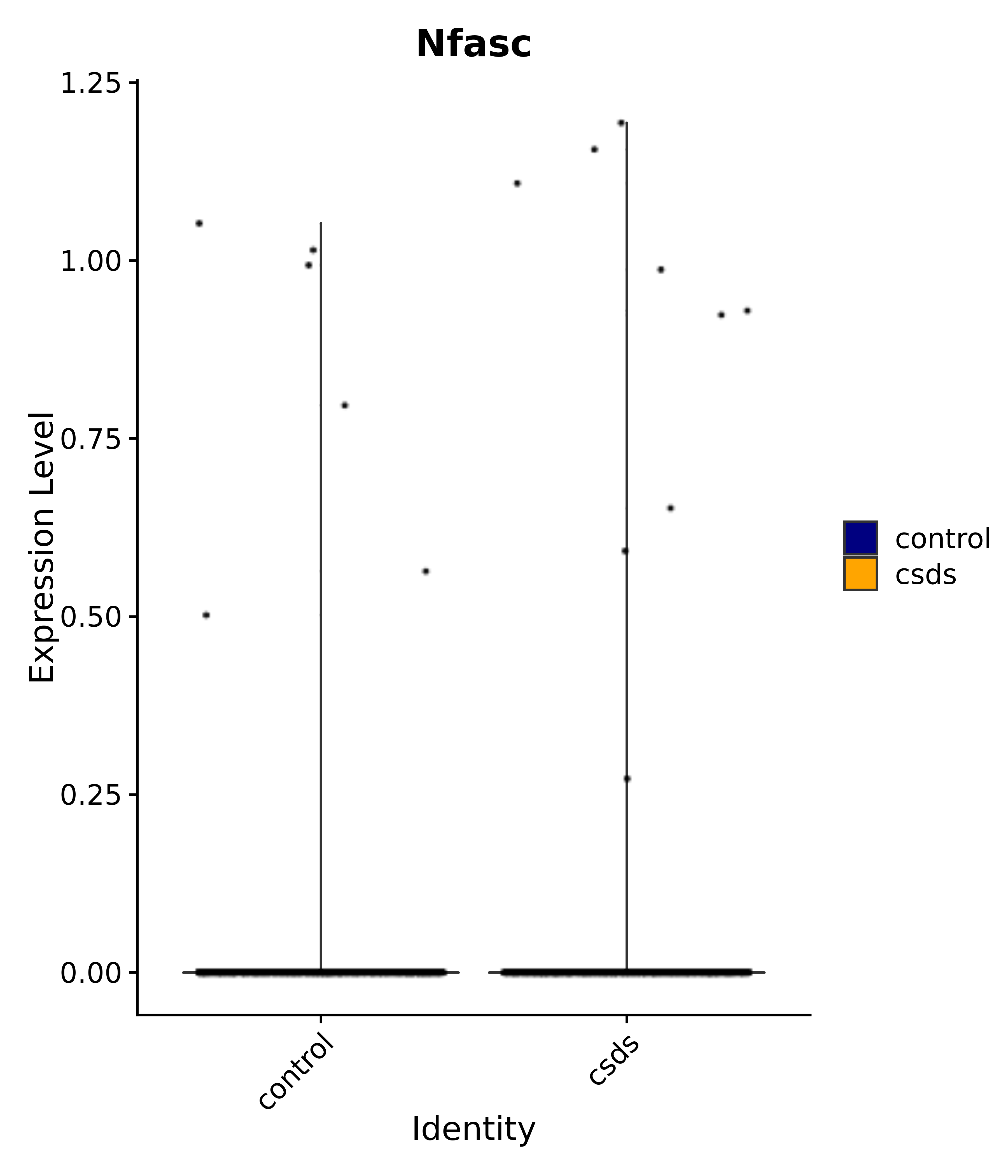
"Nfasc",
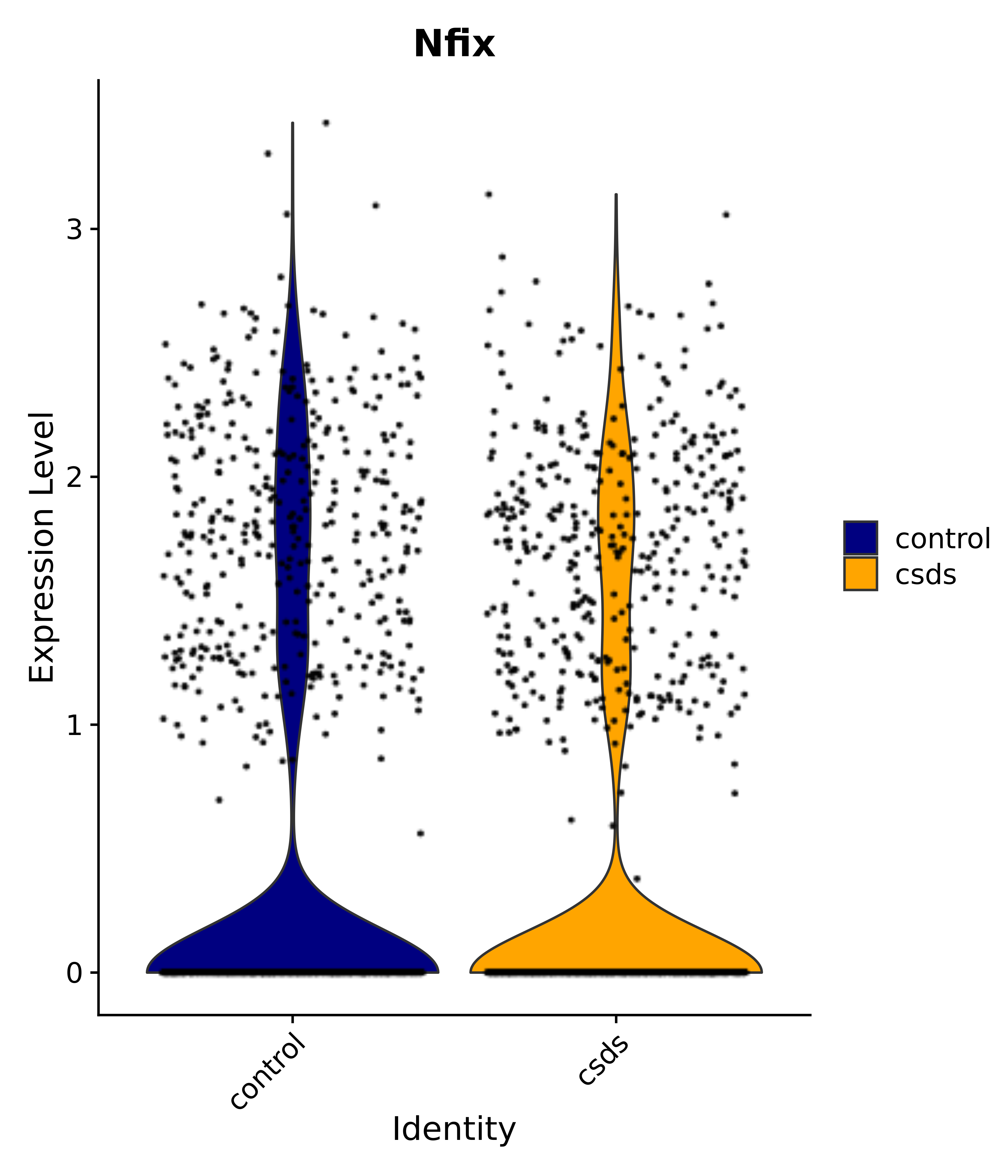
"Nfix",
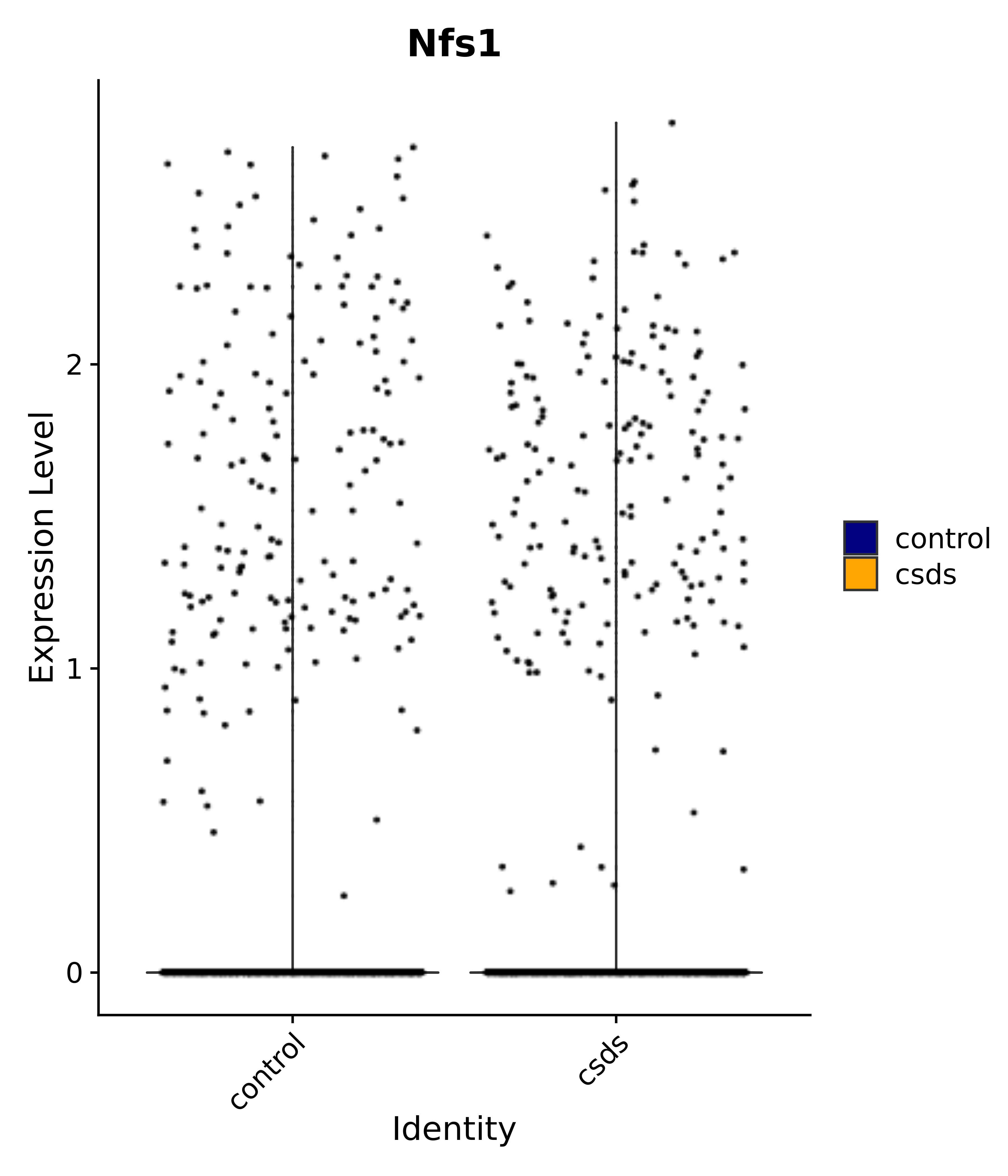
"Nfs1",
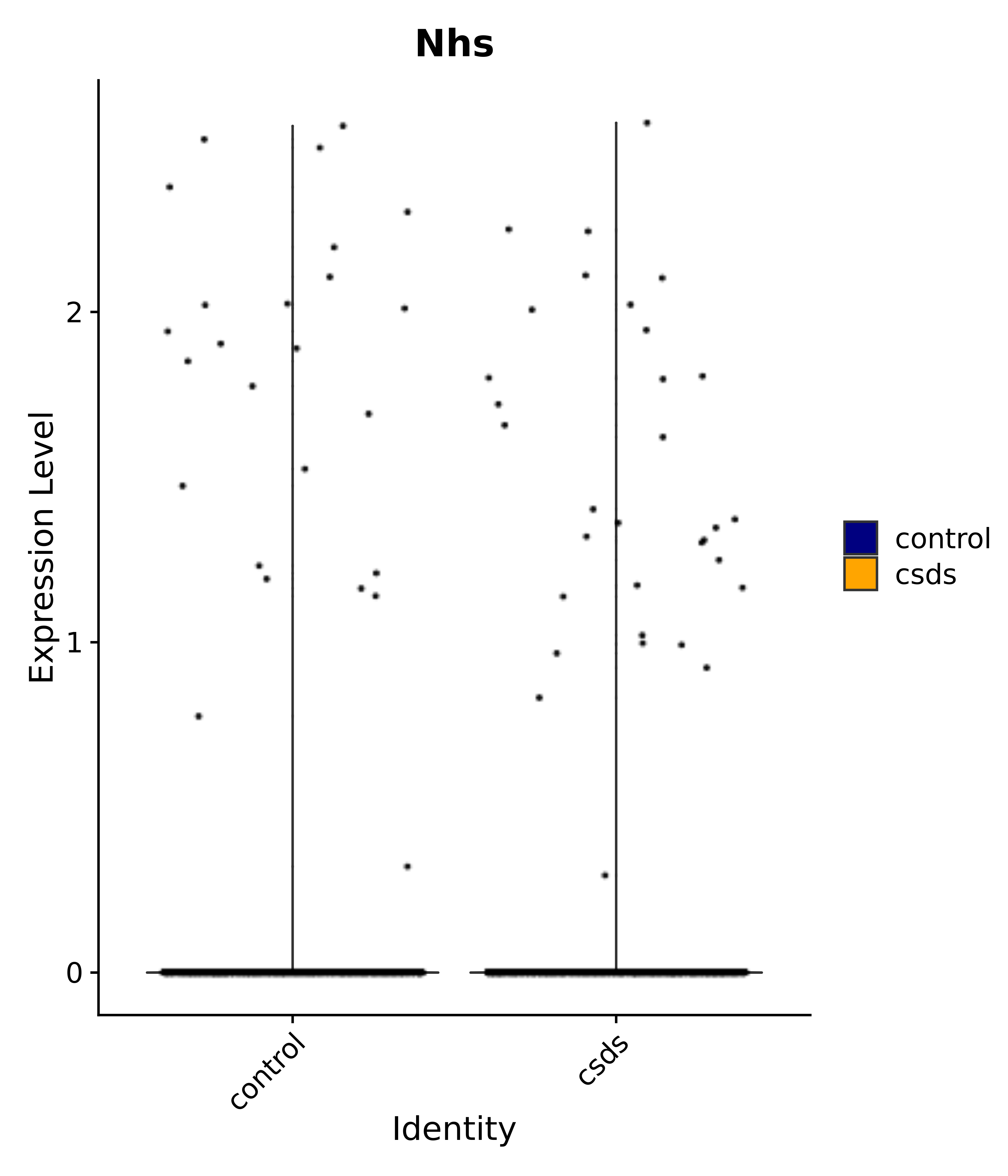
"Nhs",
"Nkx6-2",
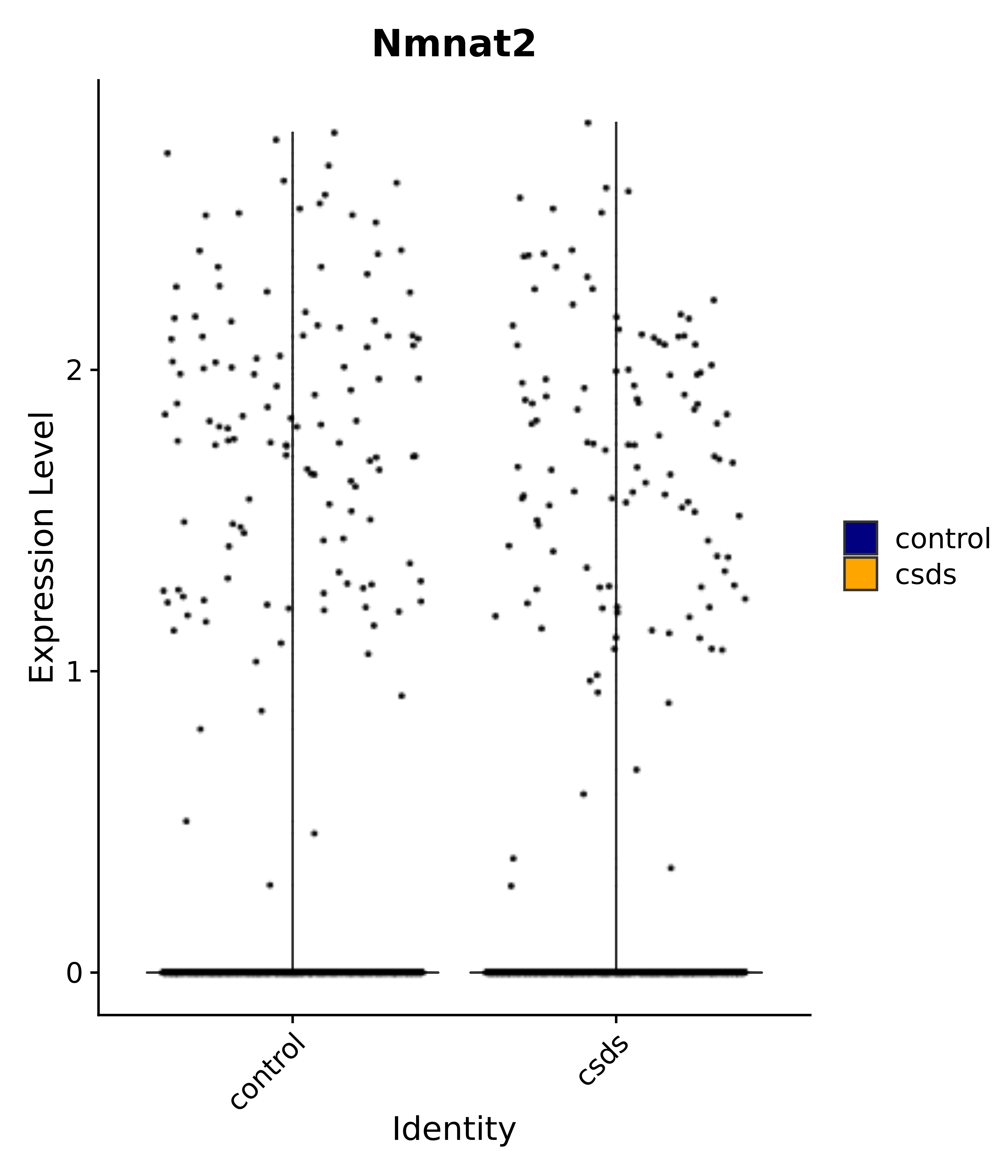
"Nmnat2",
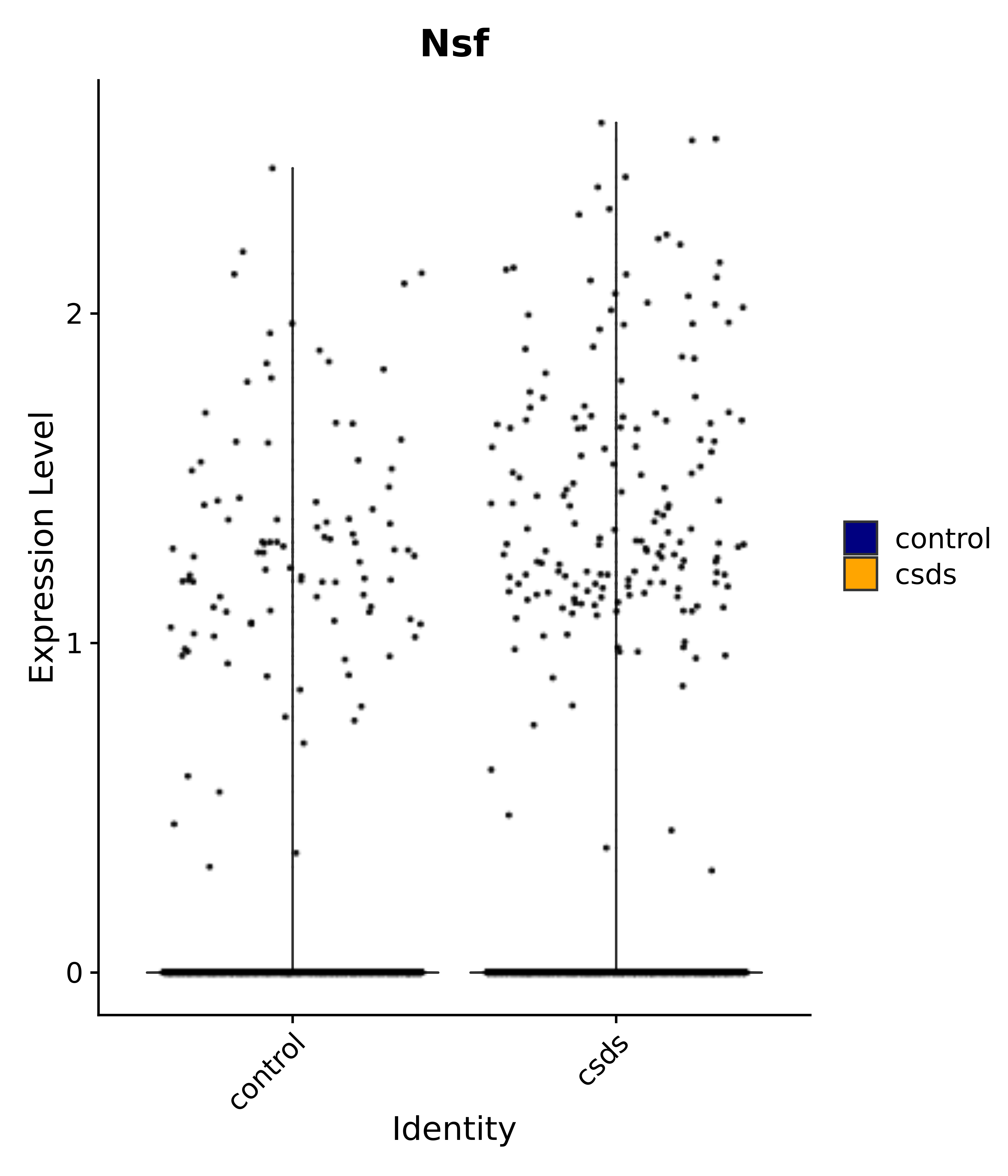
"Nsf",
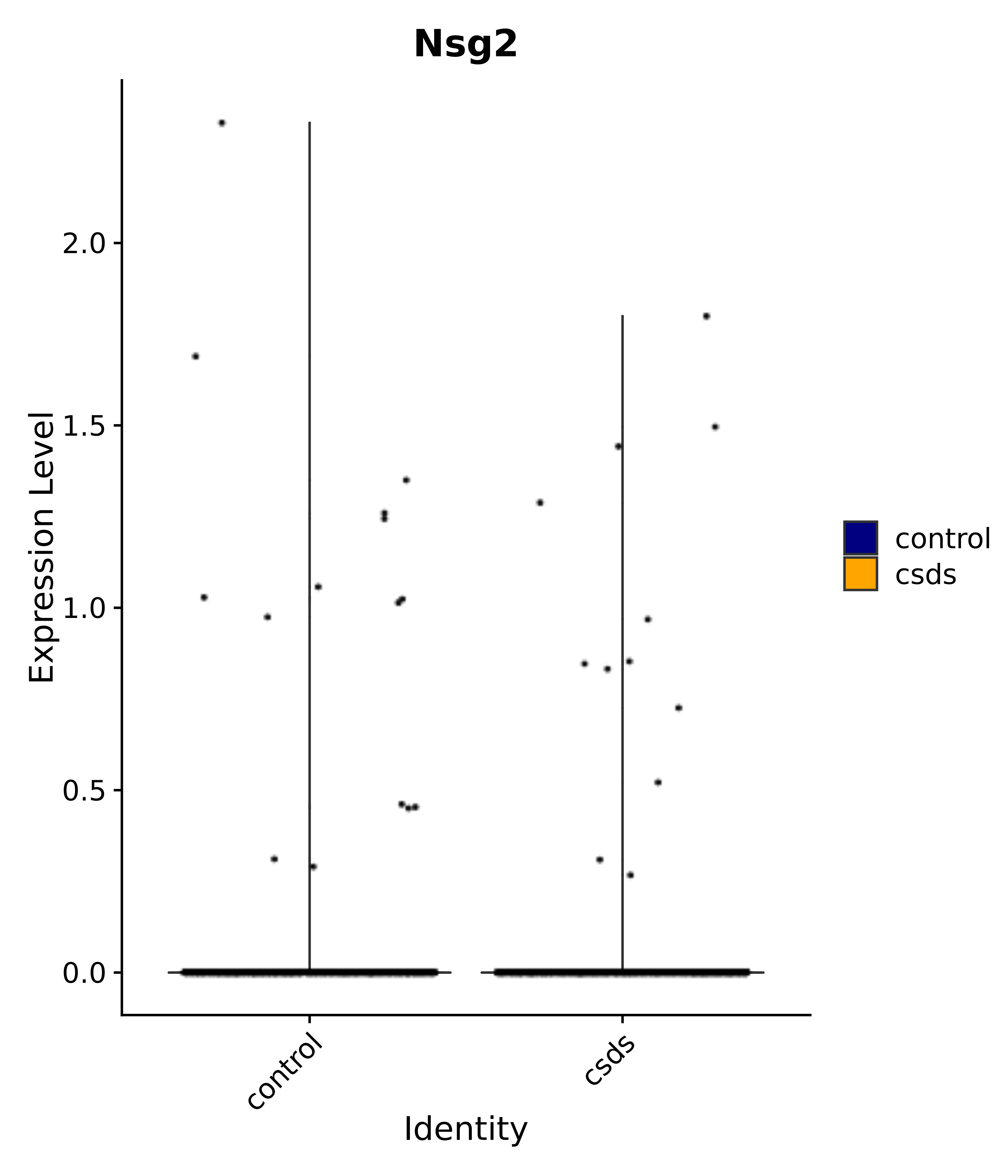
"Nsg2",
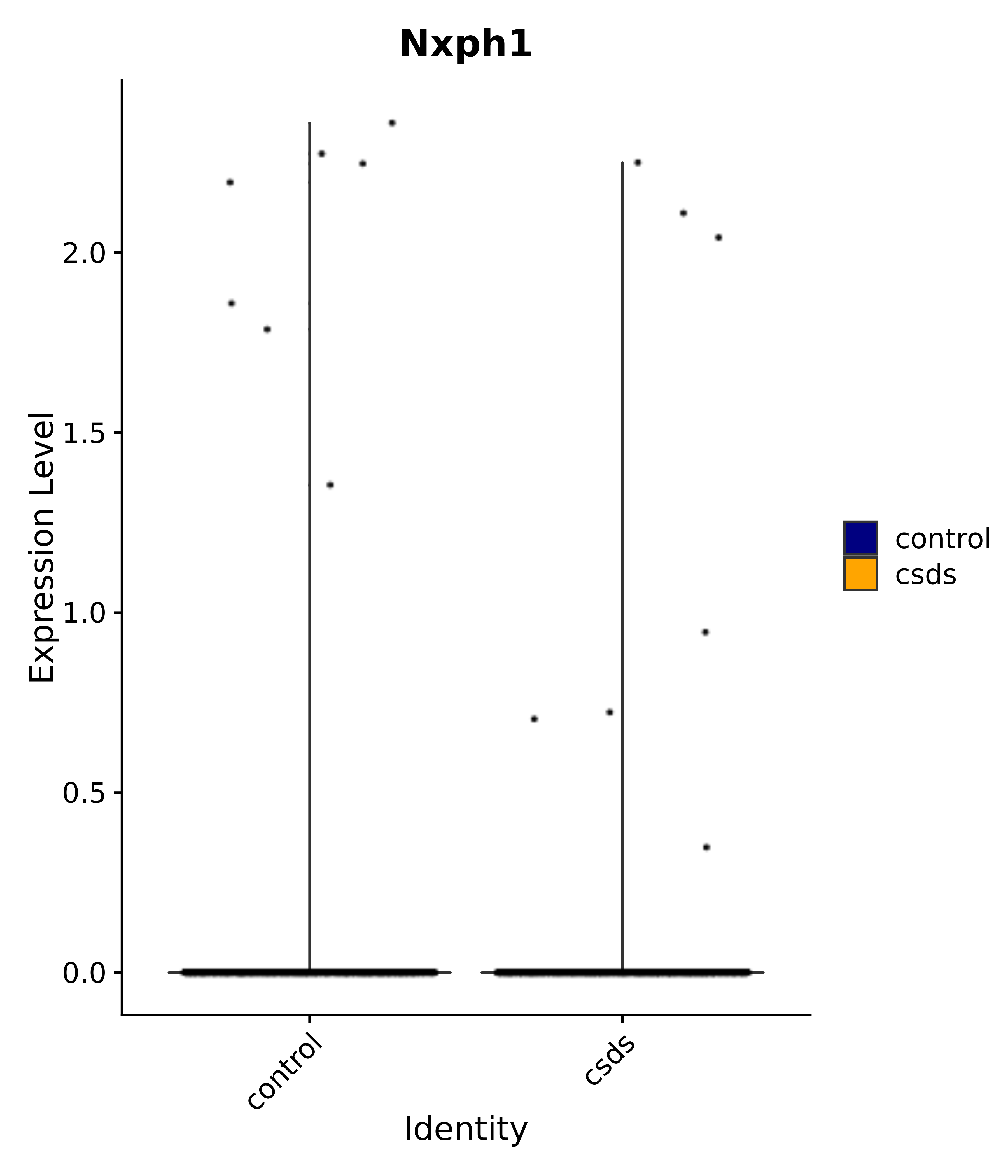
"Nxph1",
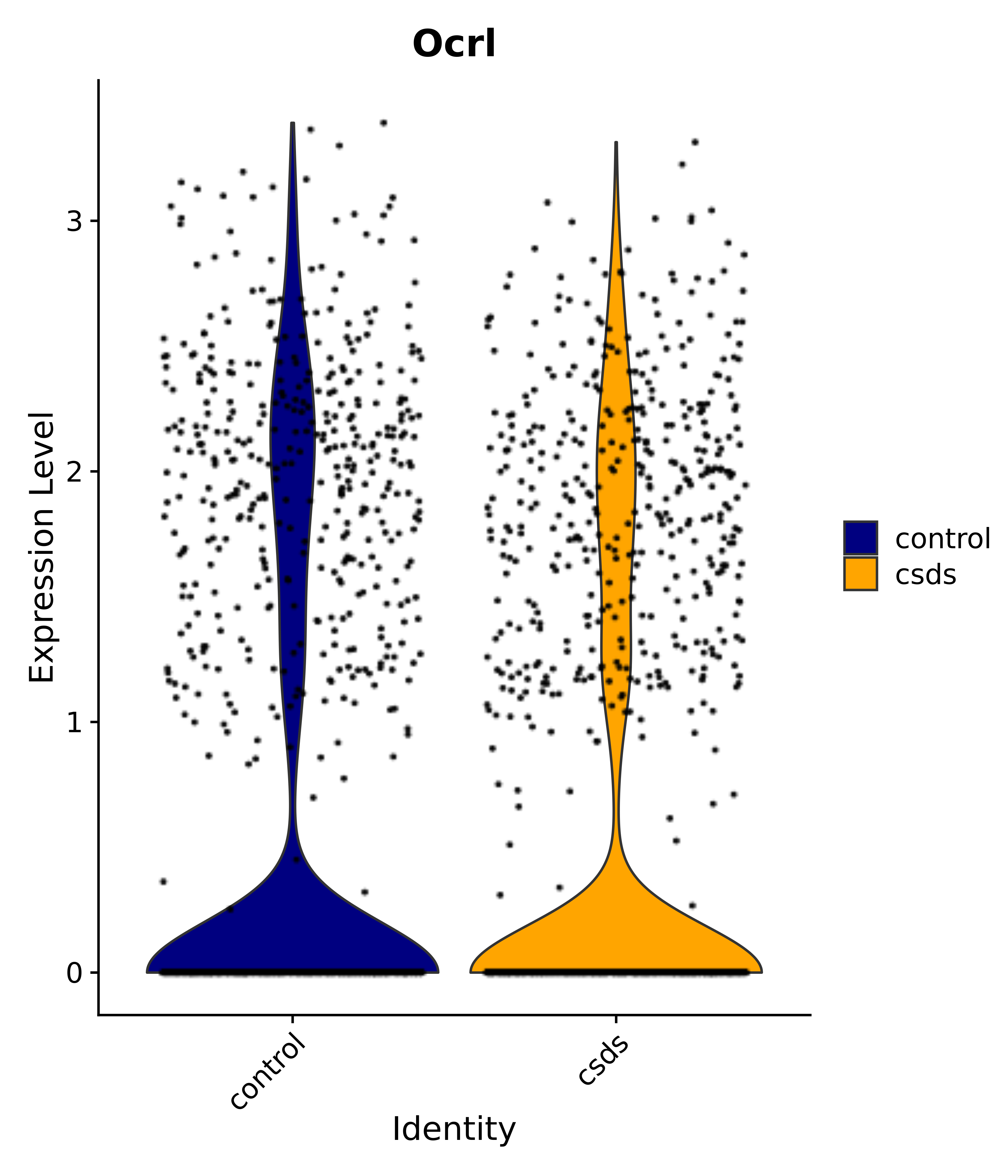
"Ocrl",
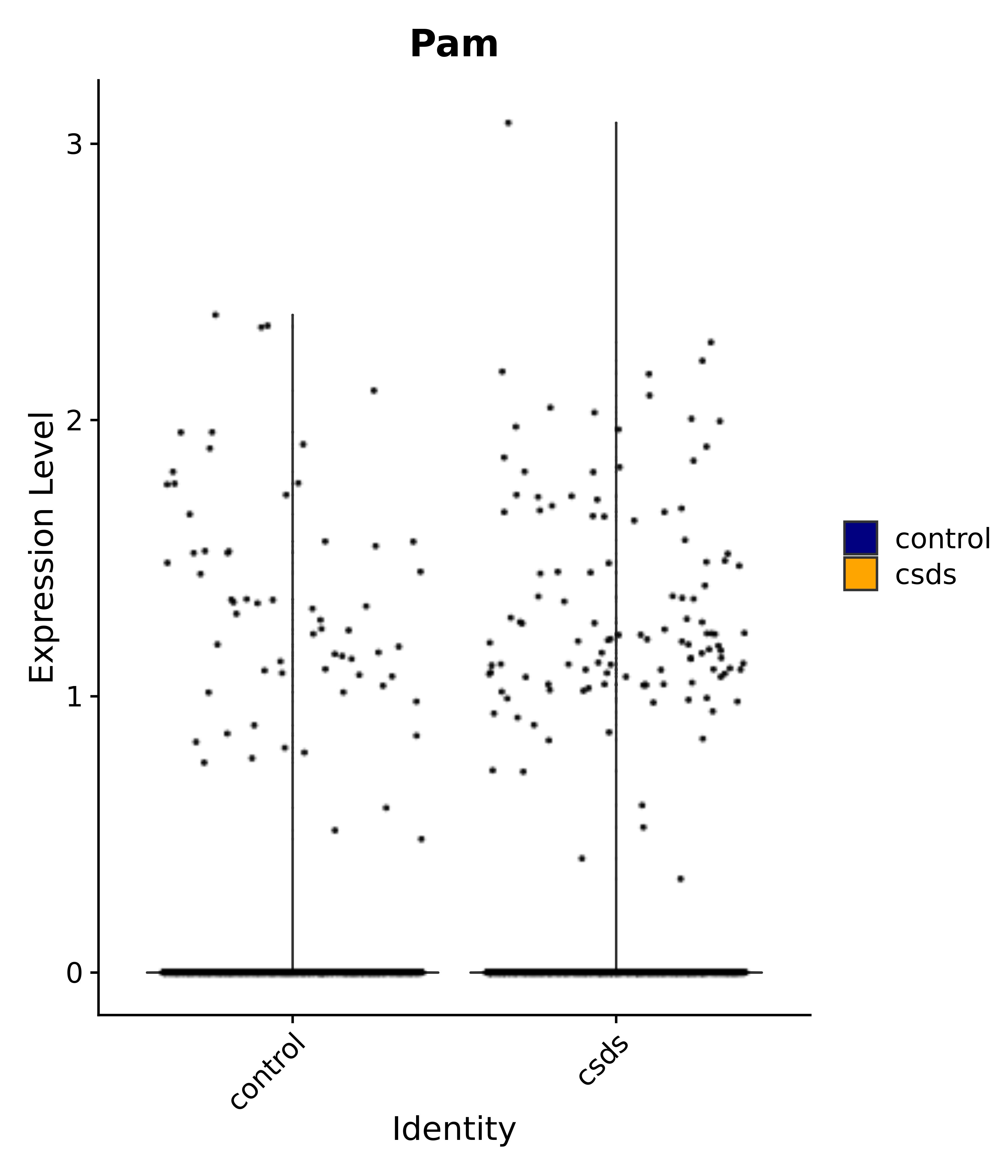
"Pam",
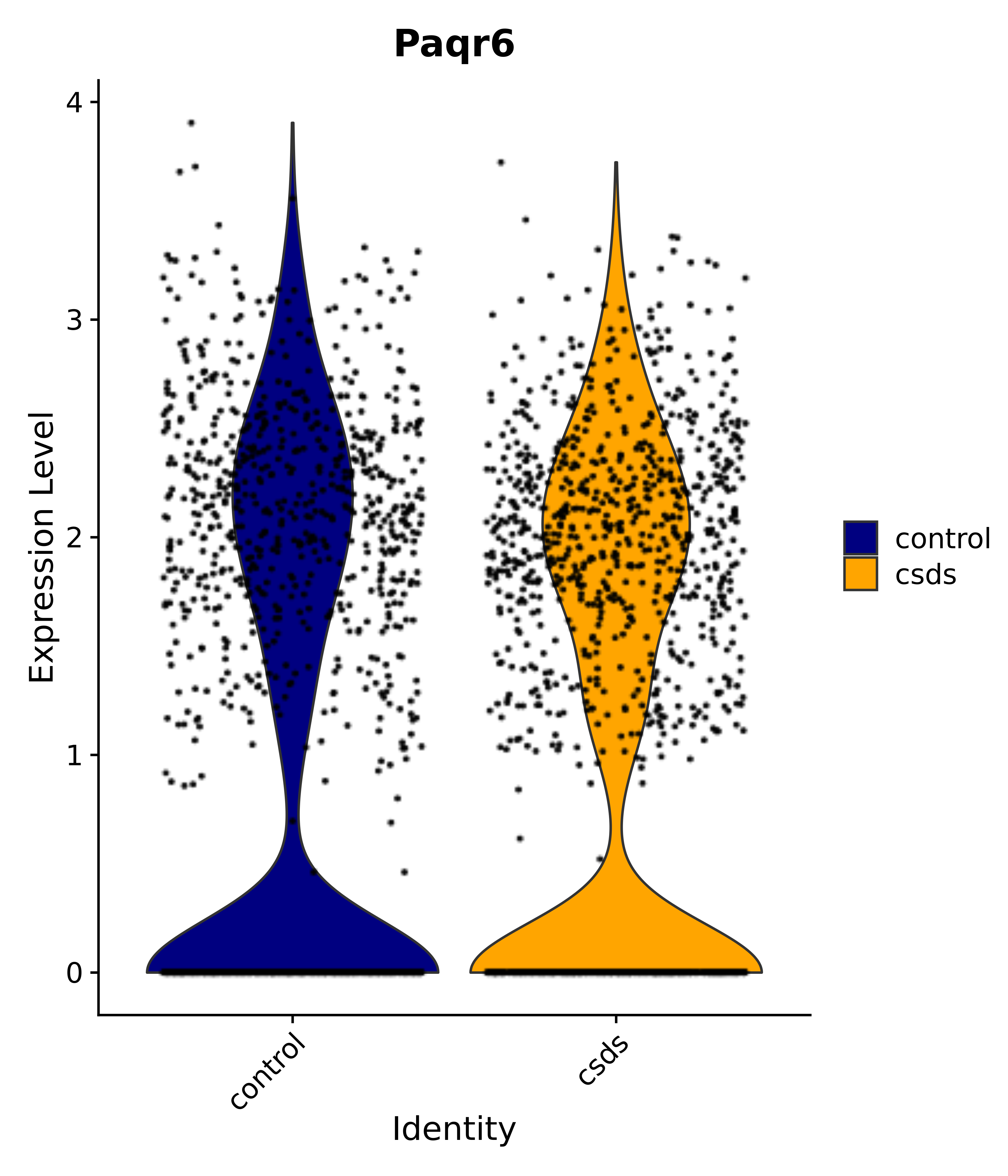
"Paqr6",
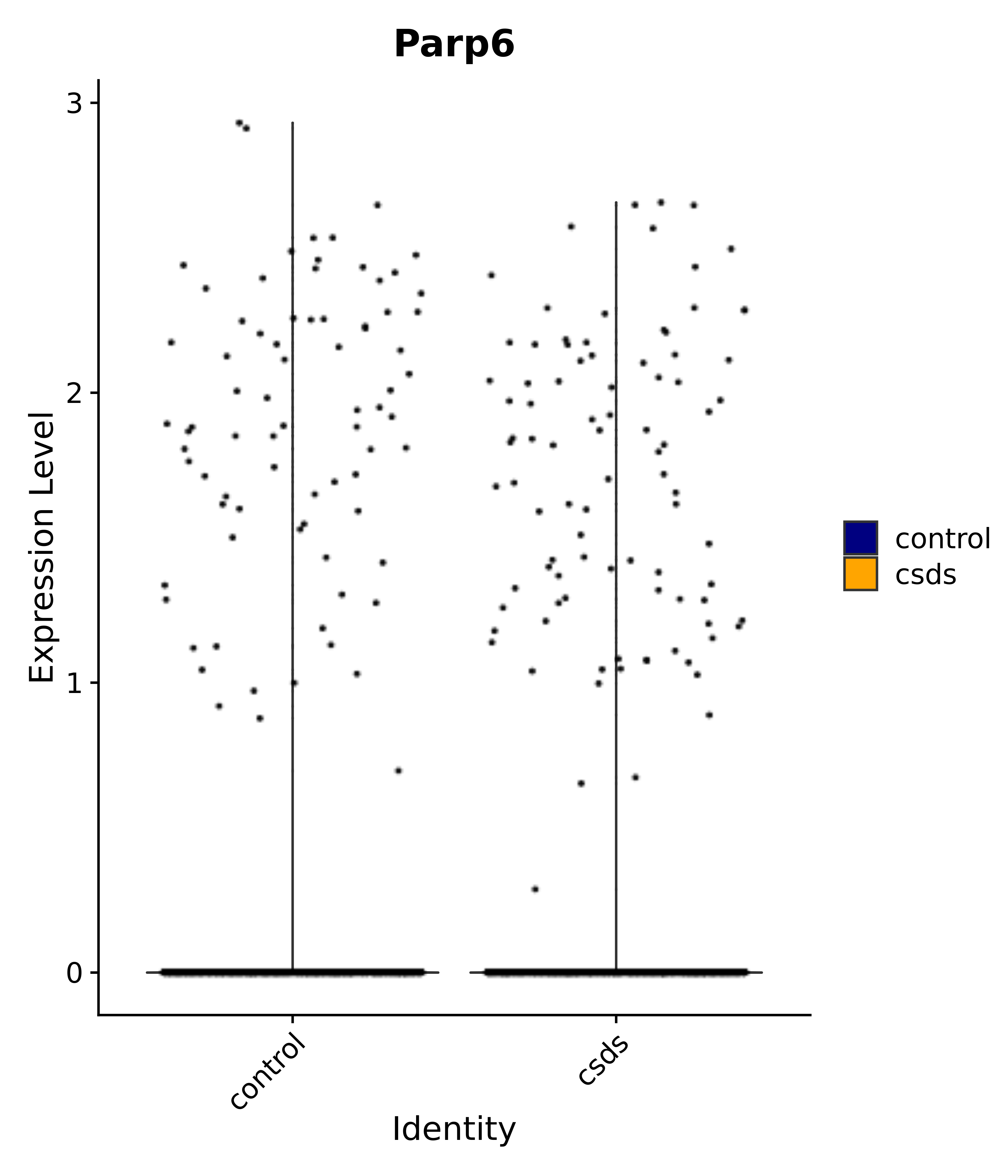
"Parp6",
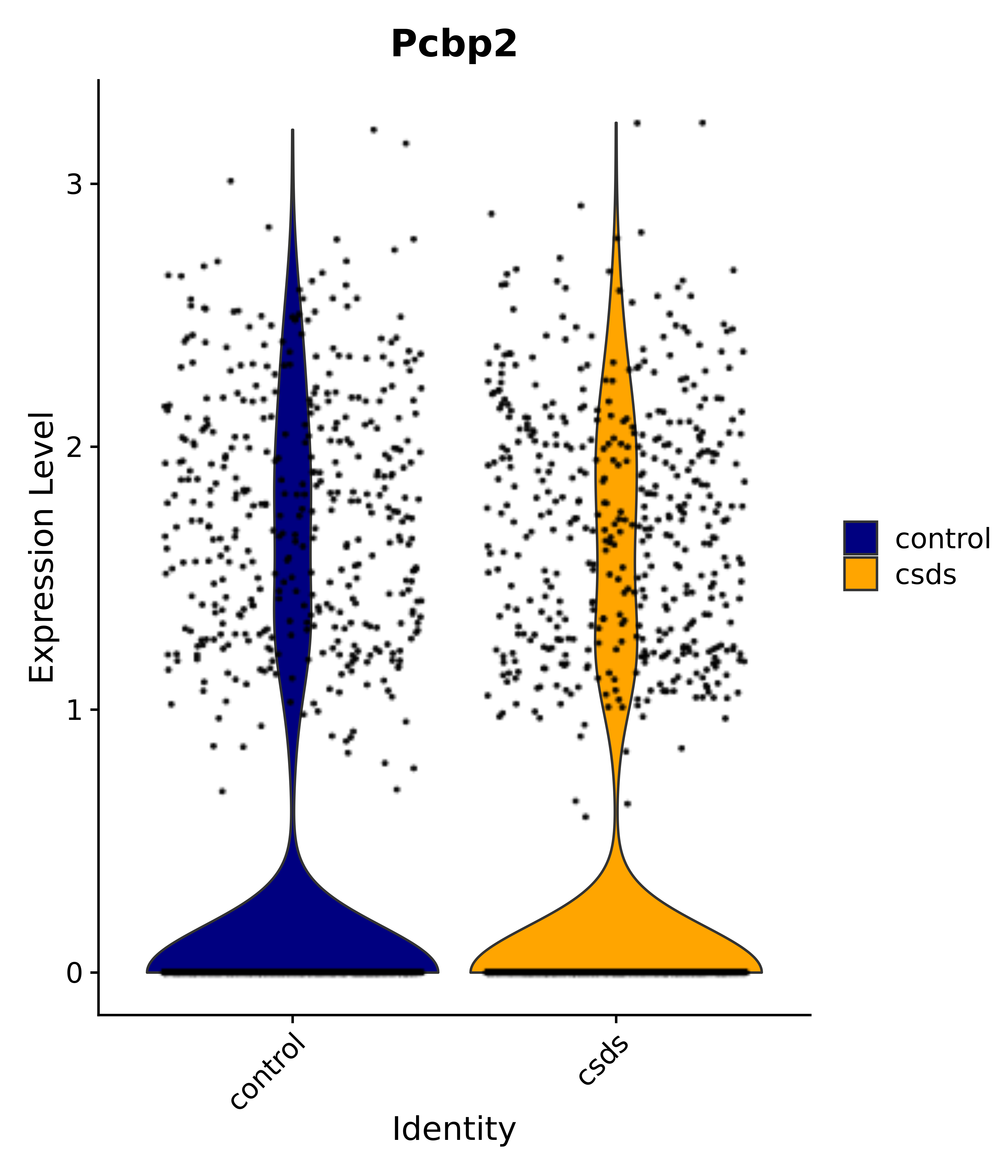
"Pcbp2",
"Pcsk1n",
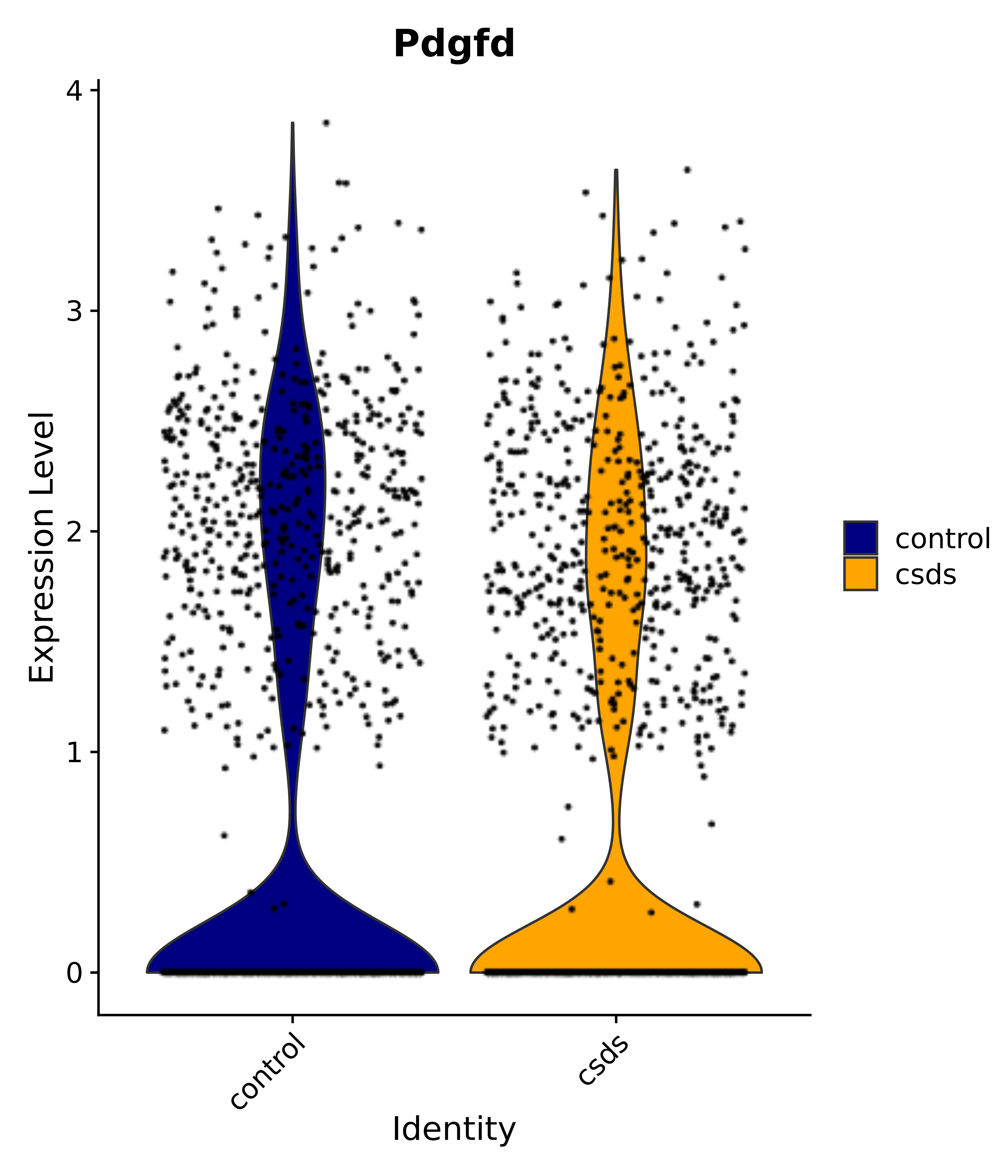
"Pdgfd",
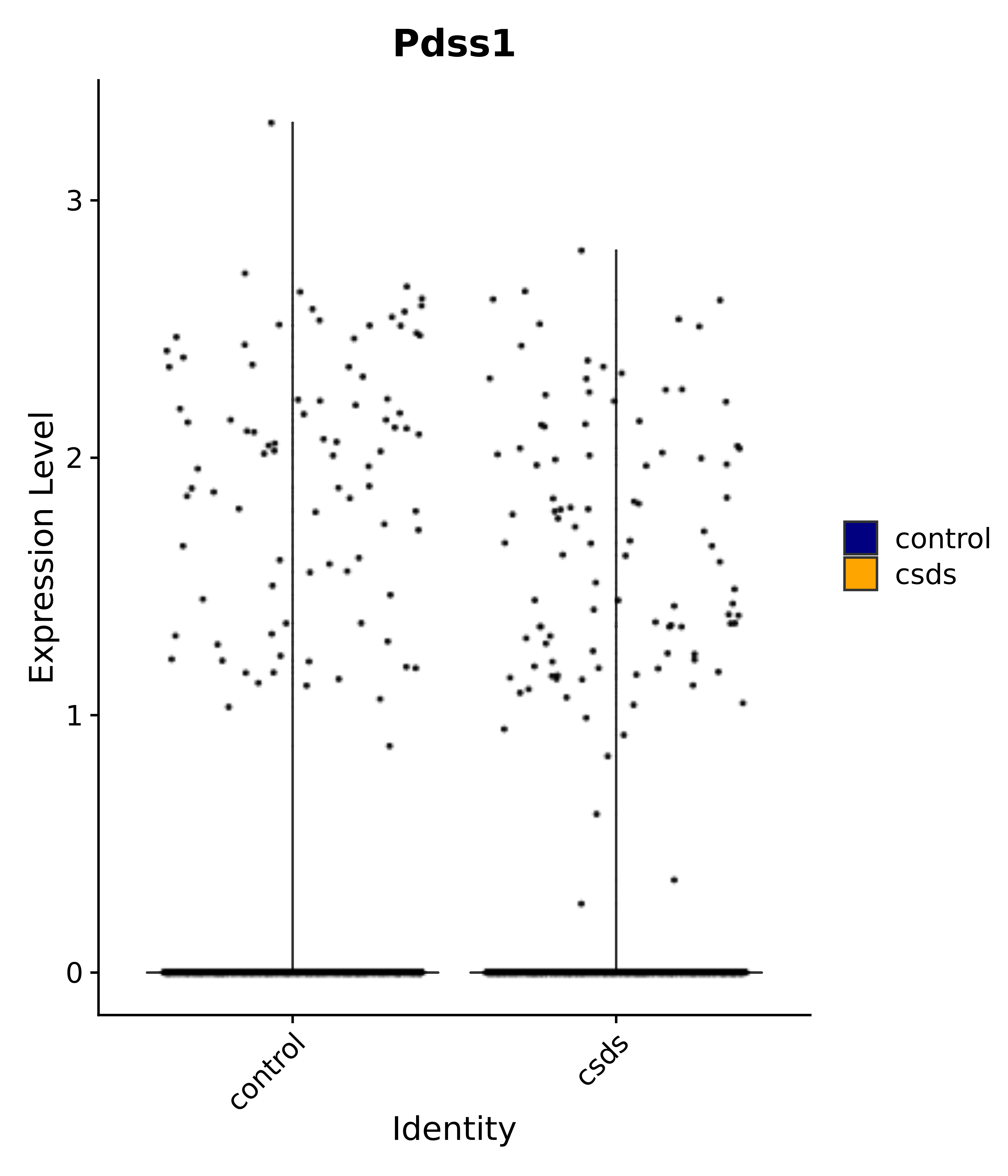
"Pdss1",
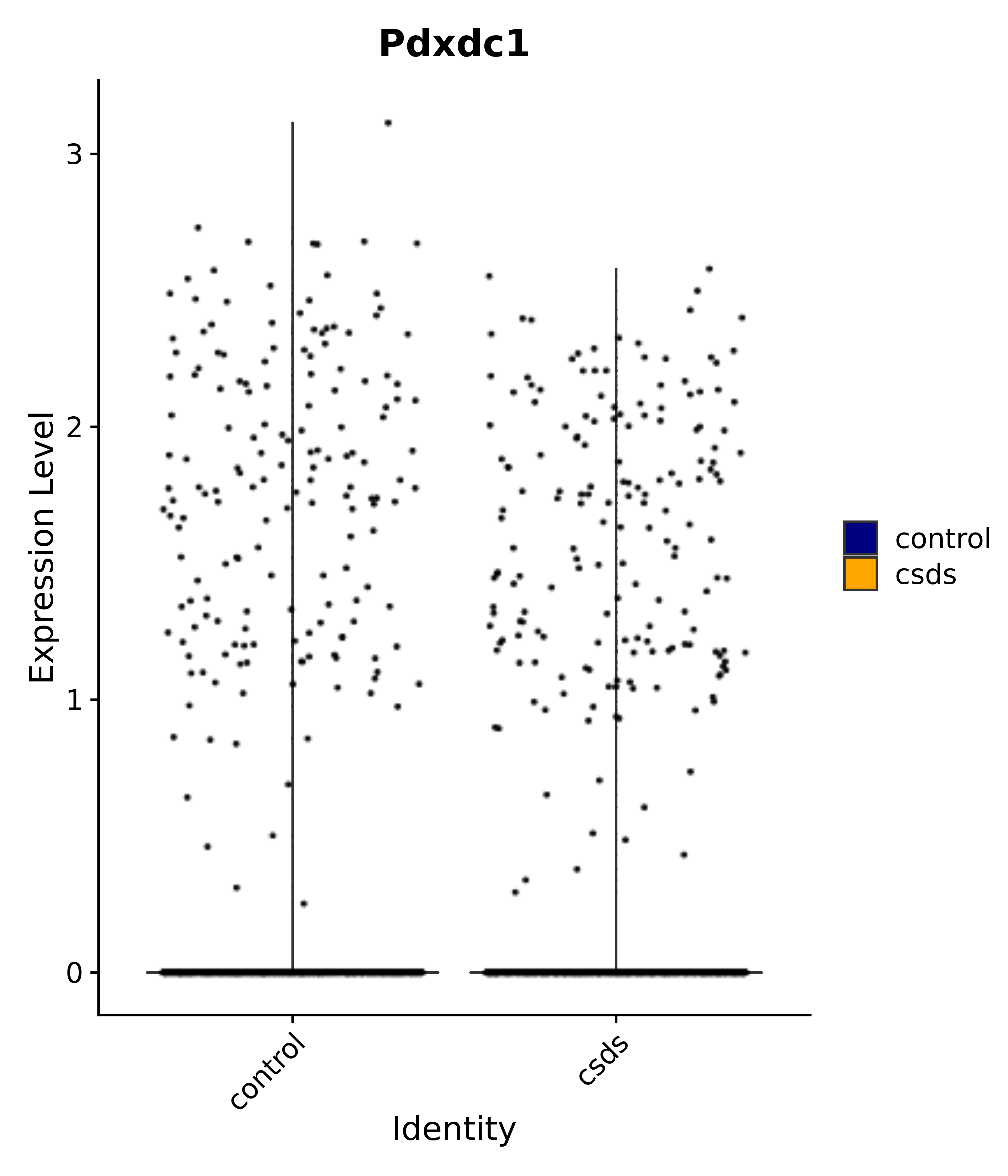
"Pdxdc1",
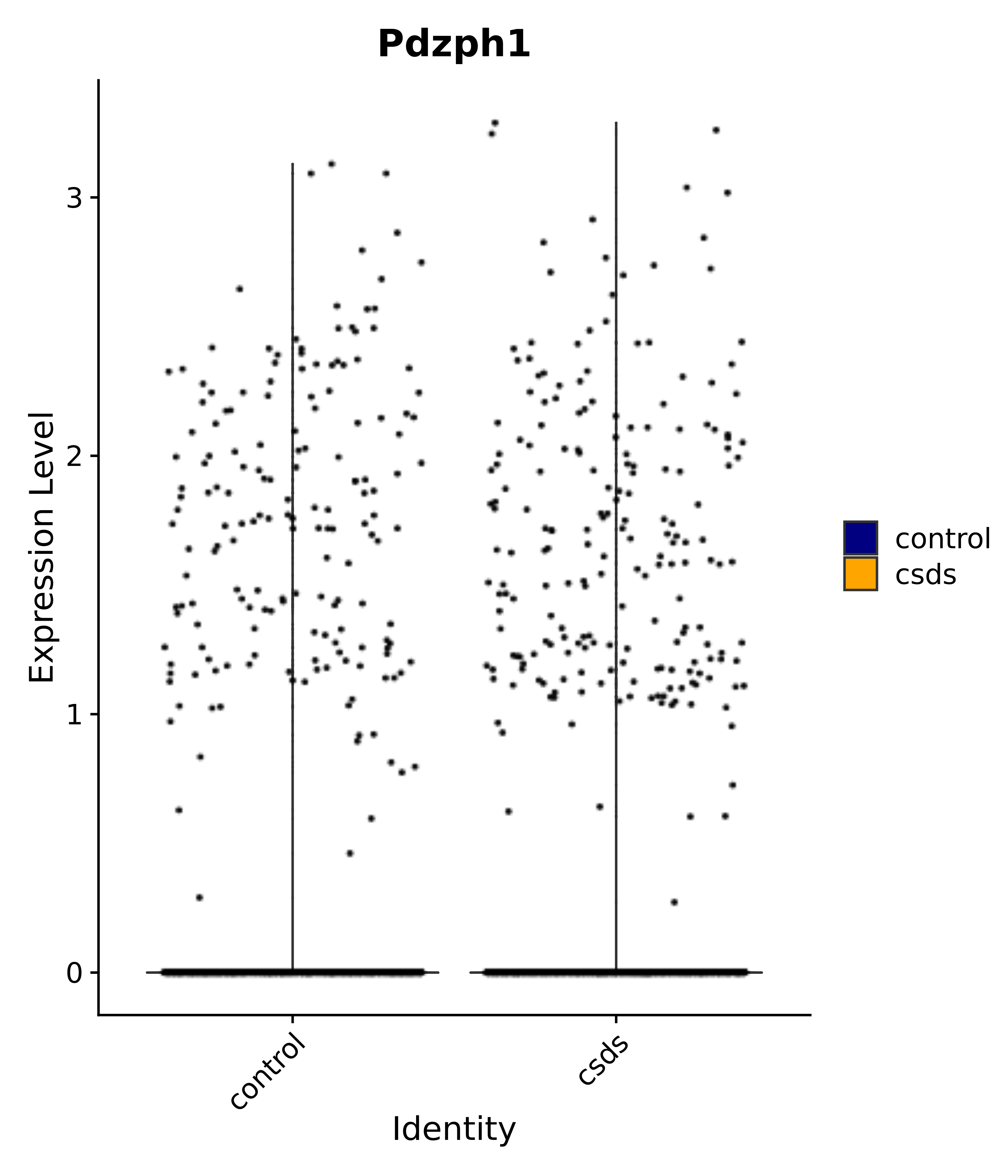
"Pdzph1",
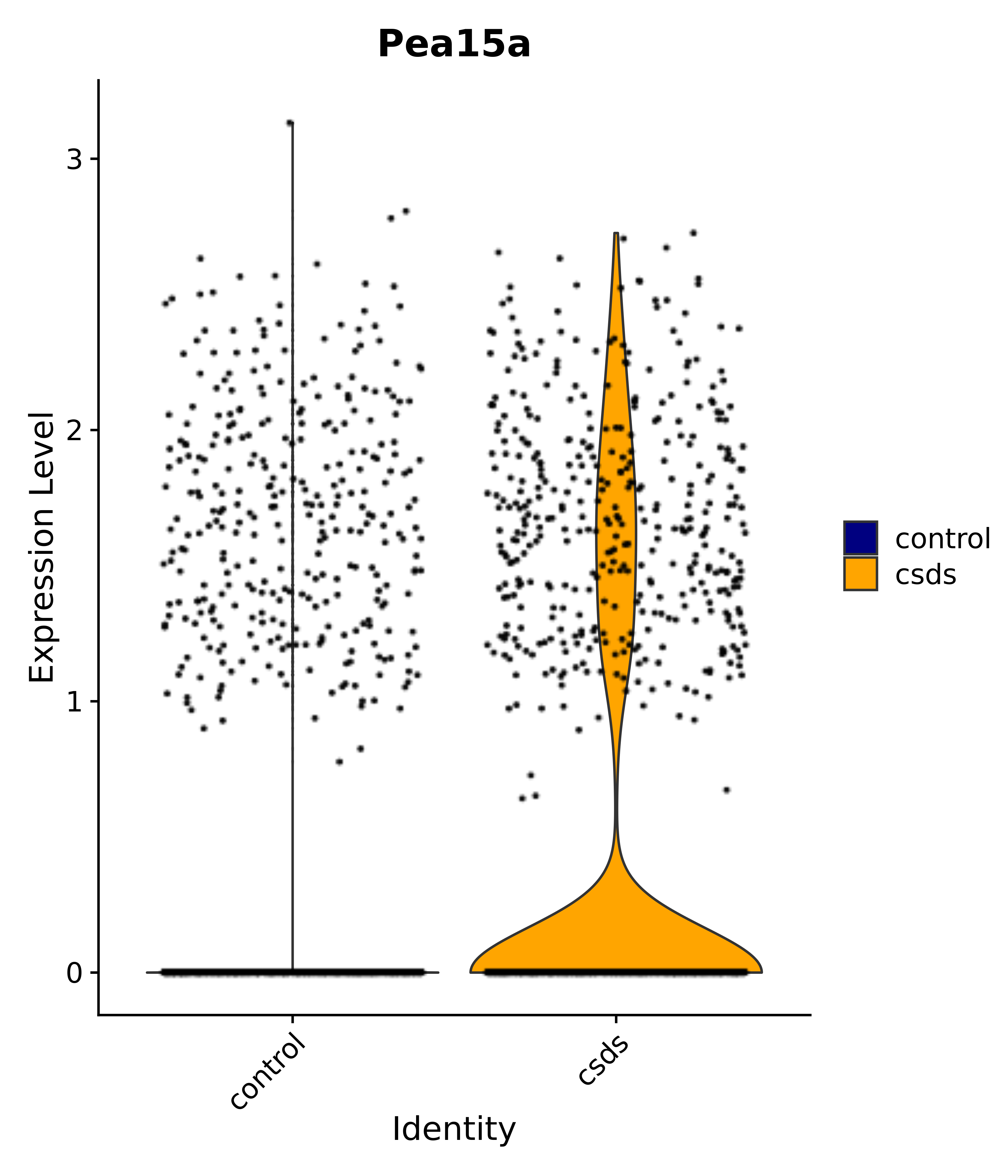
"Pea15a",
"Peg3",
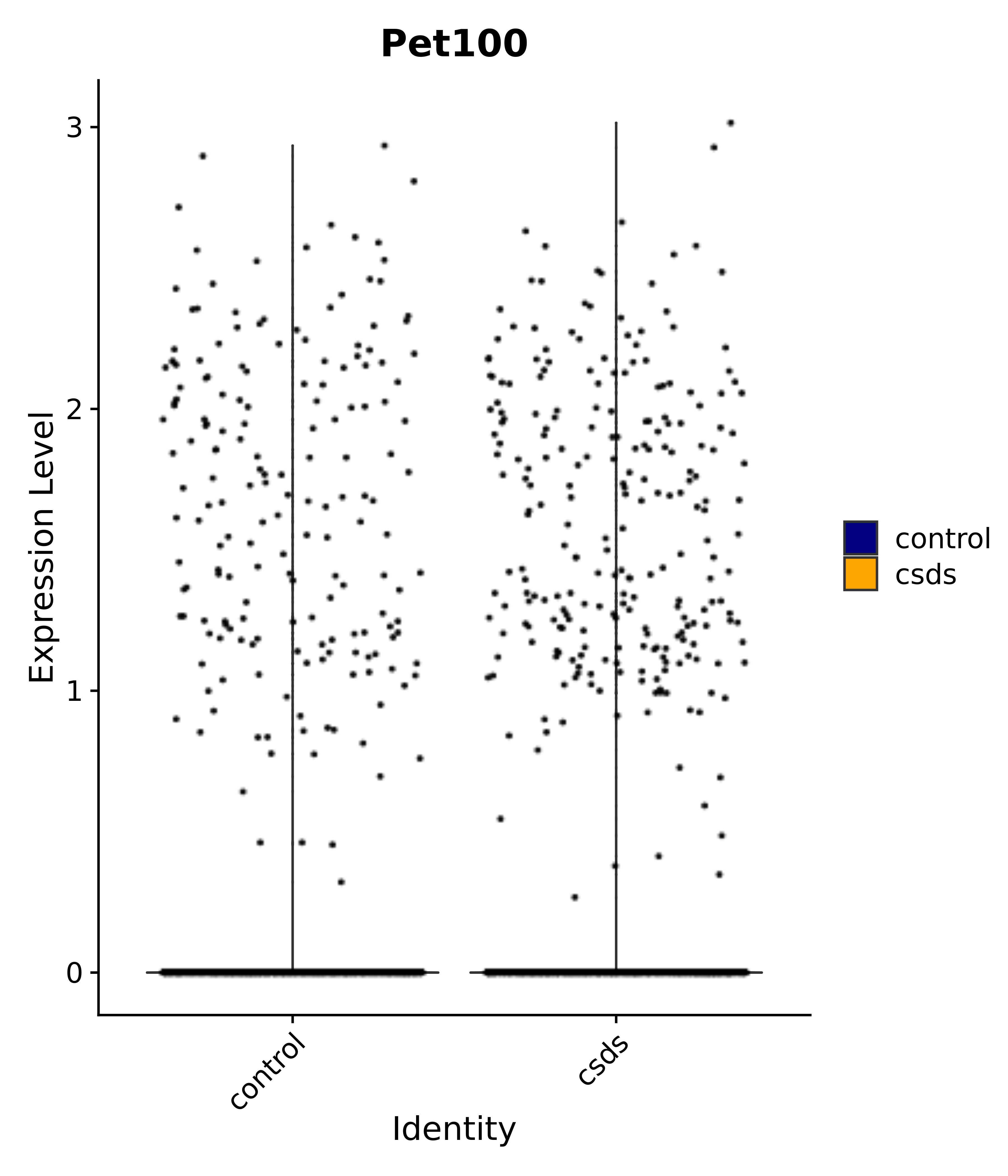
"Pet100",
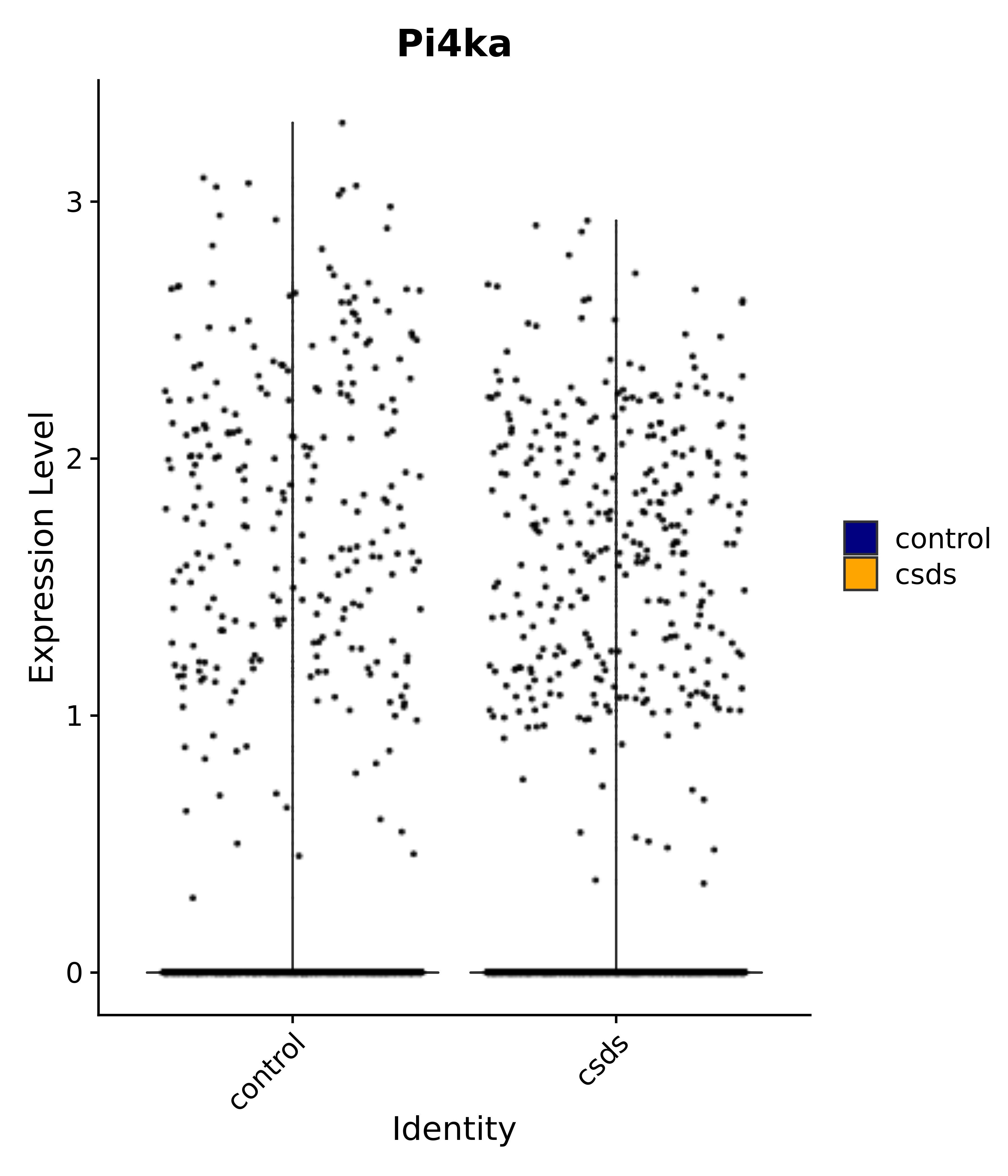
"Pi4ka",
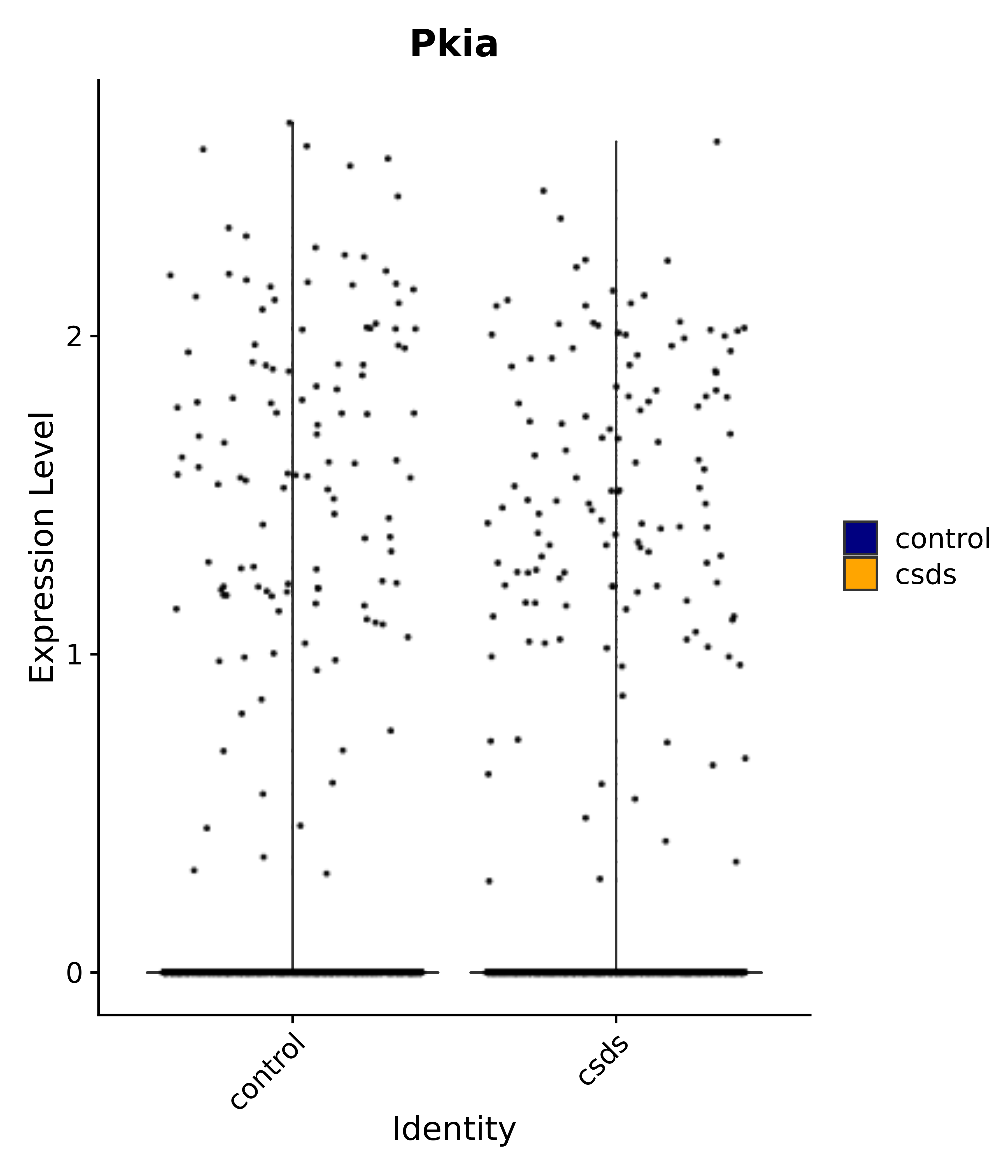
"Pkia",
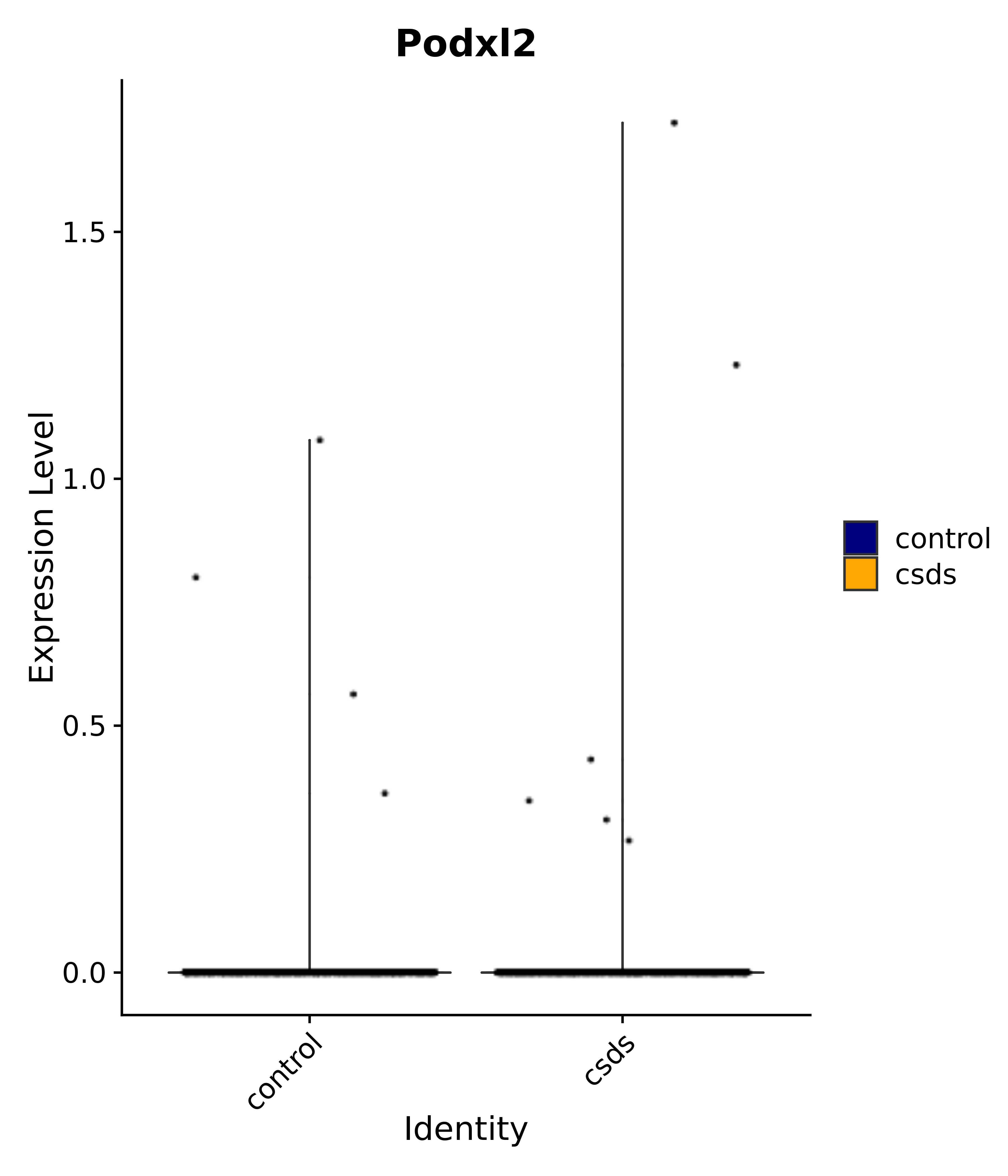
"Podxl2",
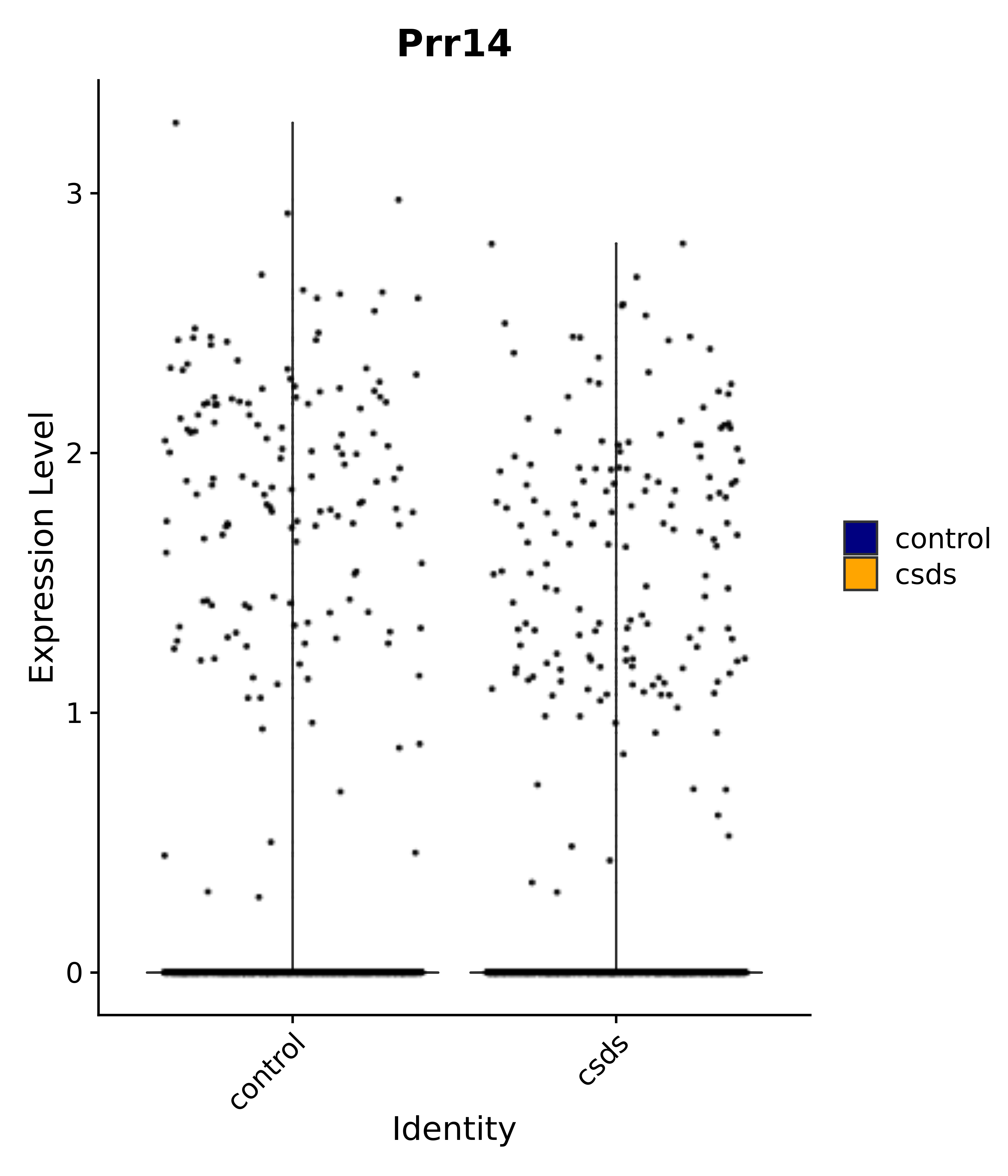
"Prr14",
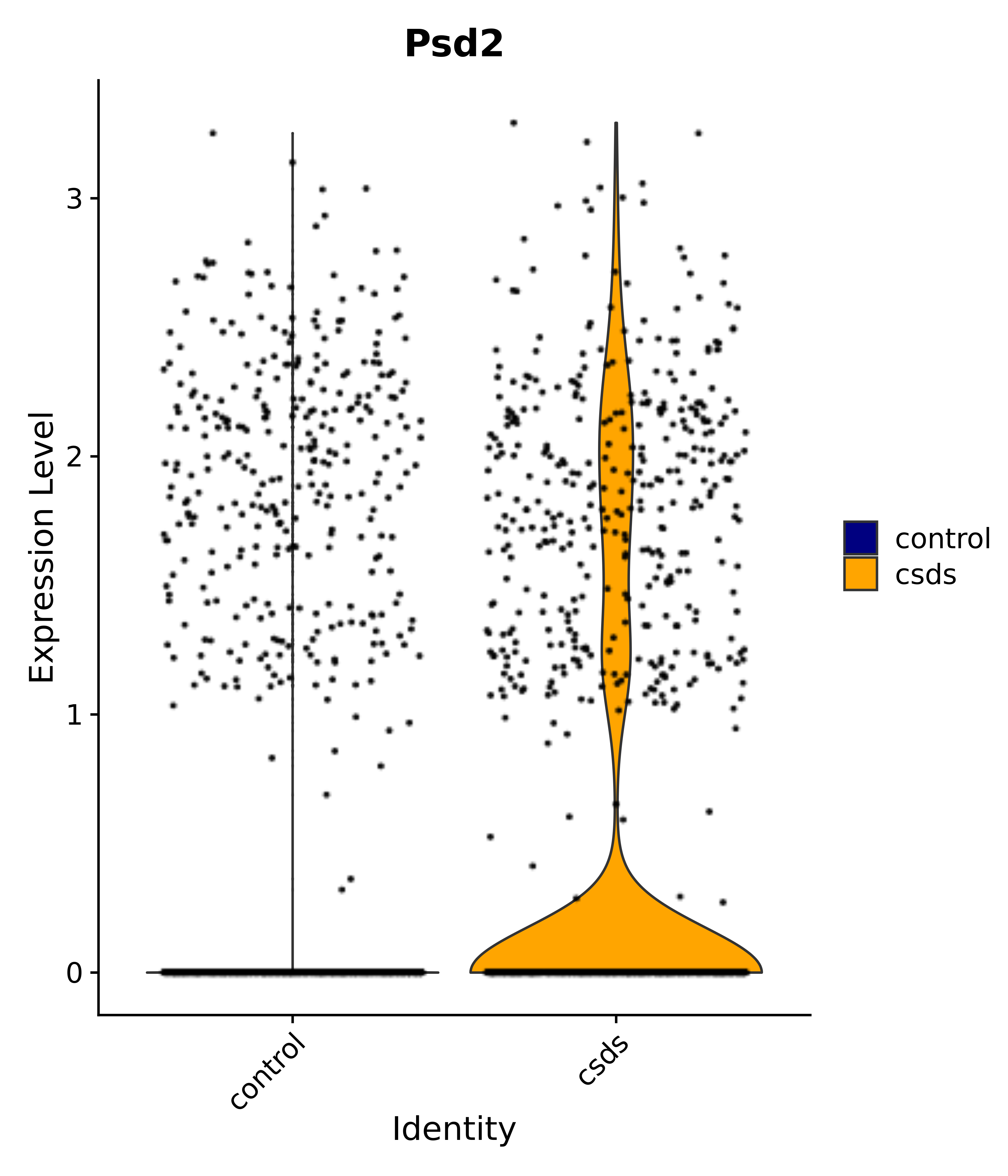
"Psd2",
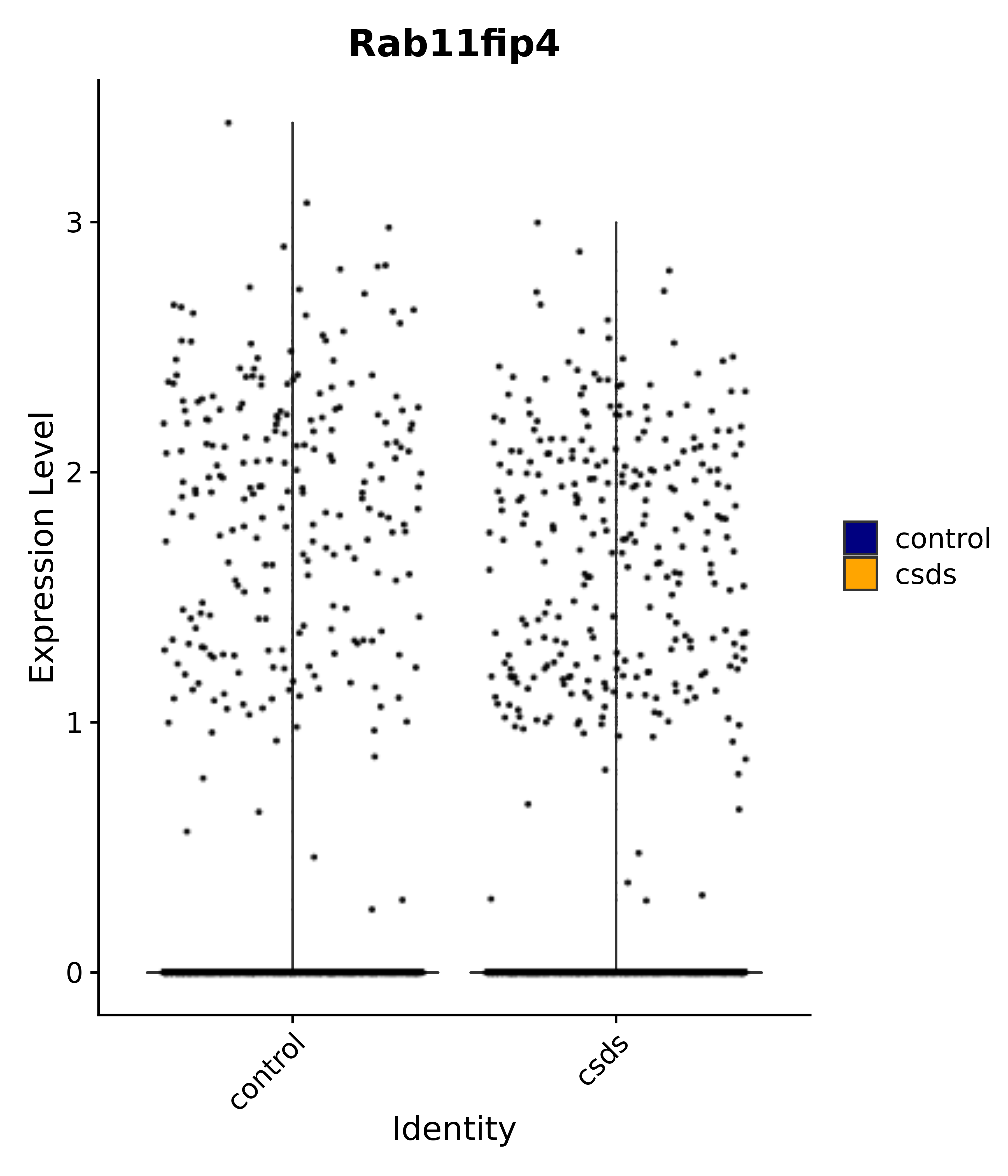
"Rab11fip4",
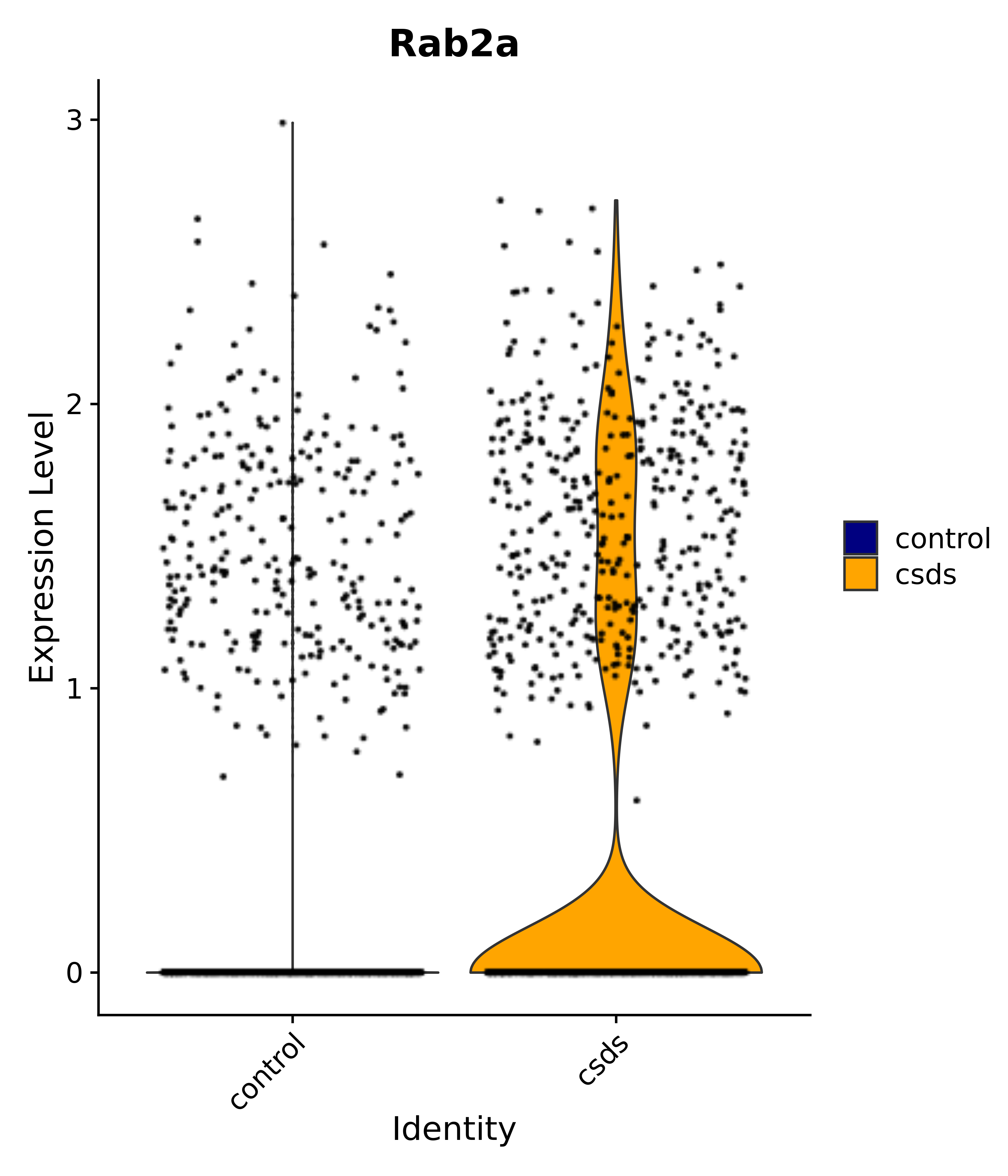
"Rab2a",
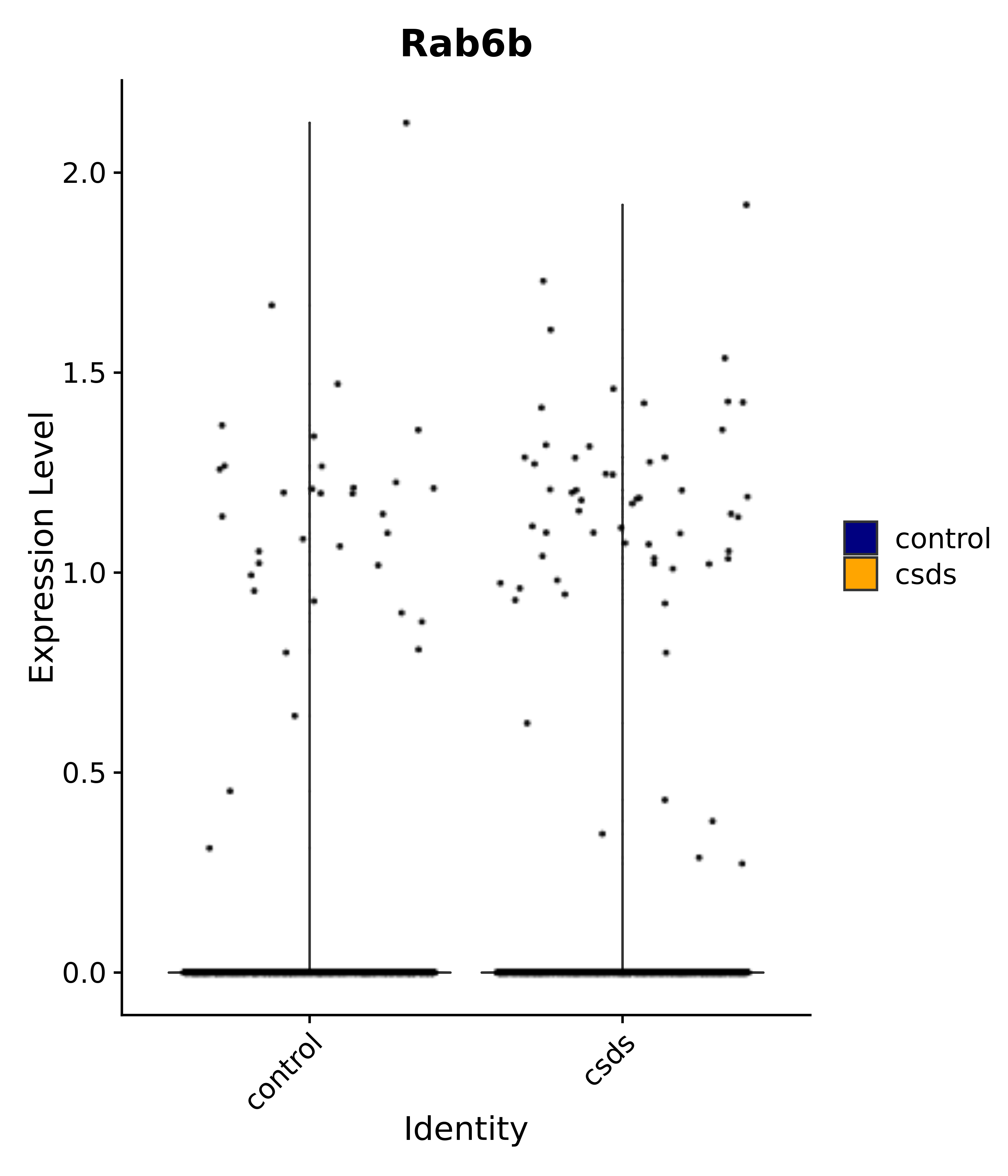
"Rab6b",
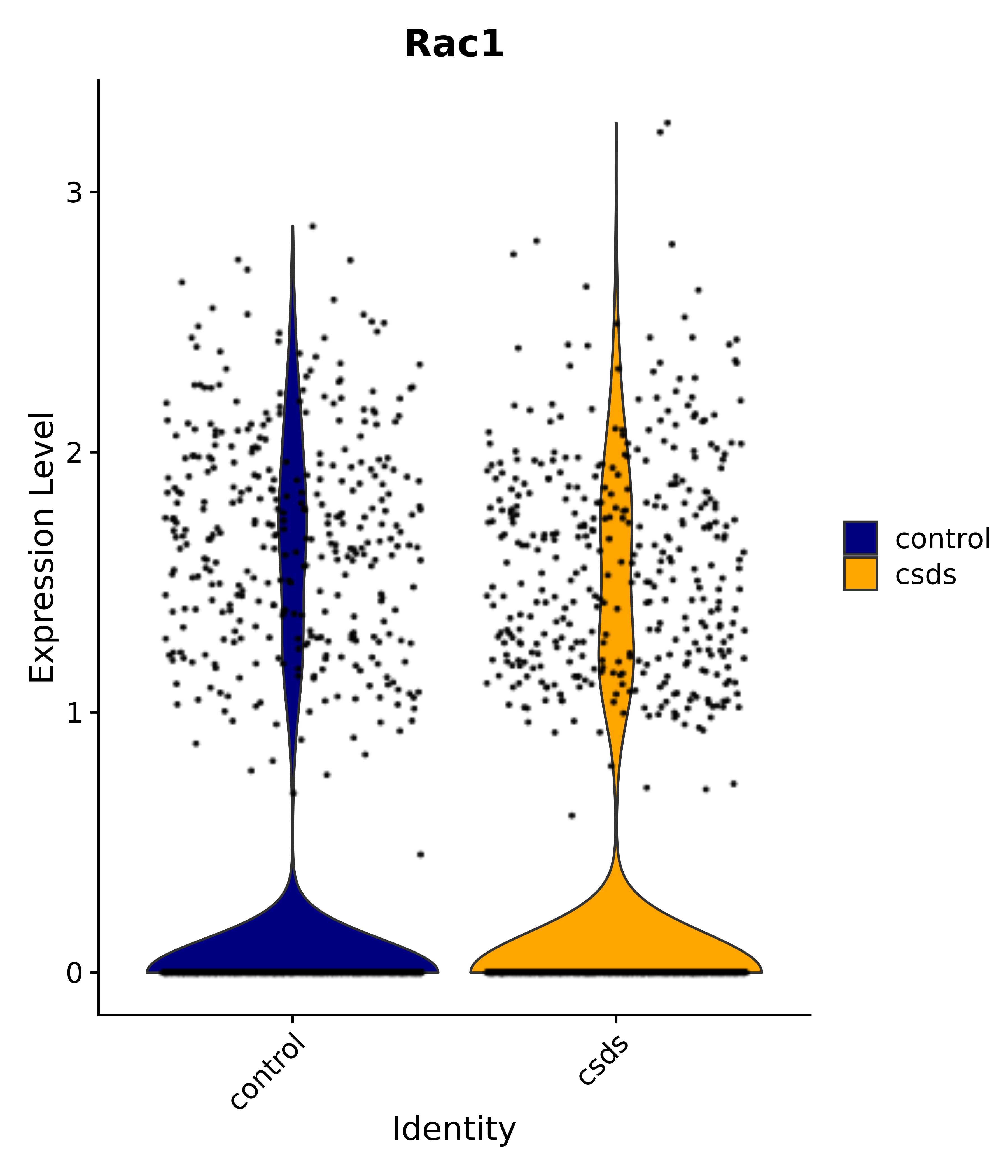
"Rac1",
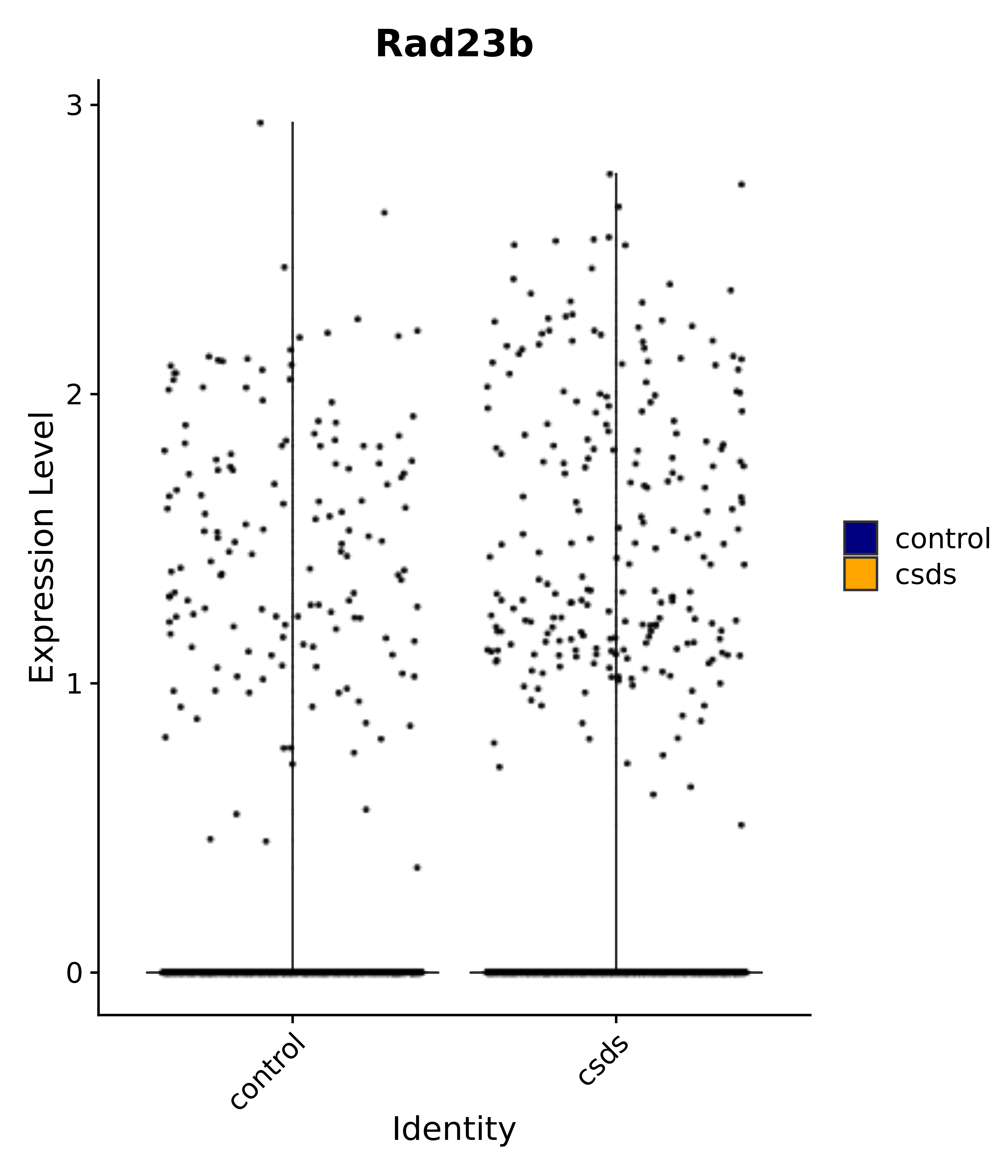
"Rad23b",
"Ralyl",
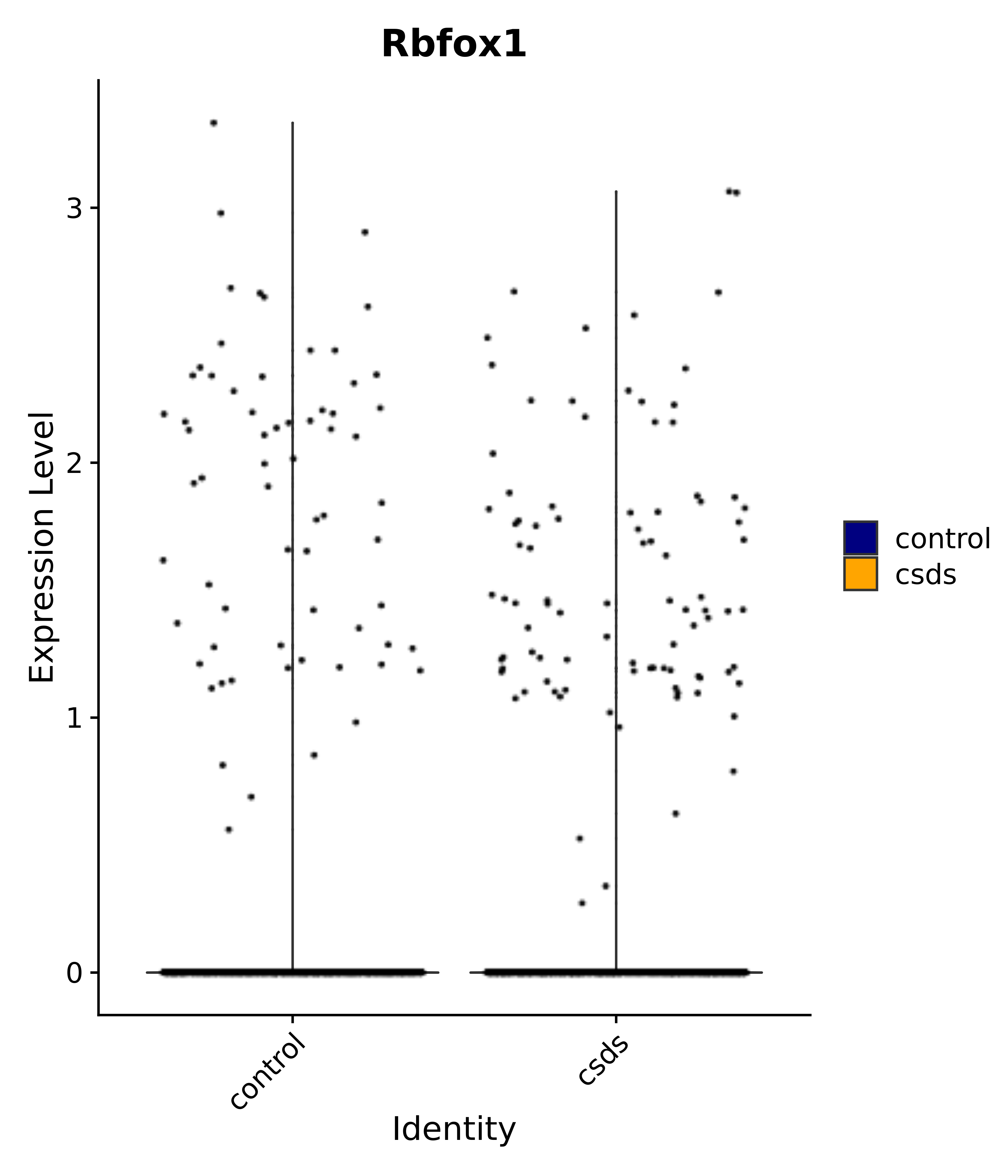
"Rbfox1",
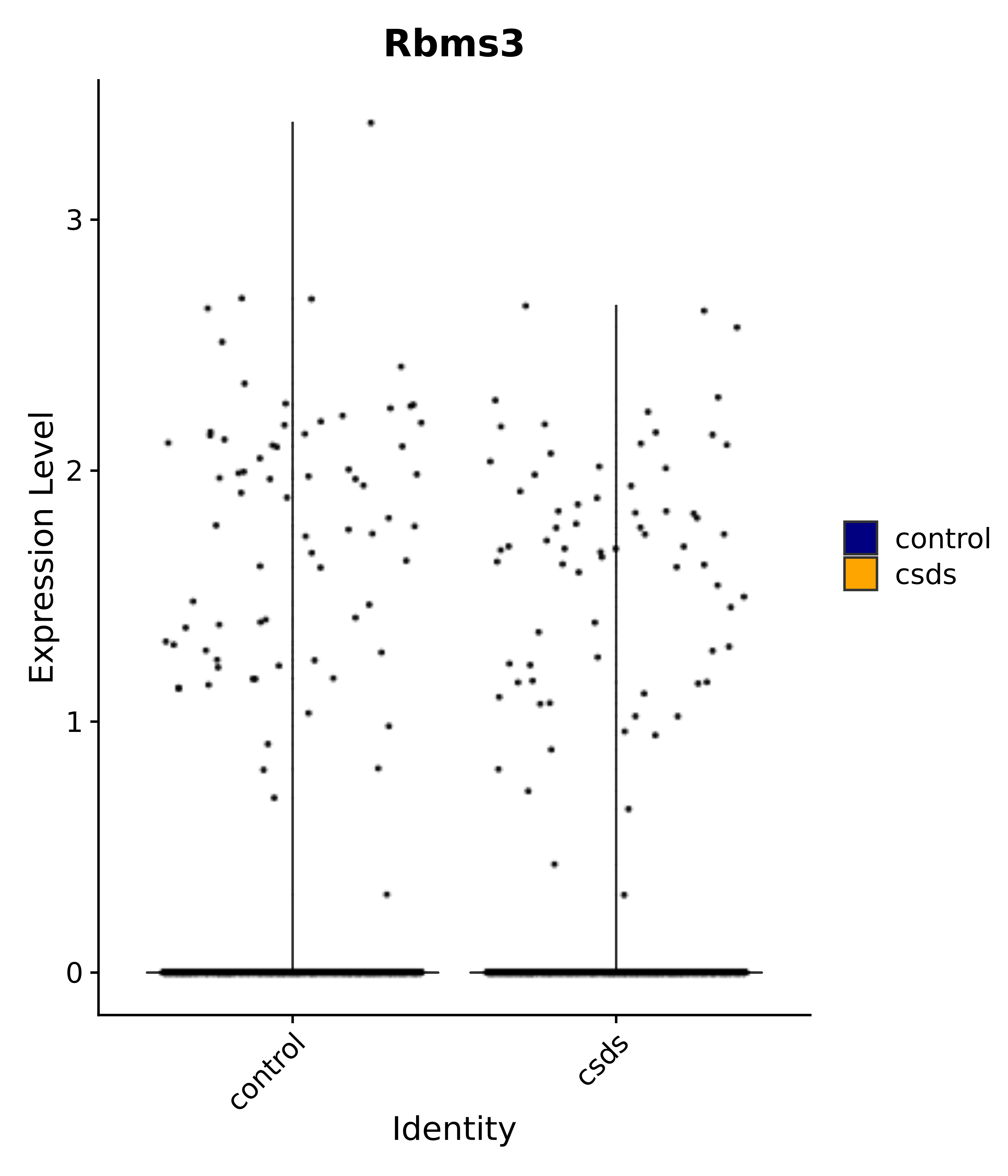
"Rbms3",
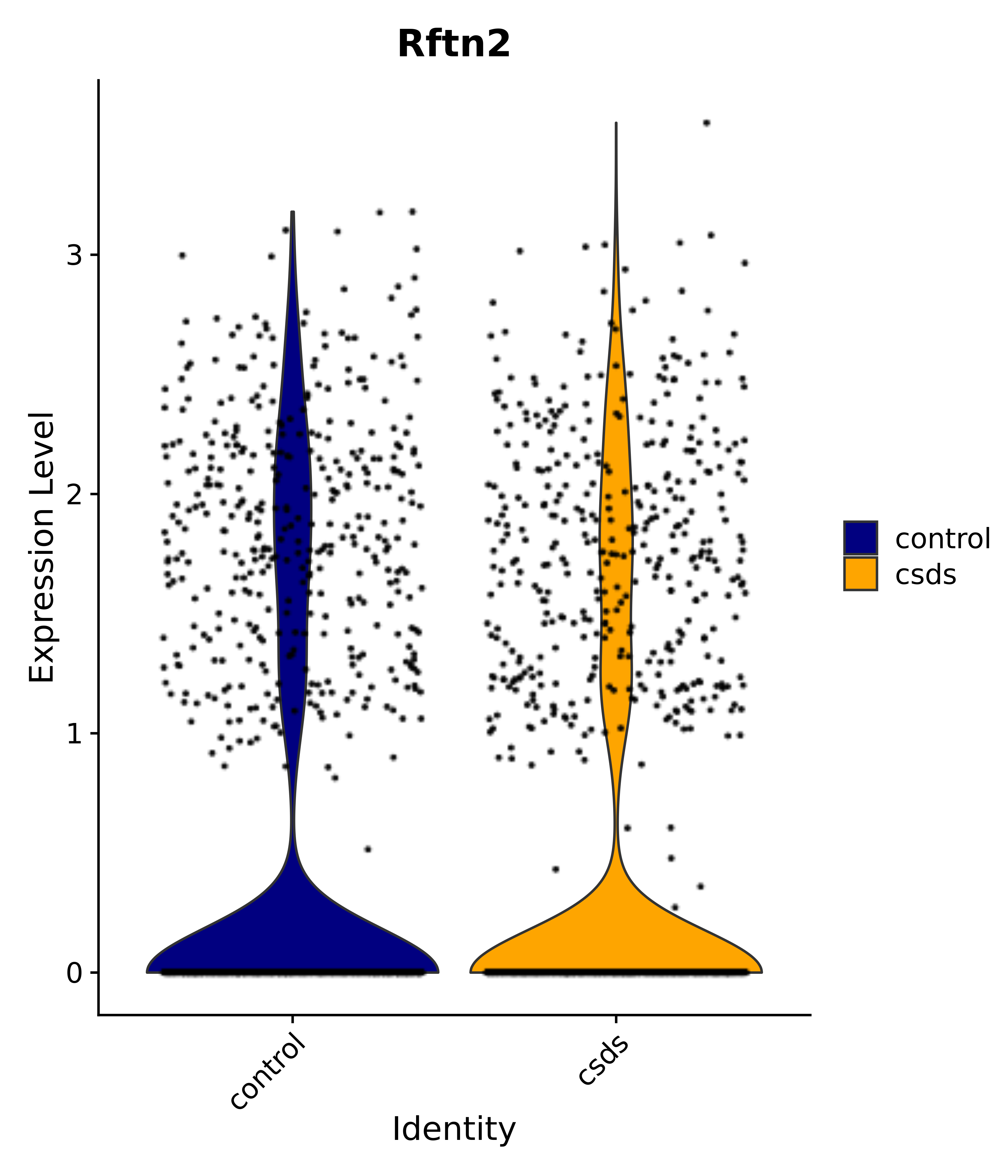
"Rftn2",
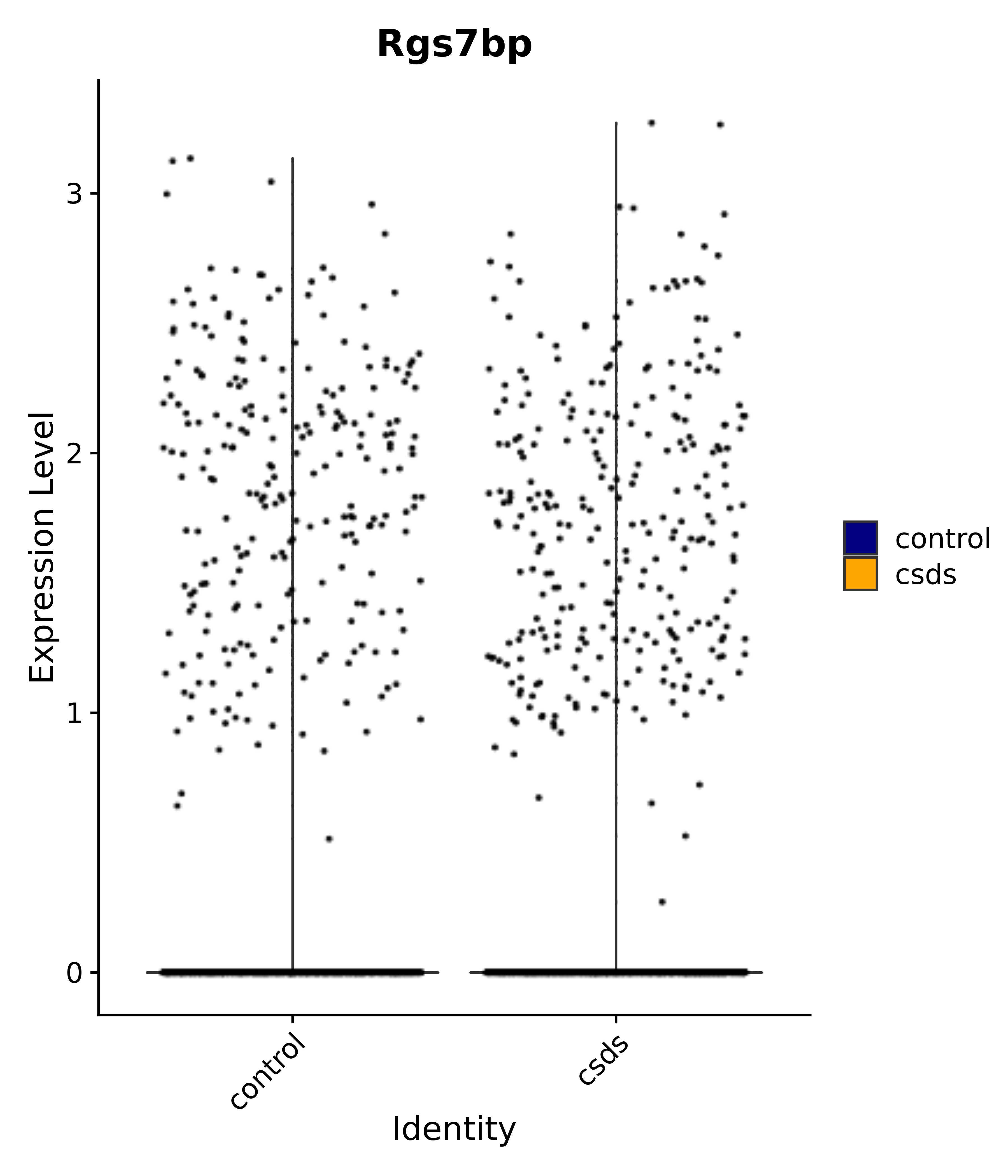
"Rgs7bp",
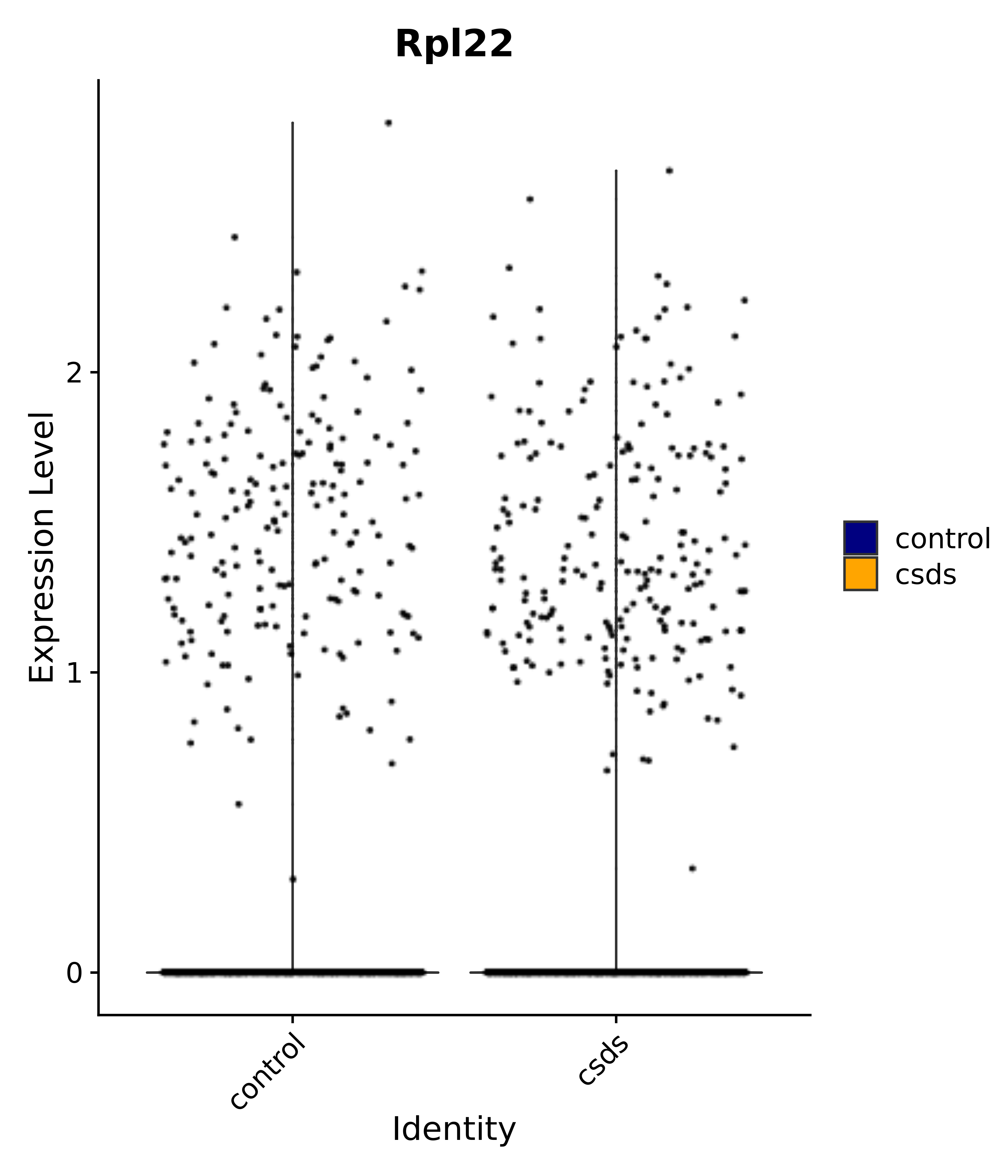
"Rpl22",
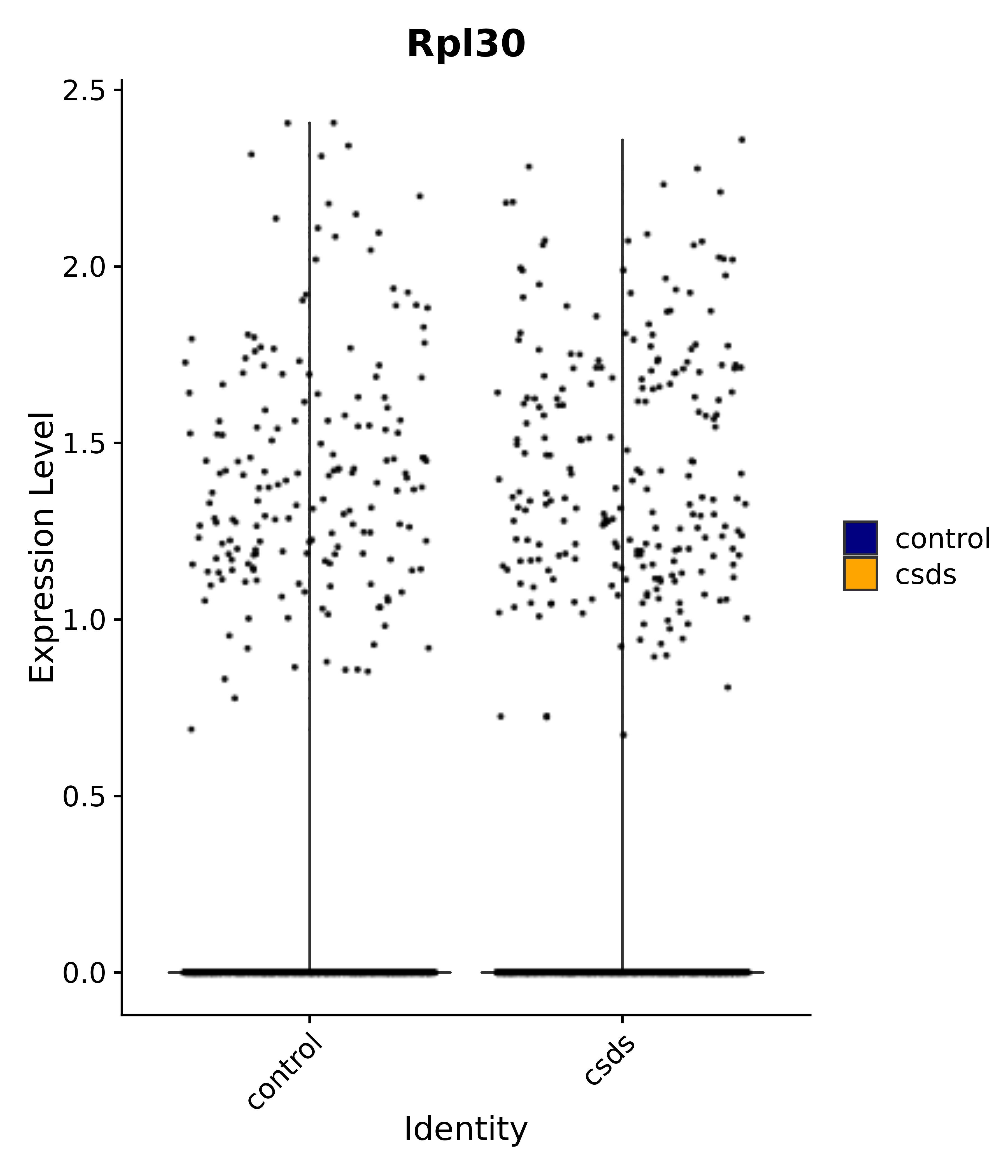
"Rpl30",
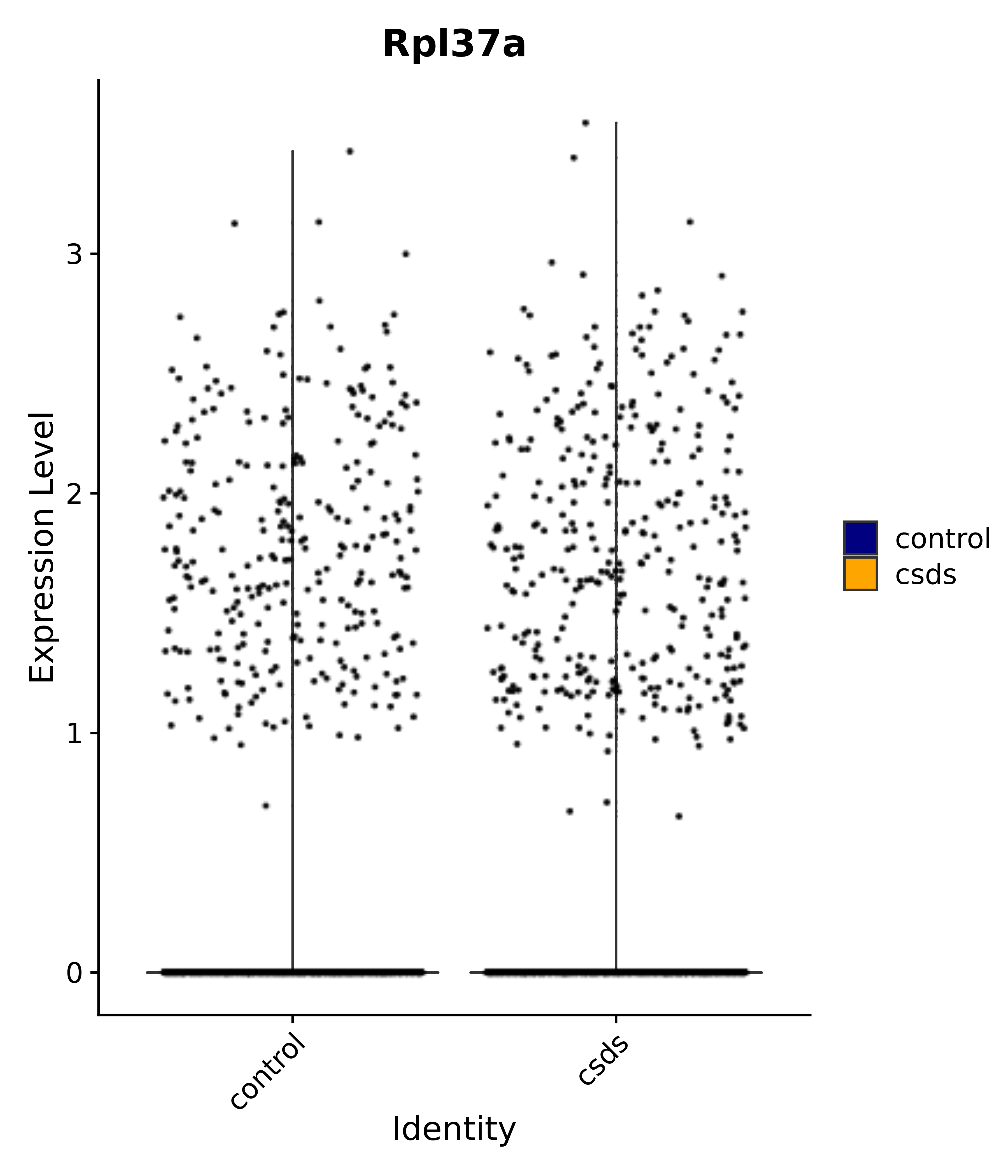
"Rpl37a",
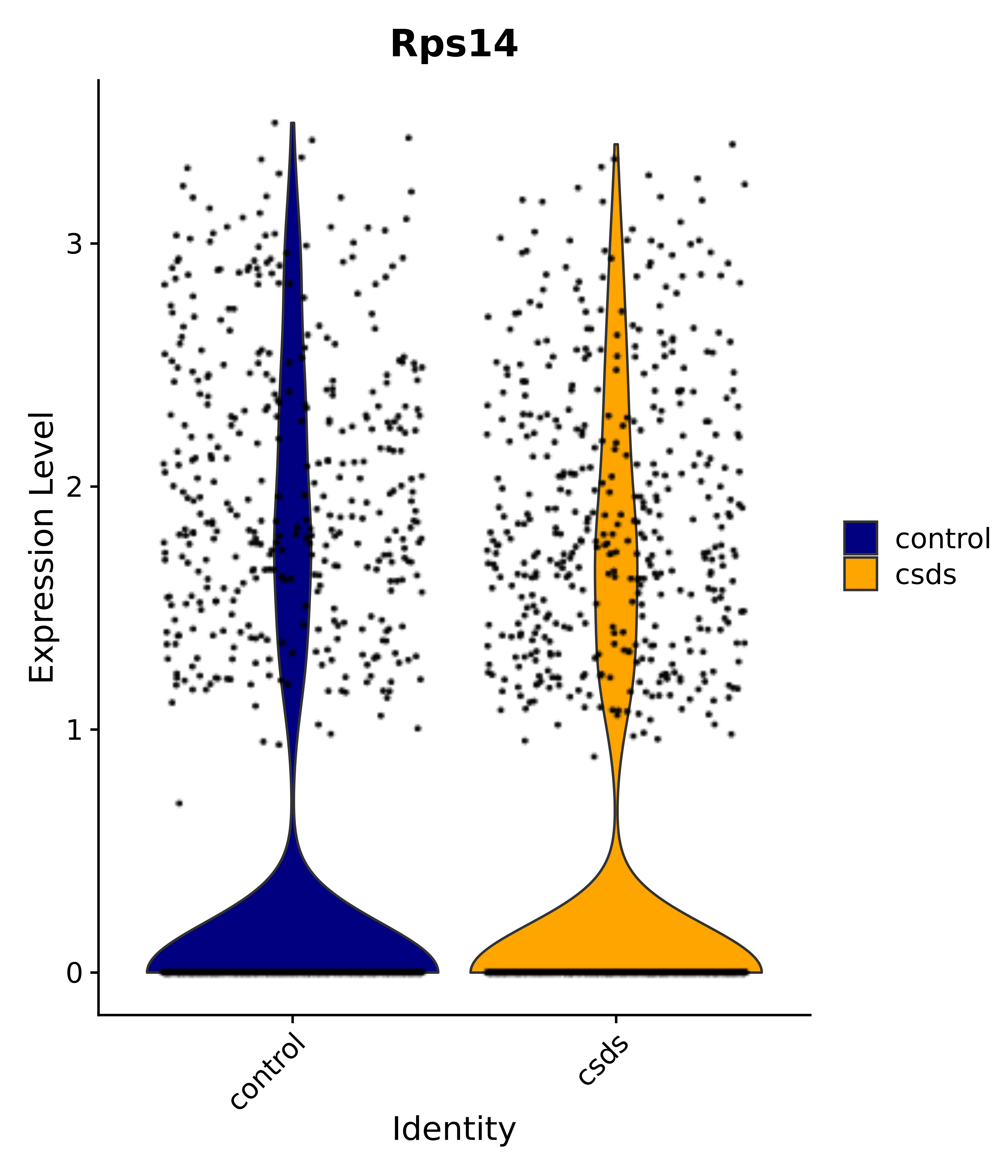
"Rps14",
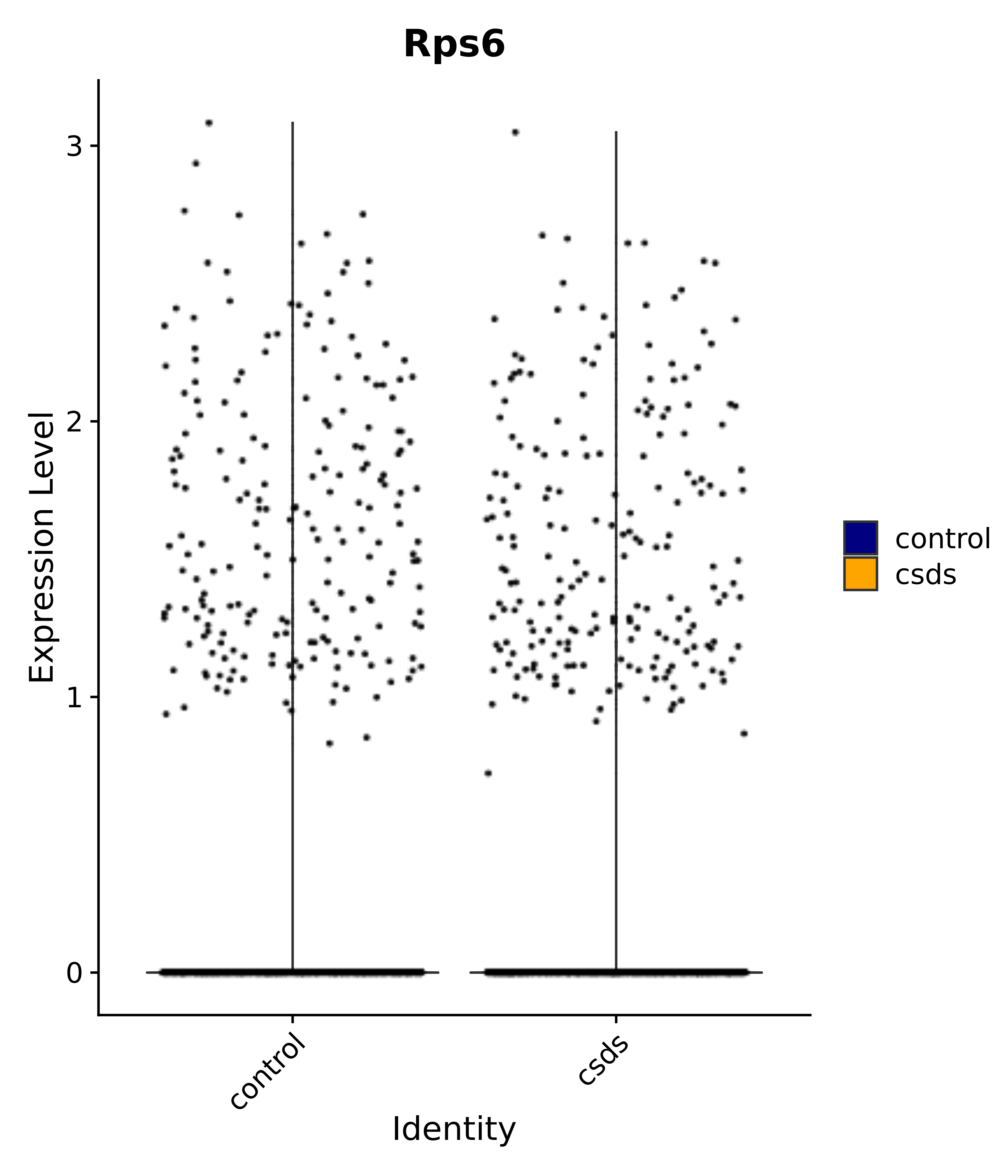
"Rps6",
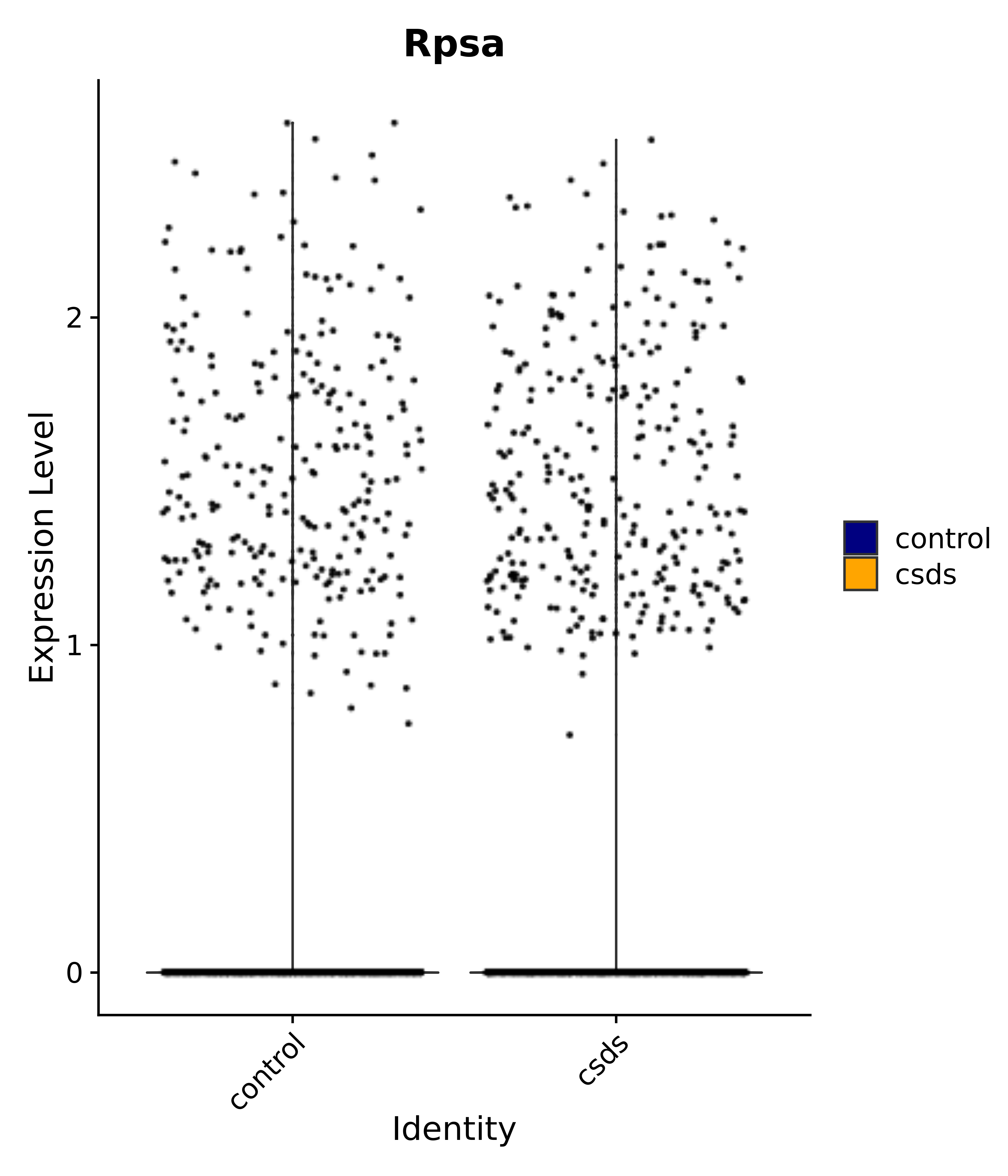
"Rpsa",
"Rtn1",
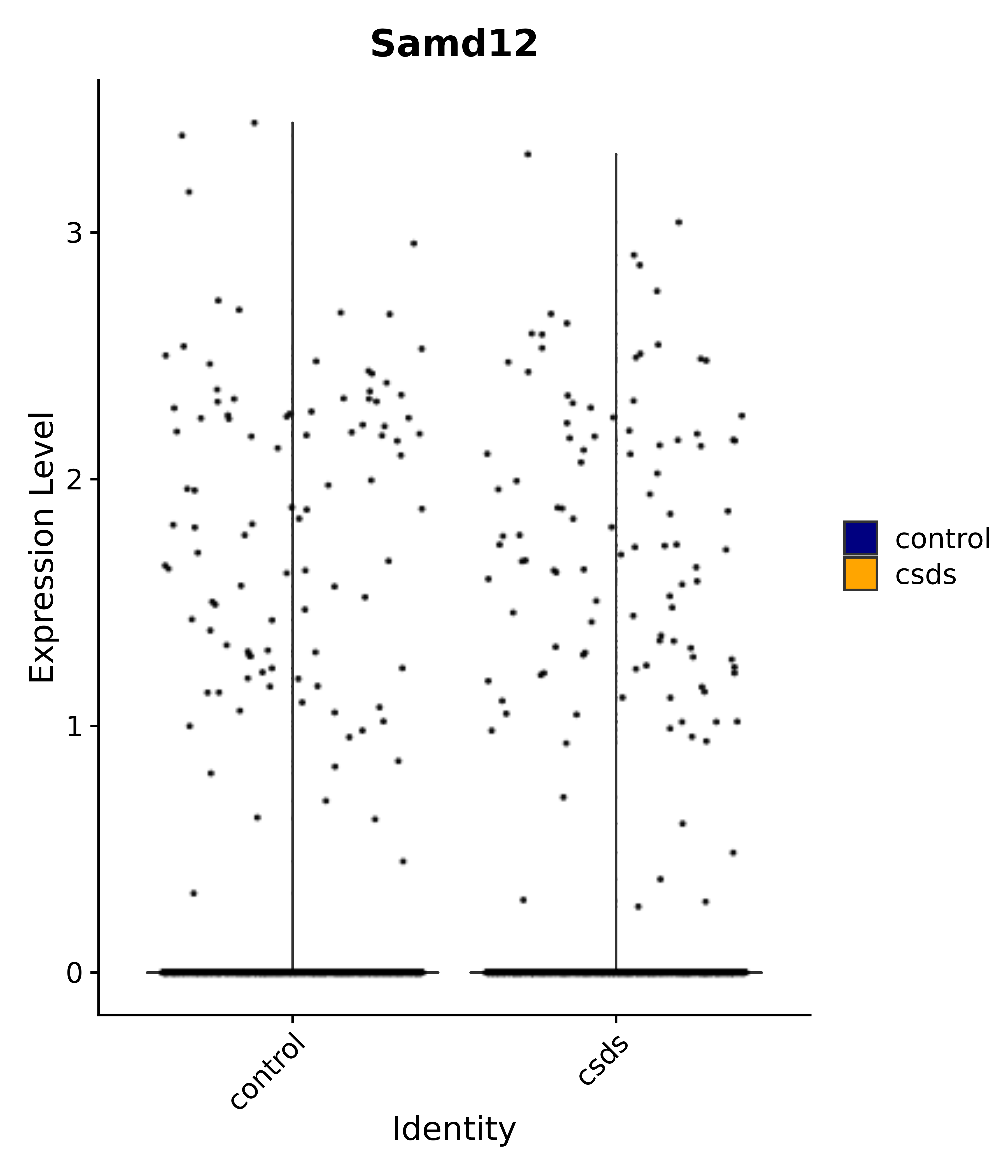
"Samd12",
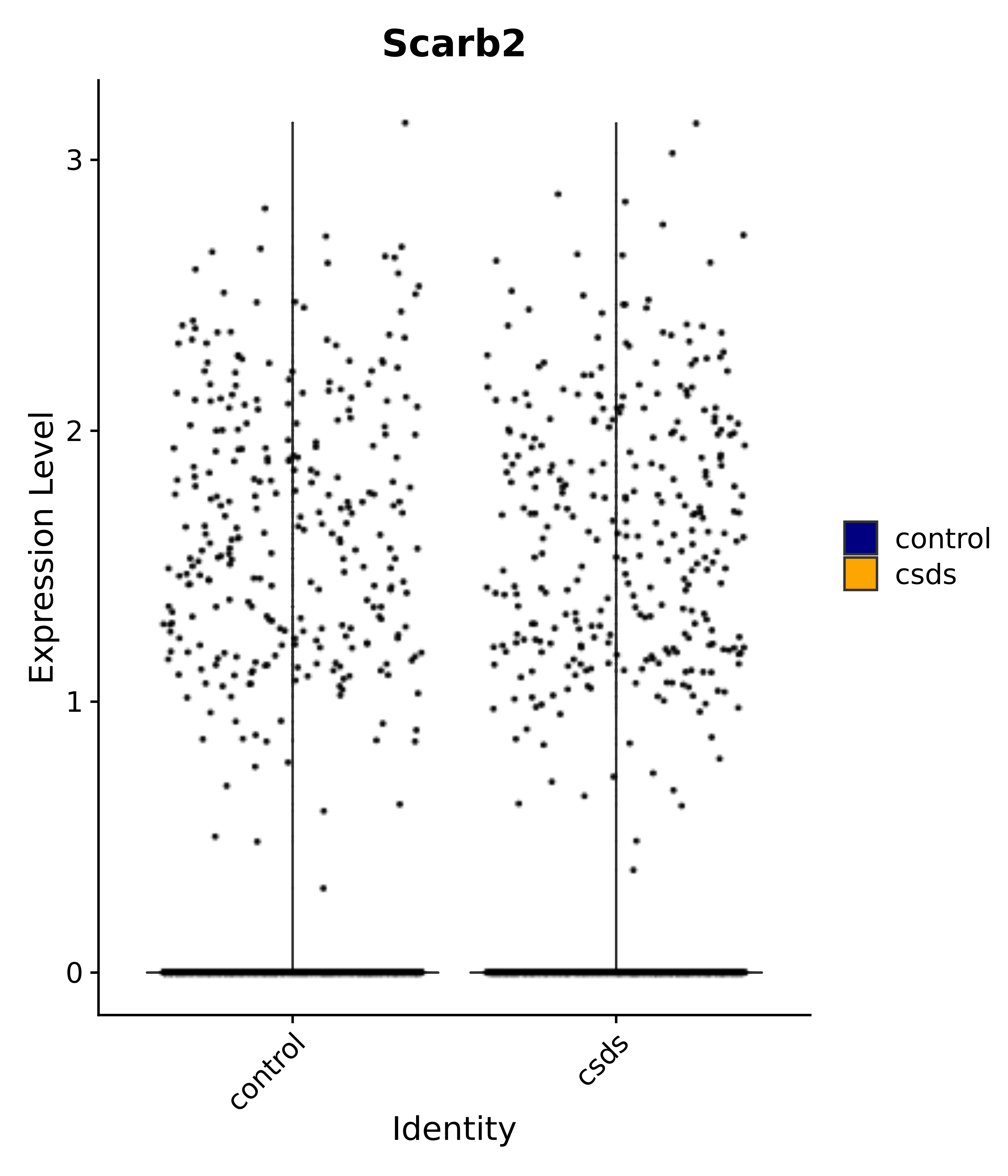
"Scarb2",
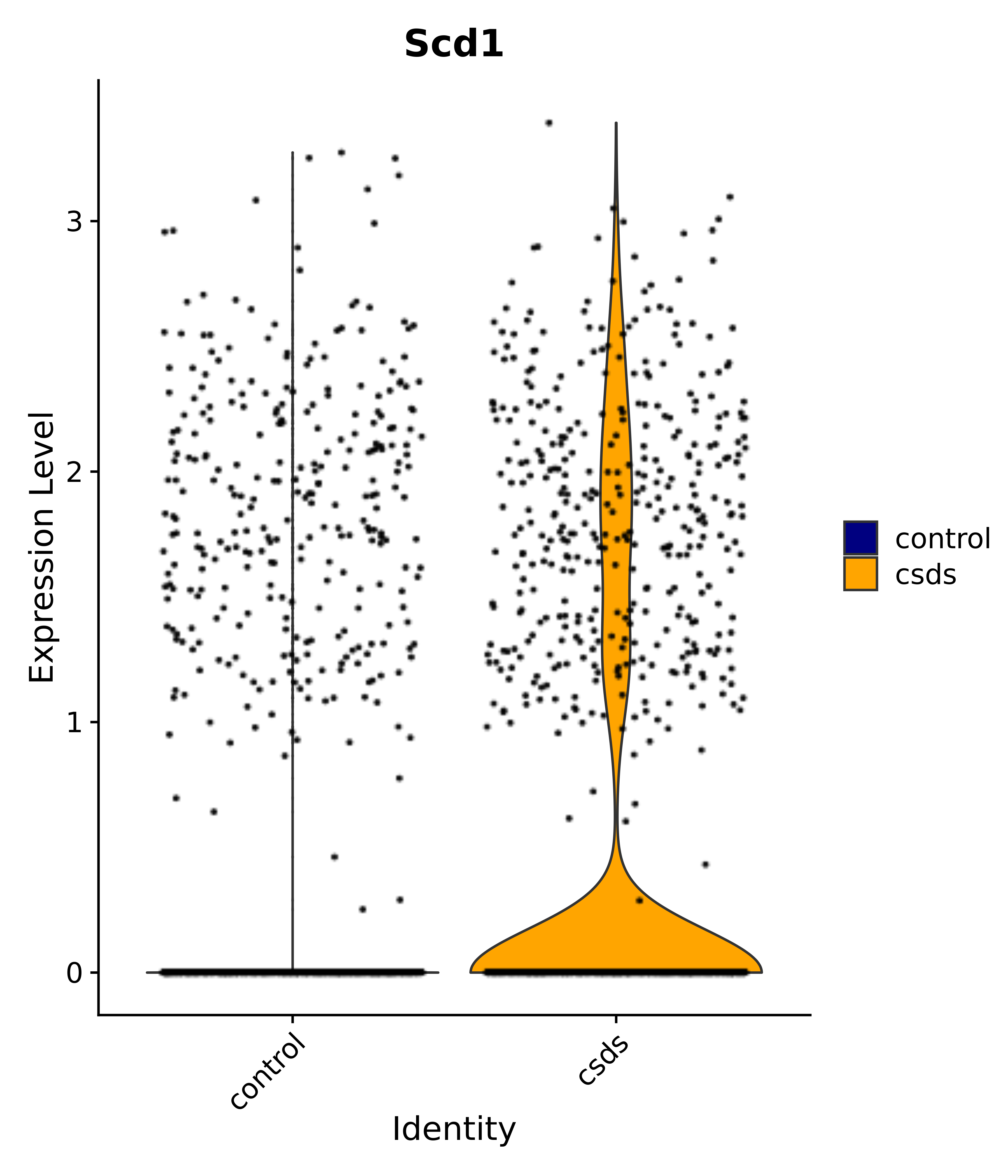
"Scd1",
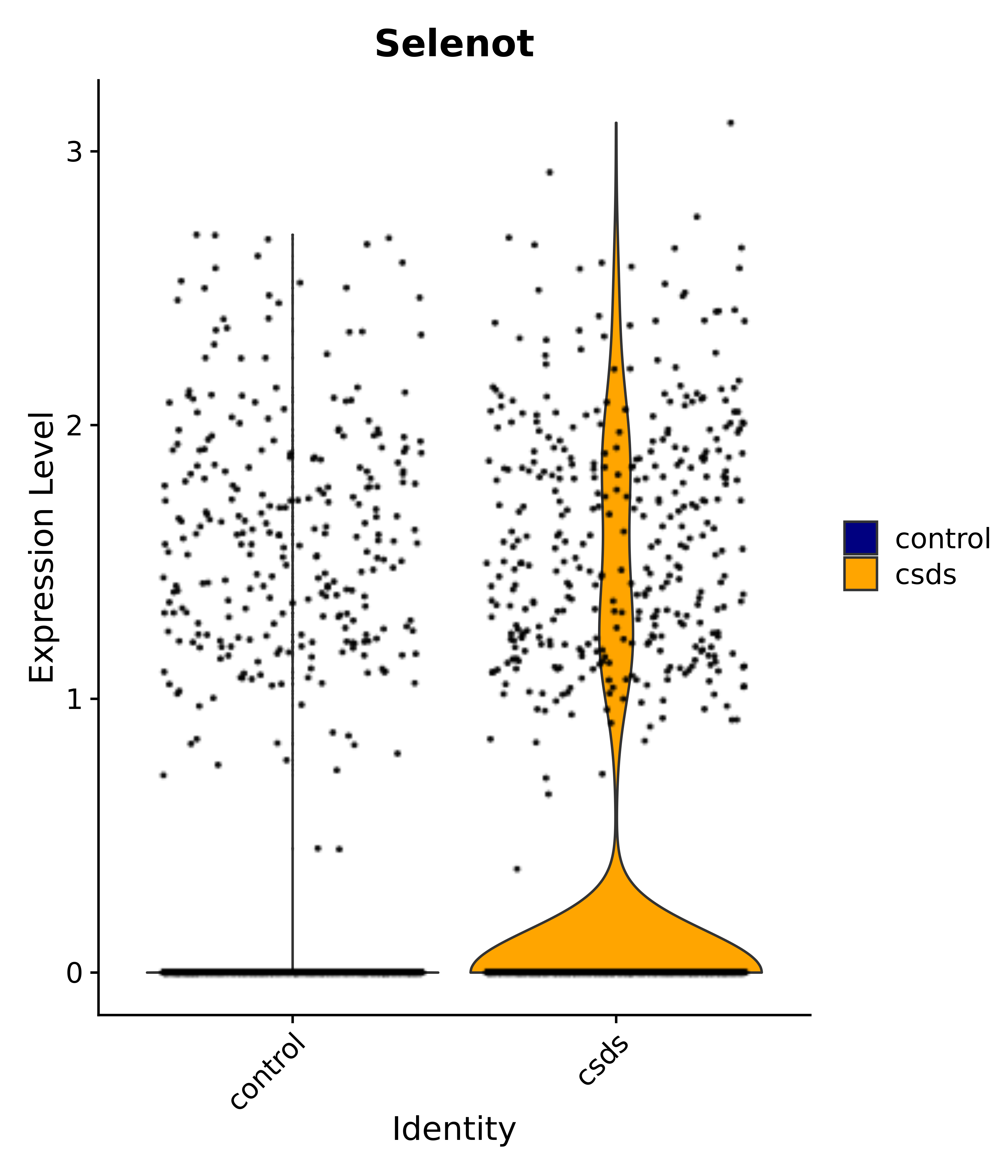
"Selenot",
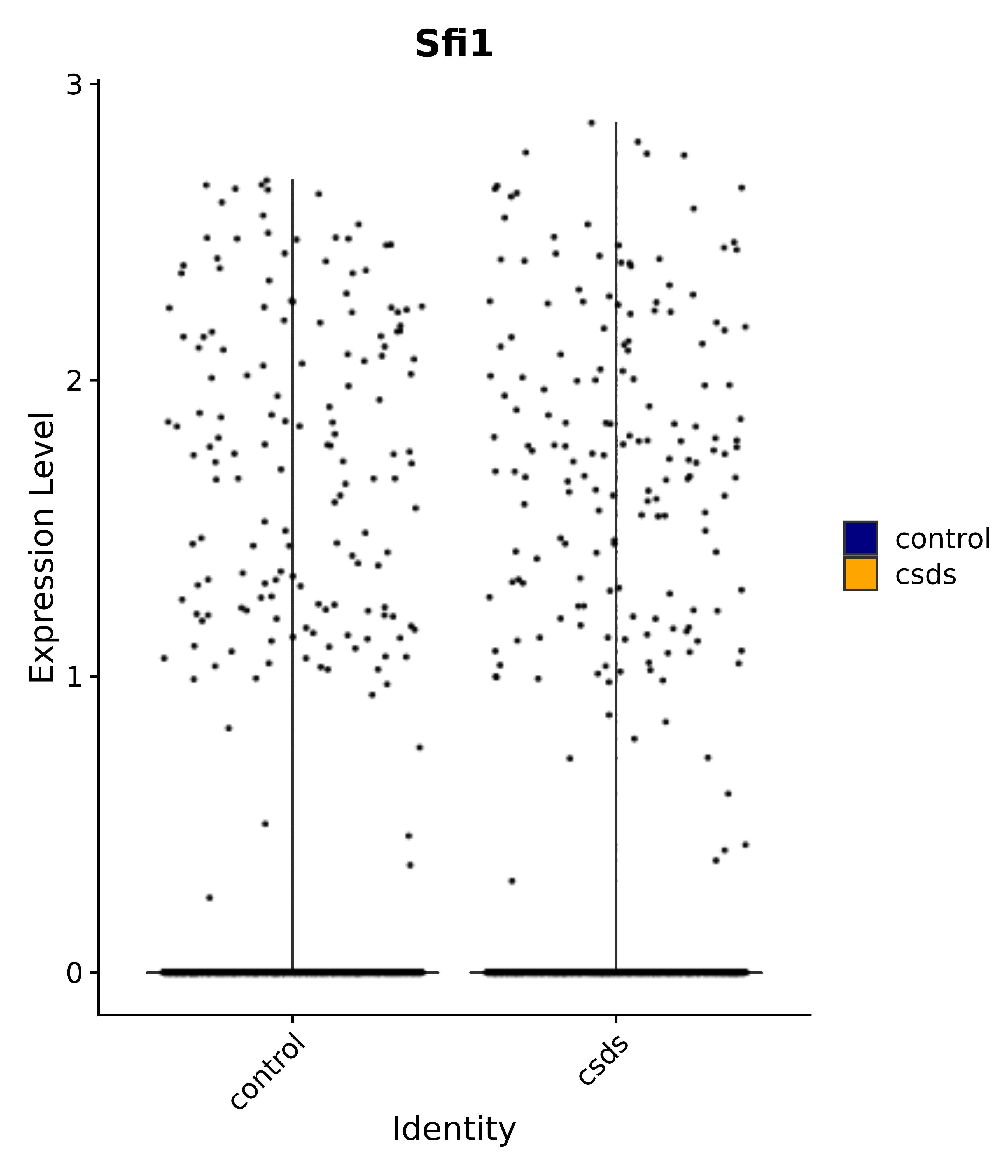
"Sfi1",
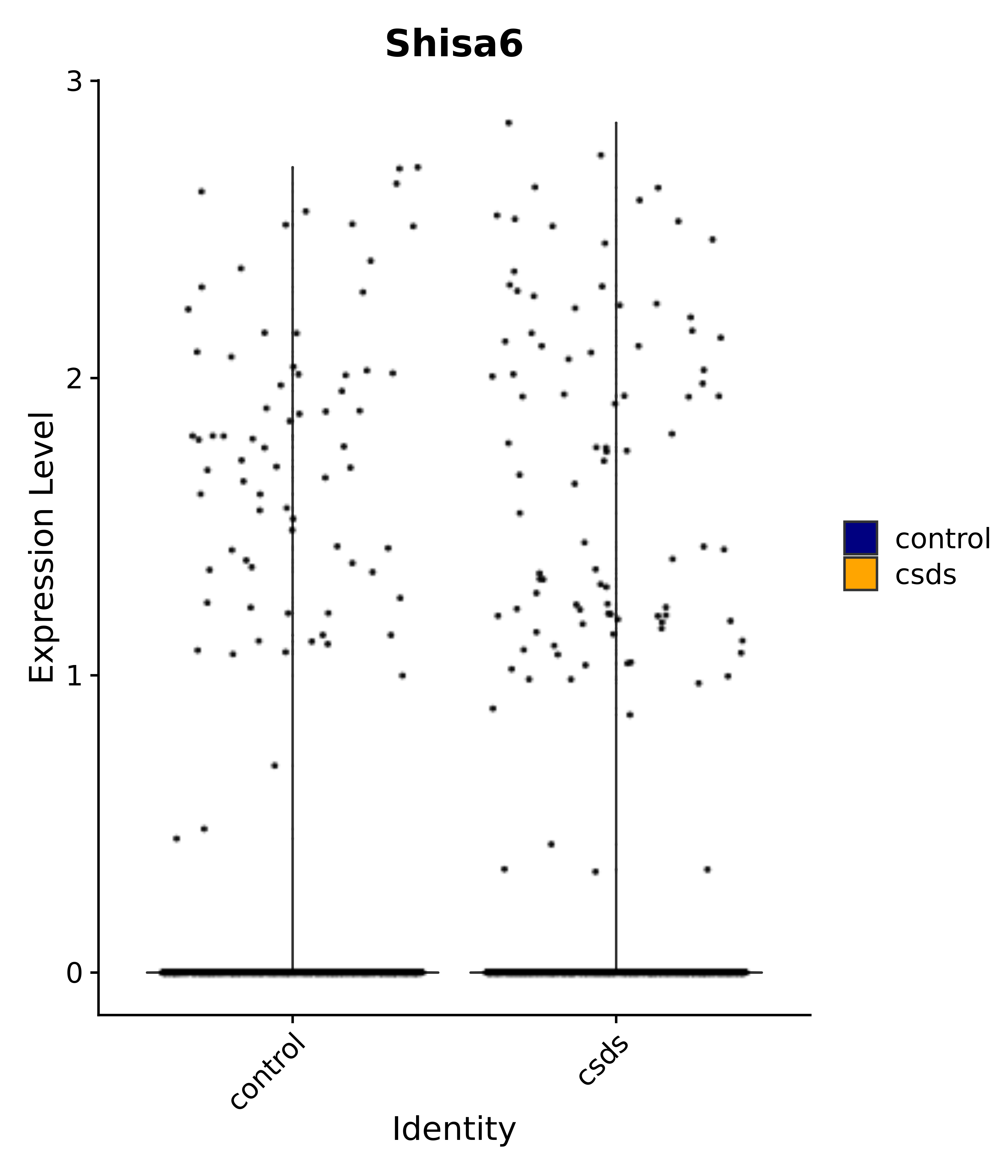
"Shisa6",
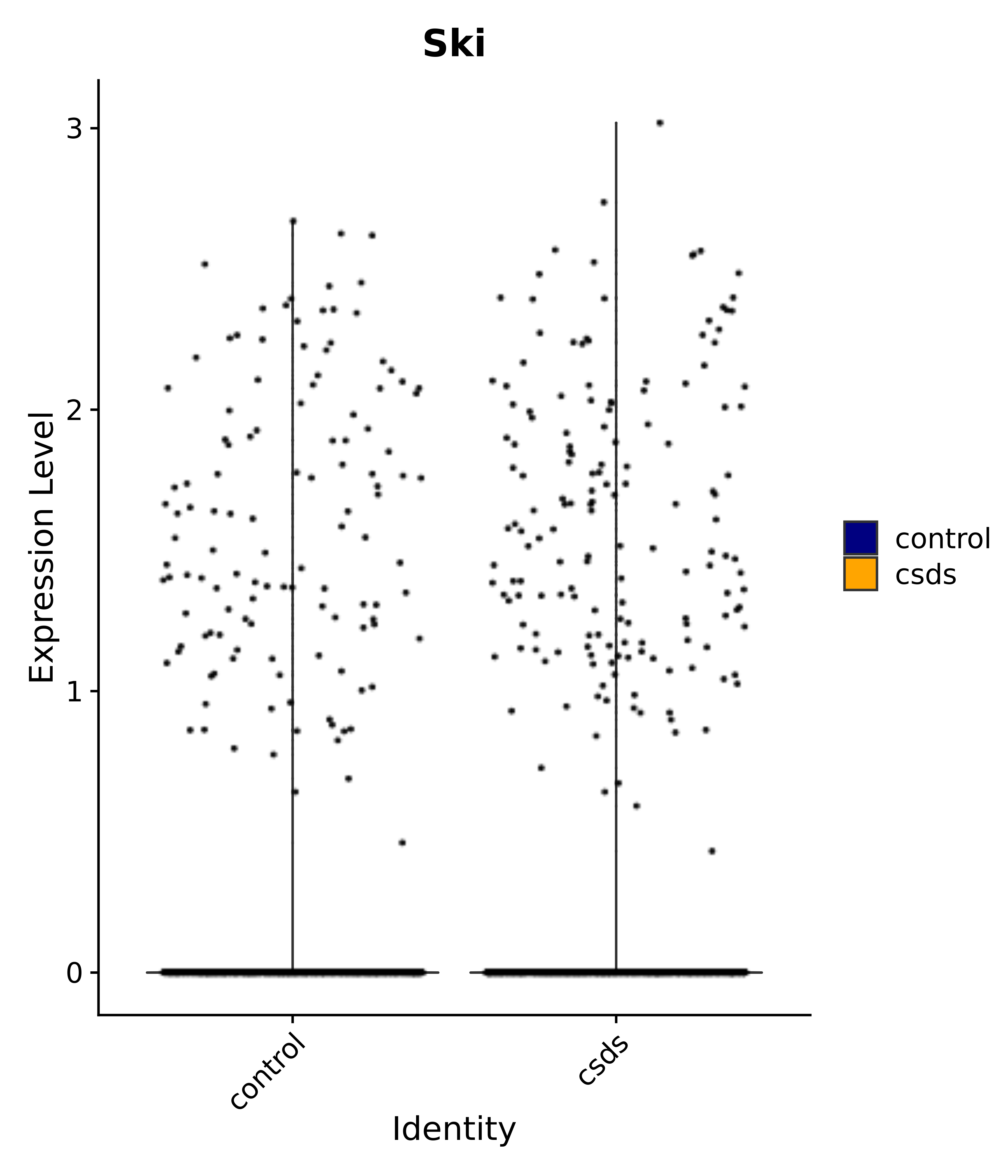
"Ski",
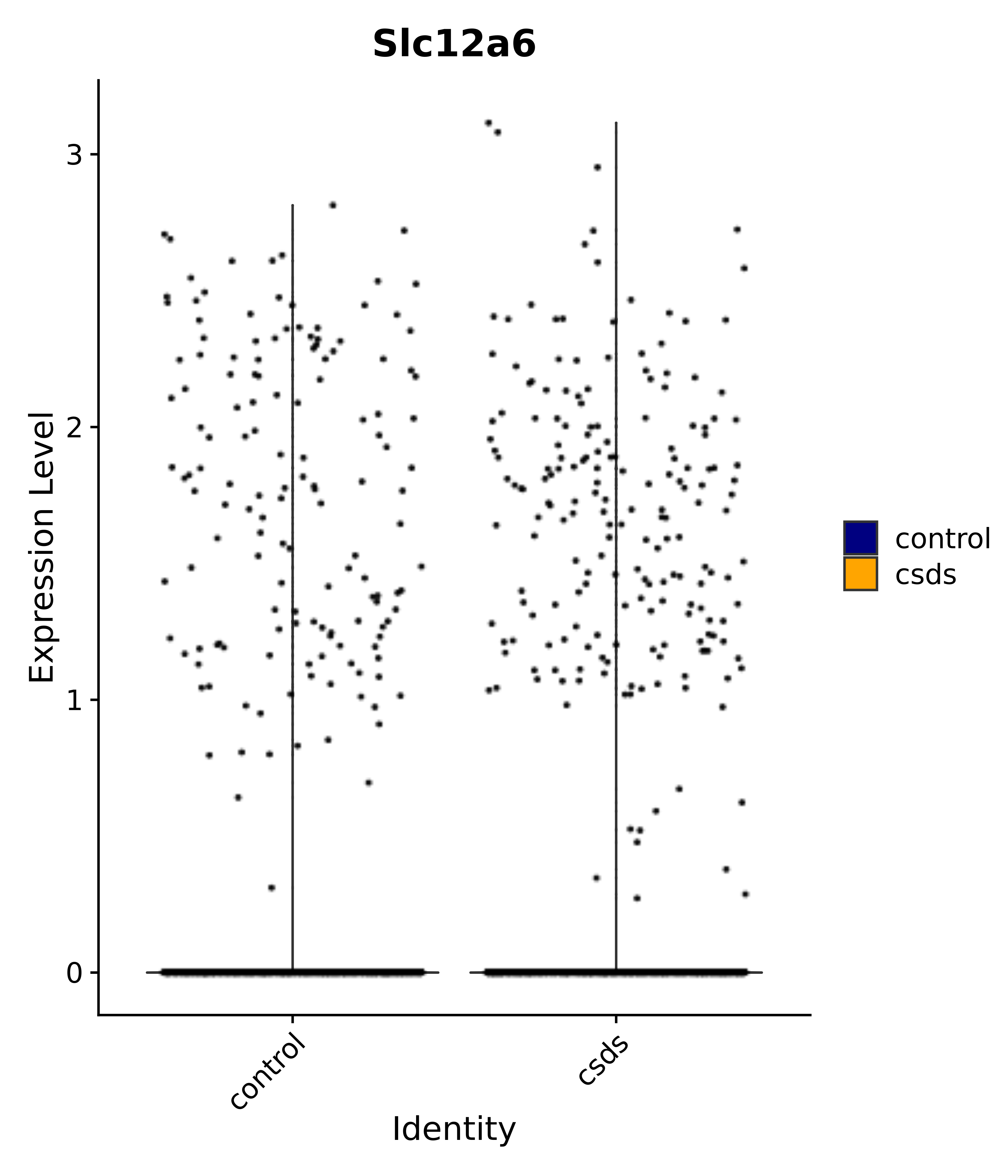
"Slc12a6",
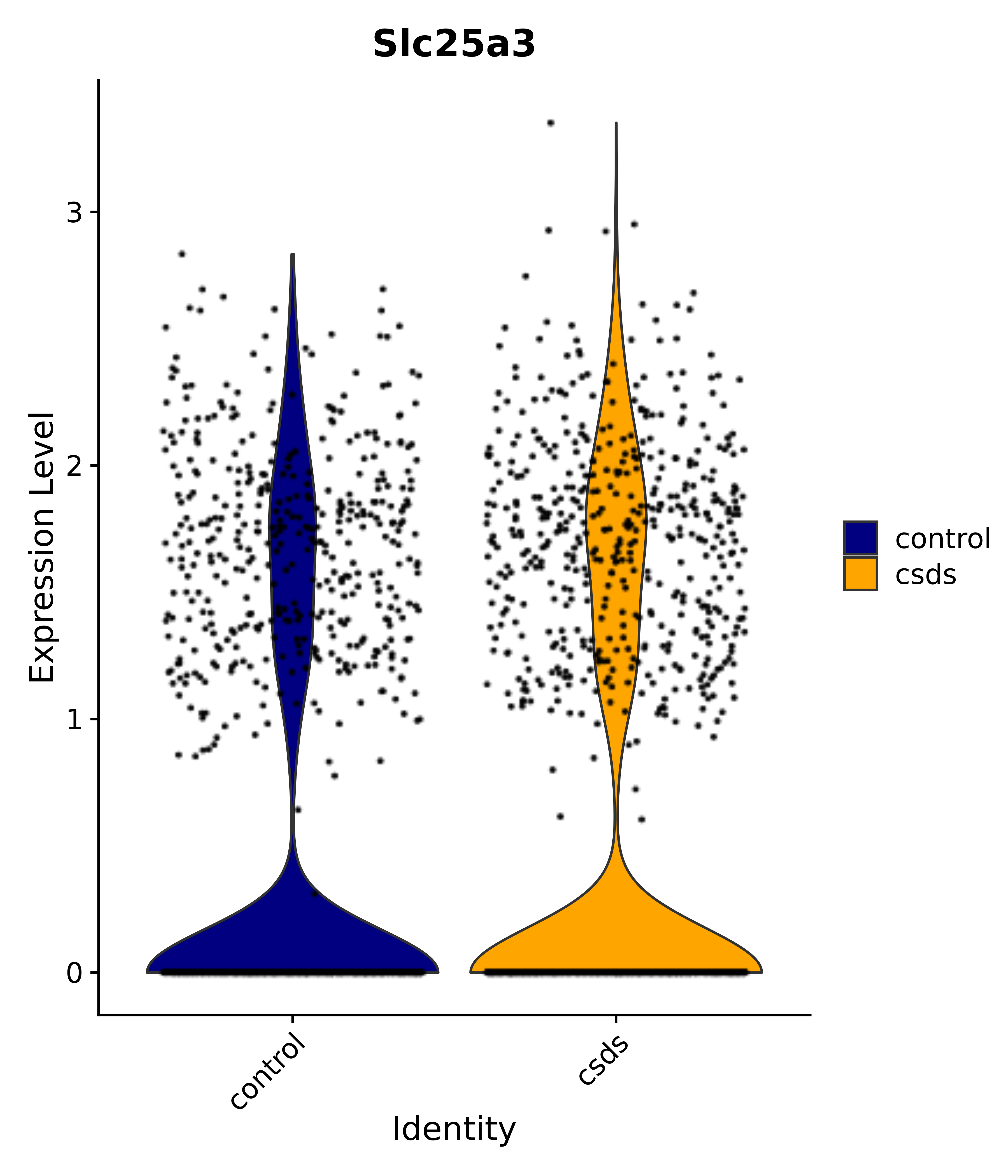
"Slc25a3",
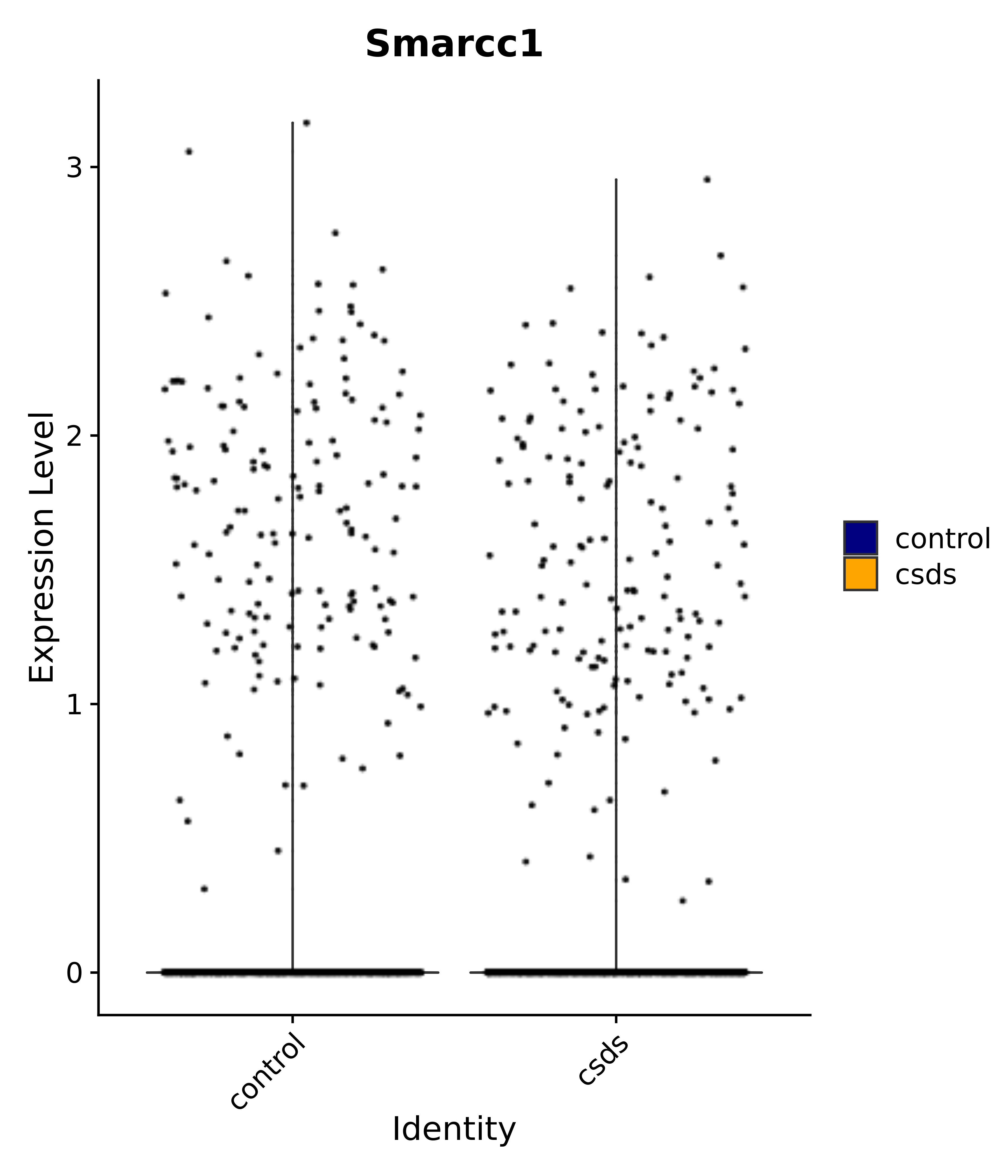
"Smarcc1",
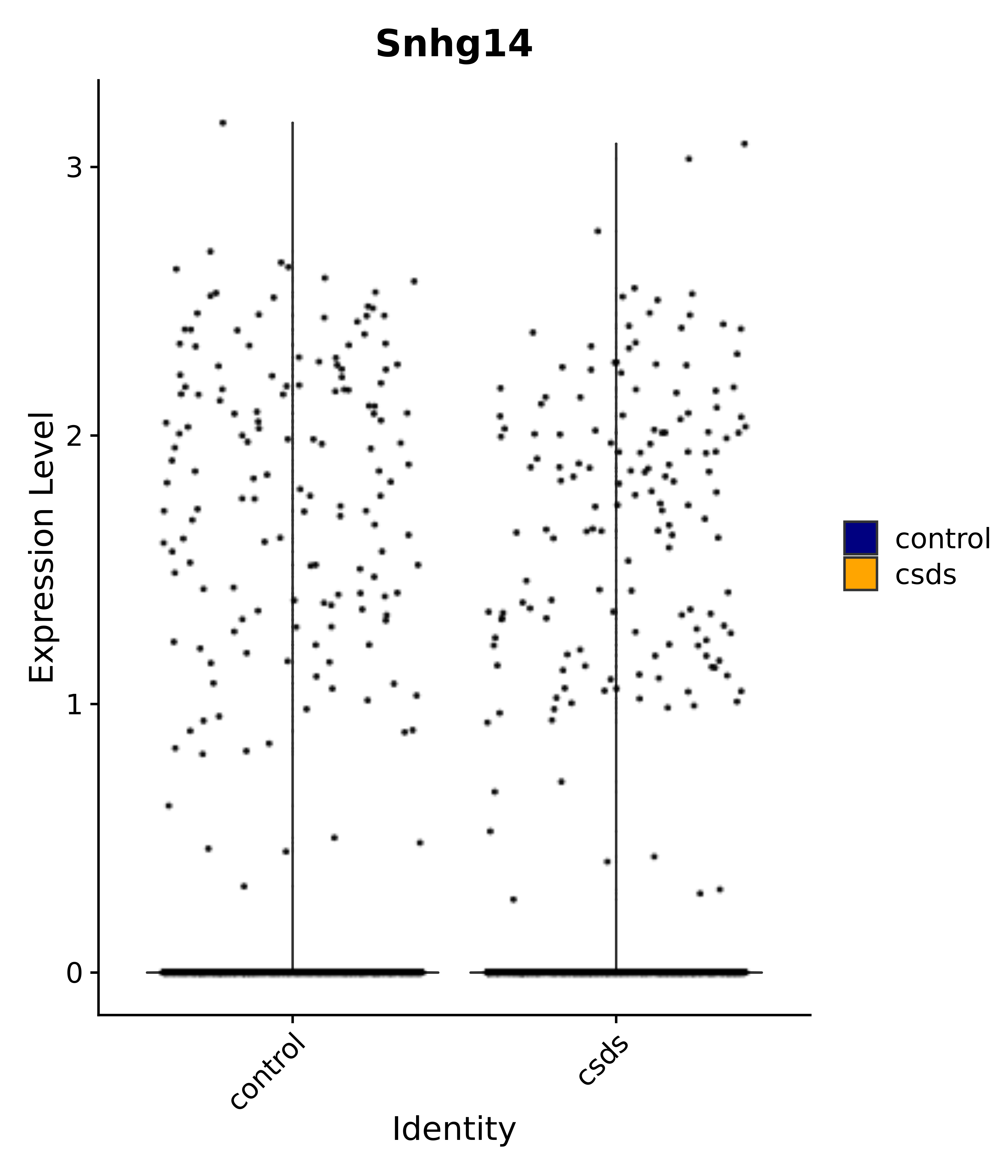
"Snhg14",
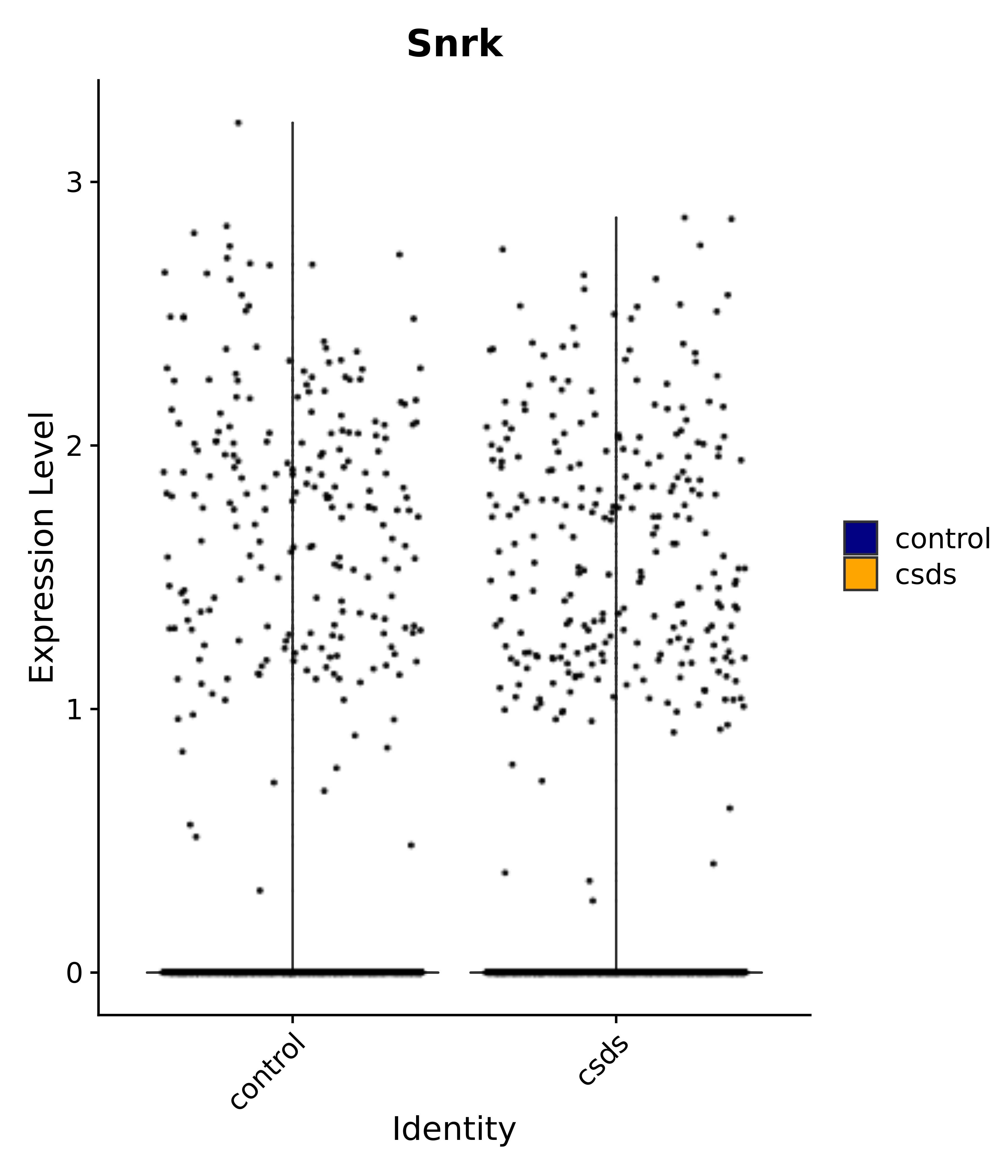
"Snrk",
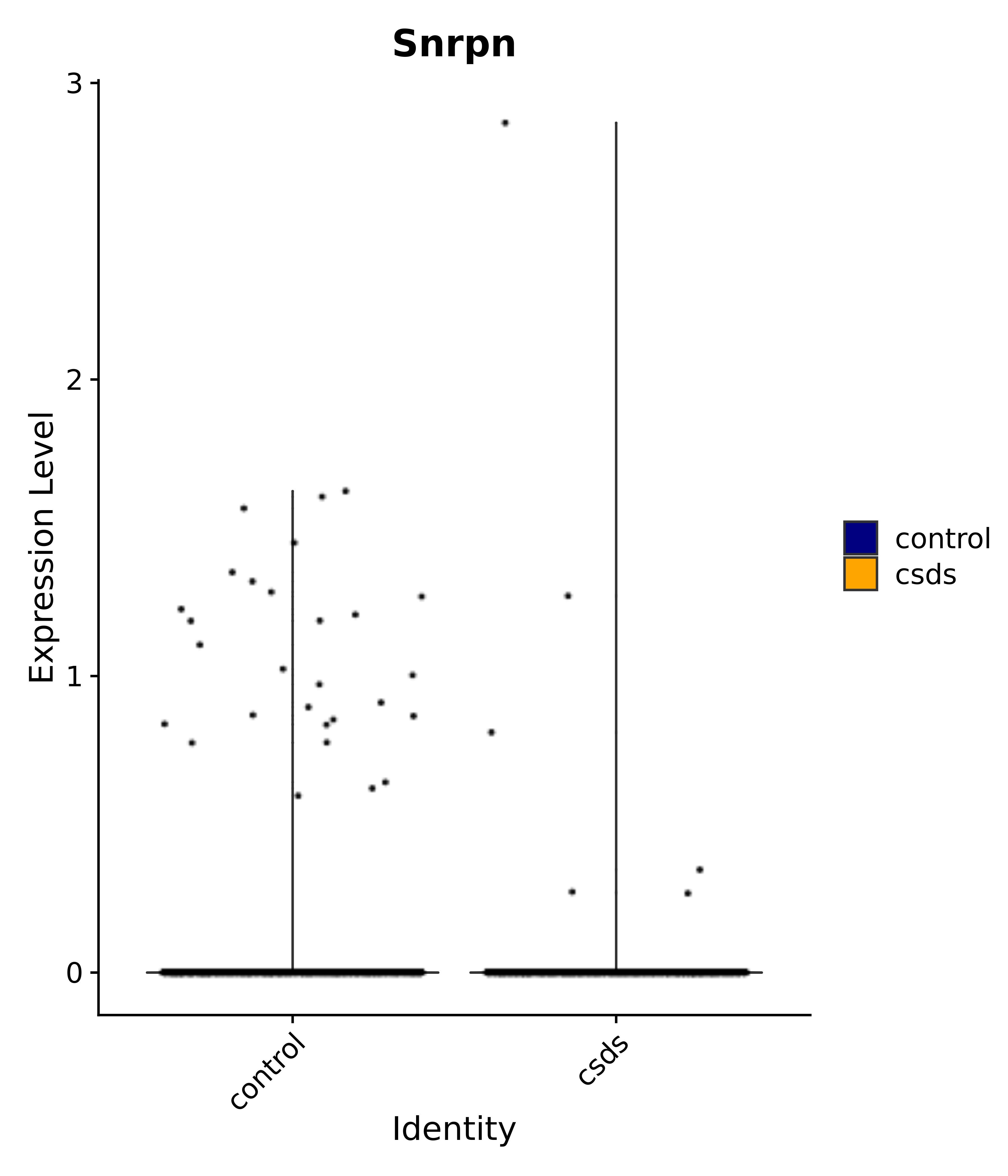
"Snrpn",
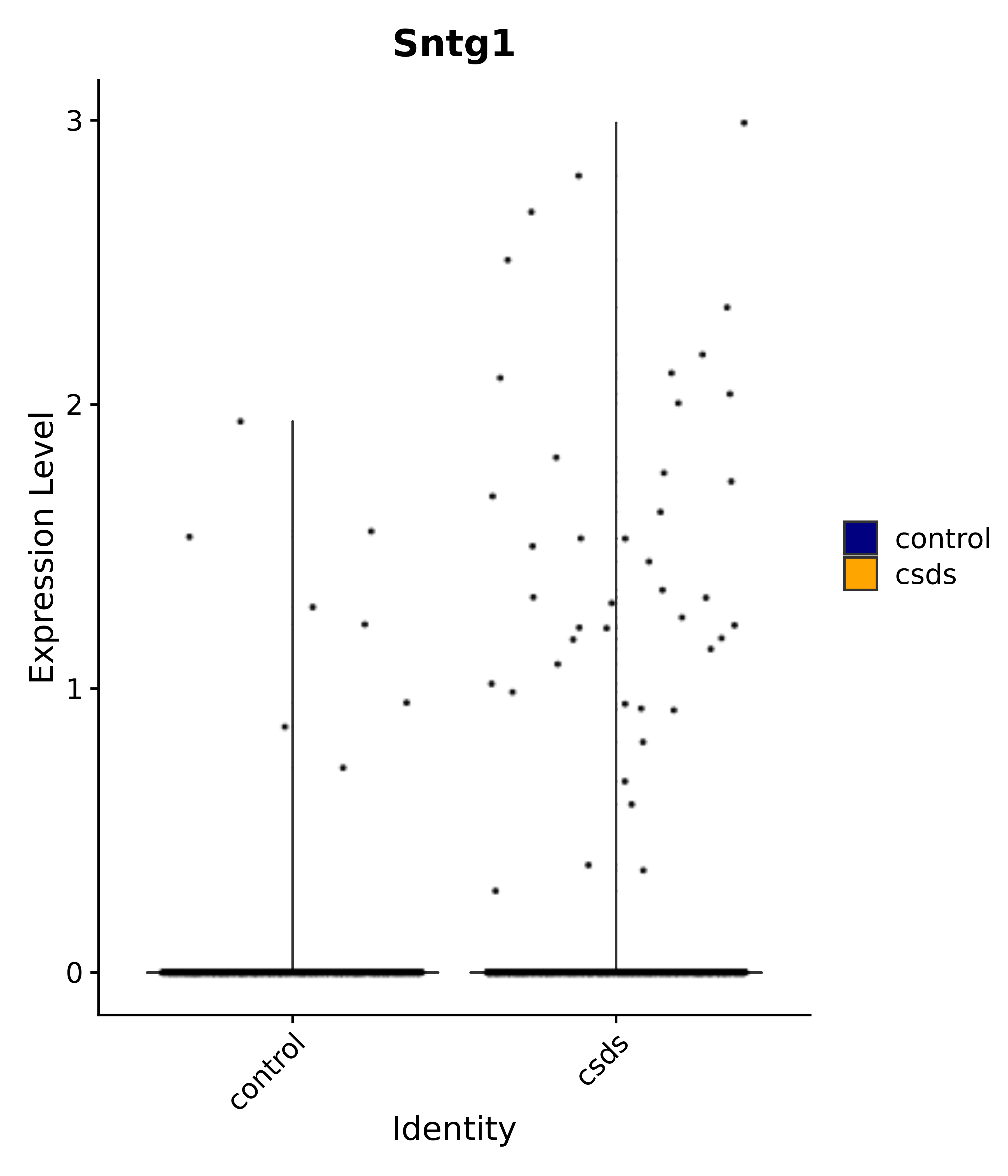
"Sntg1",
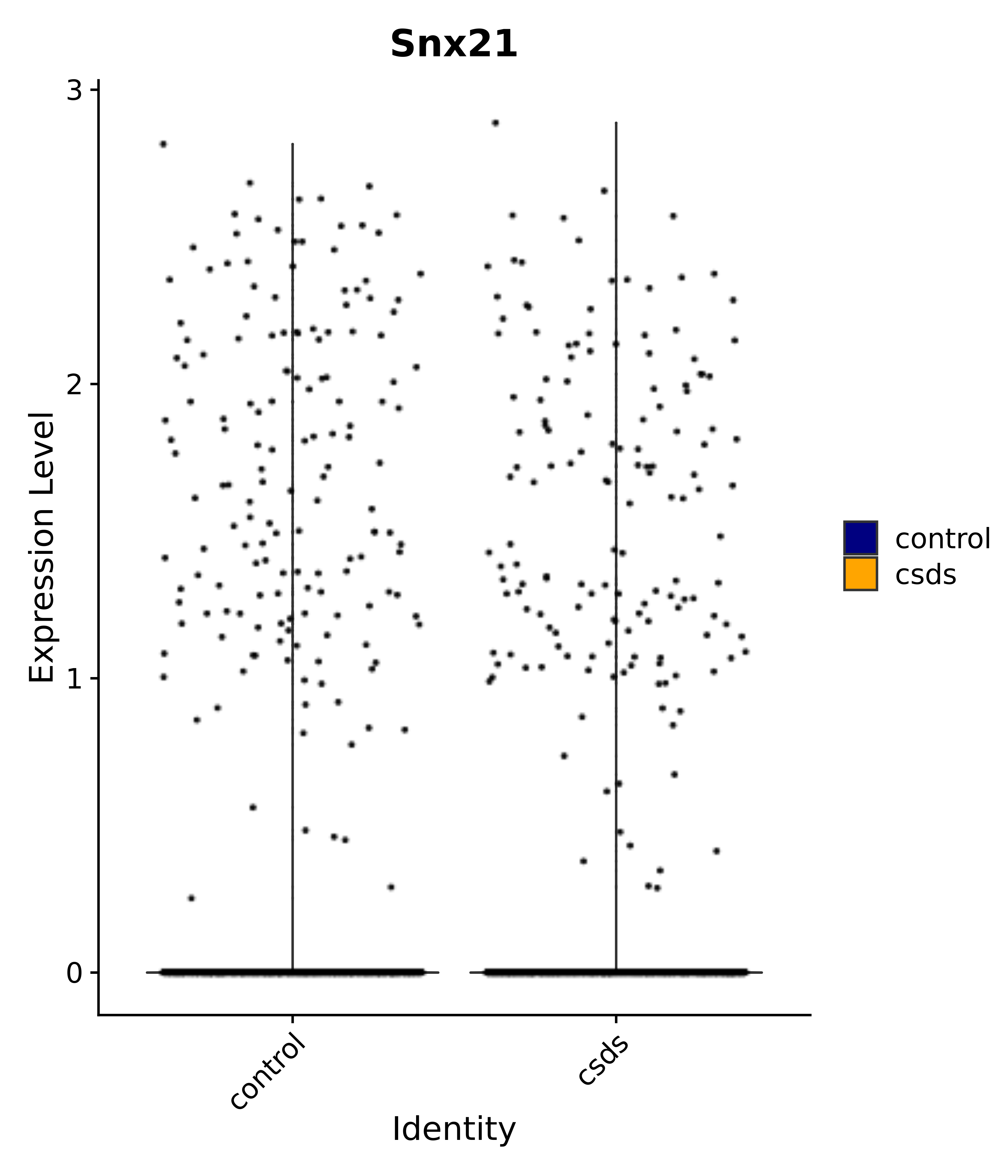
"Snx21",
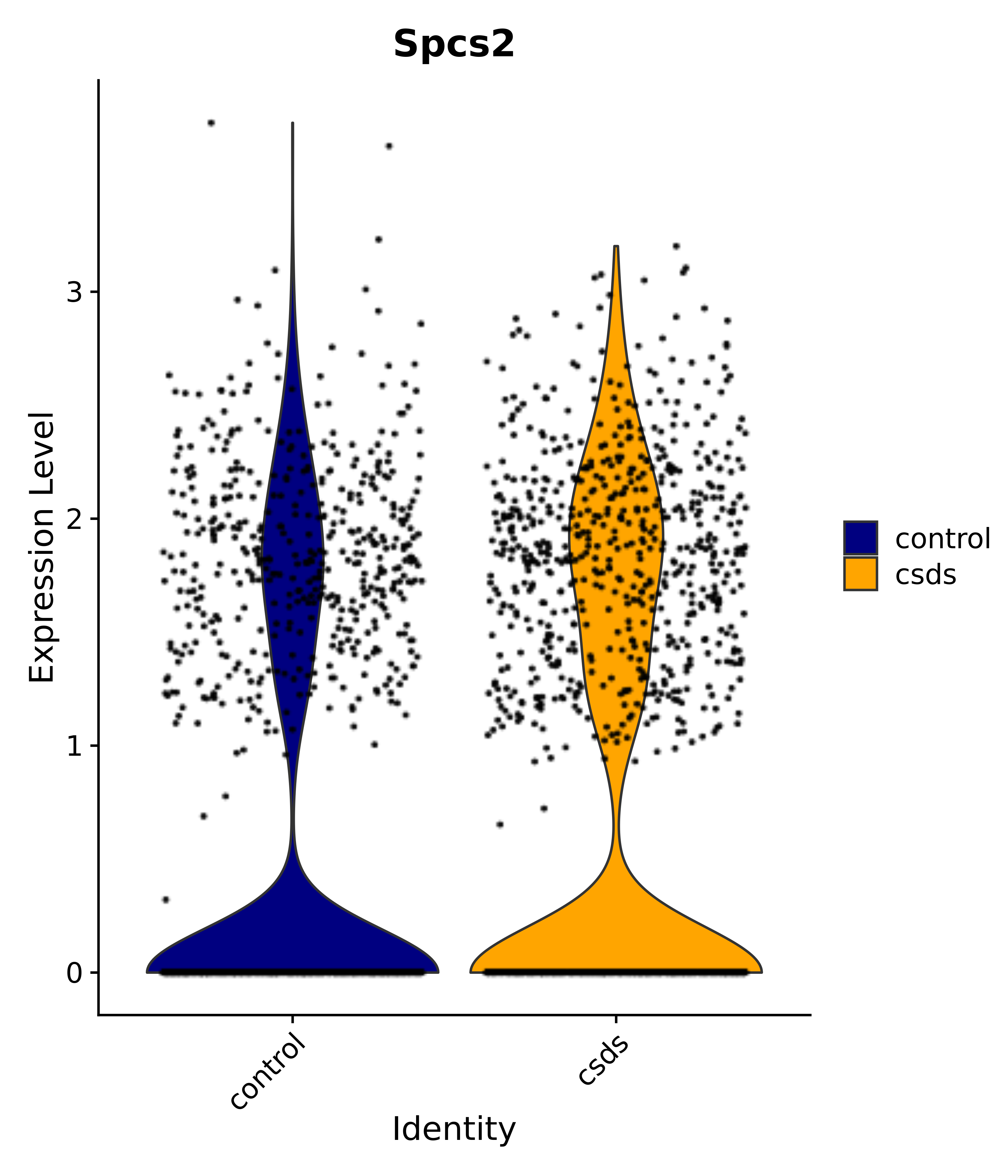
"Spcs2",
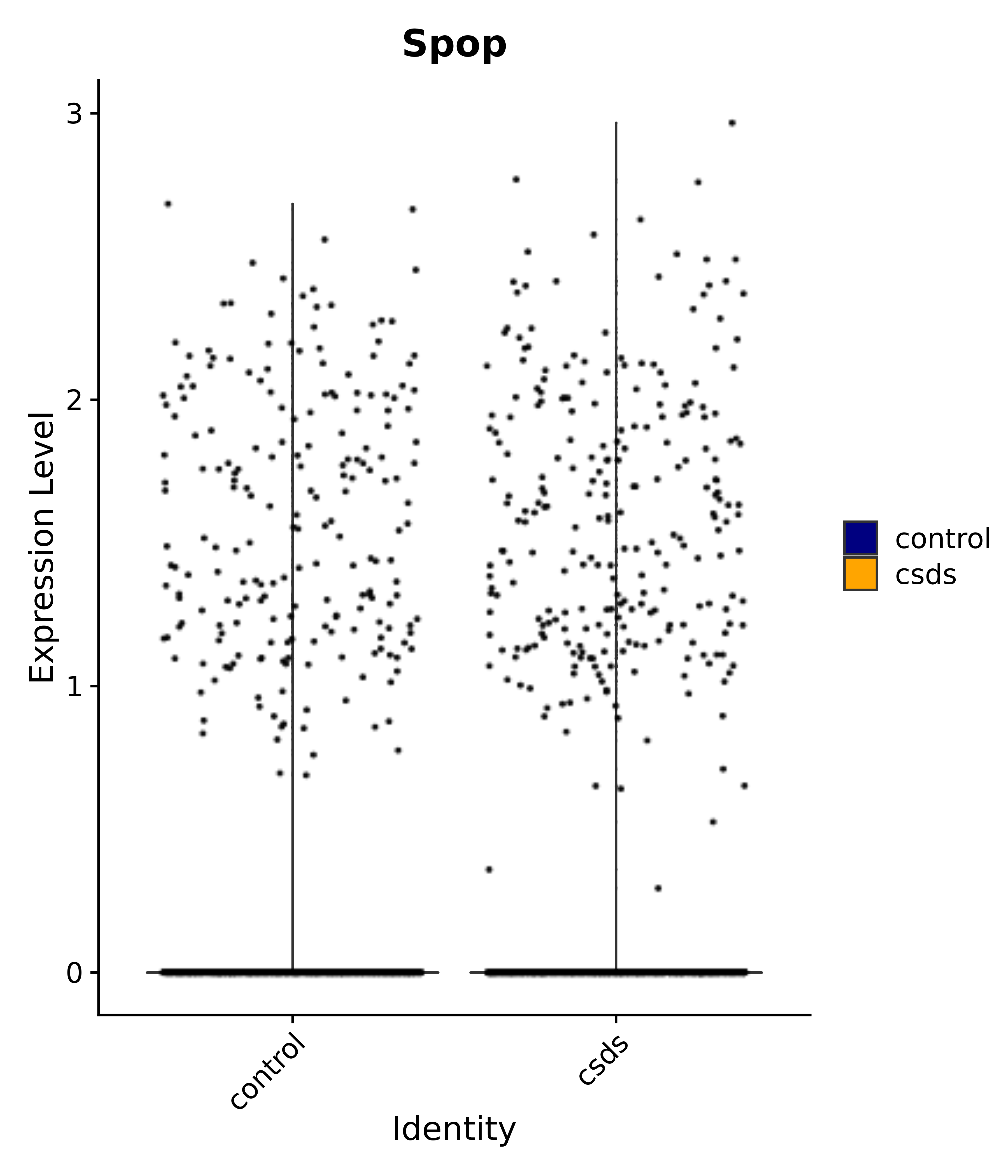
"Spop",
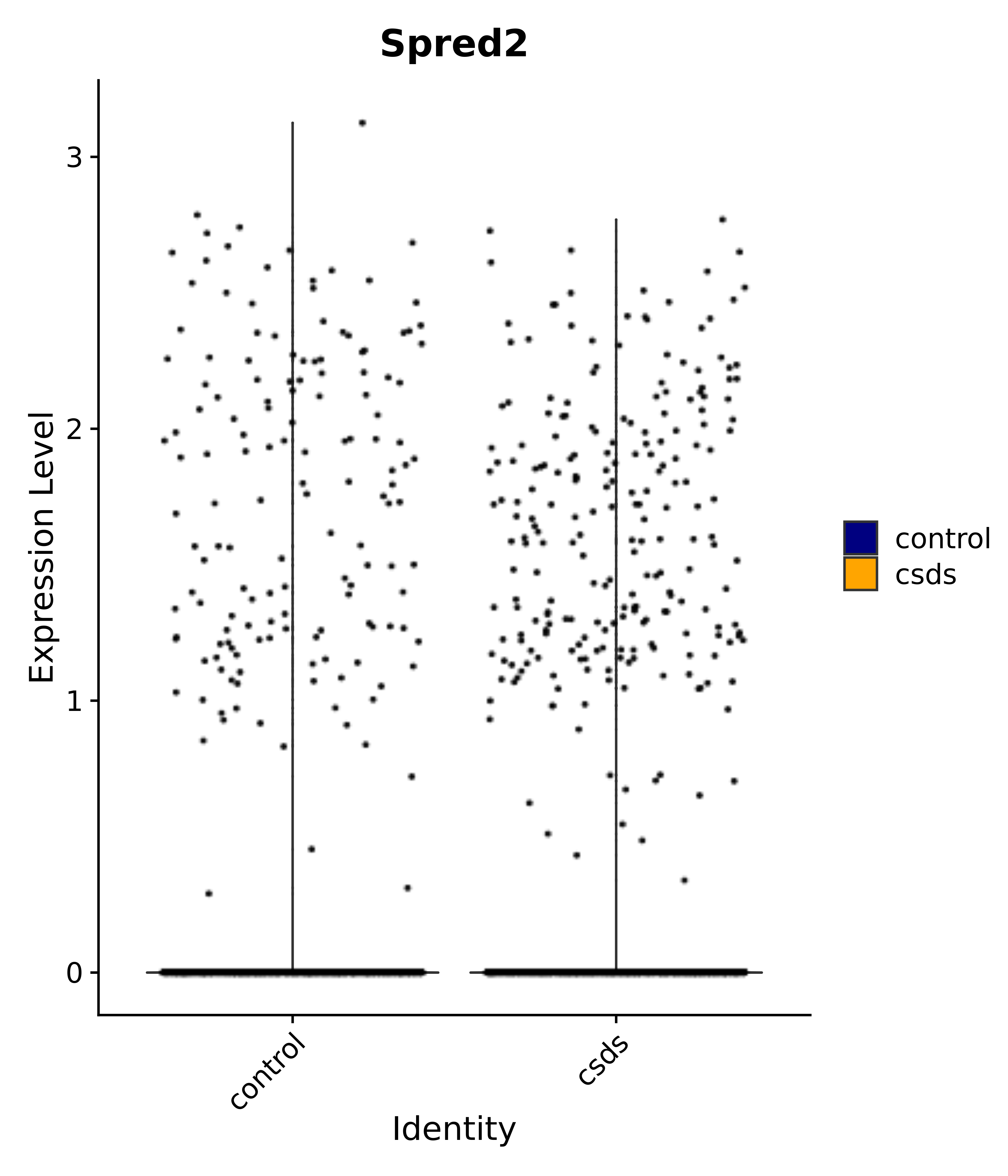
"Spred2",
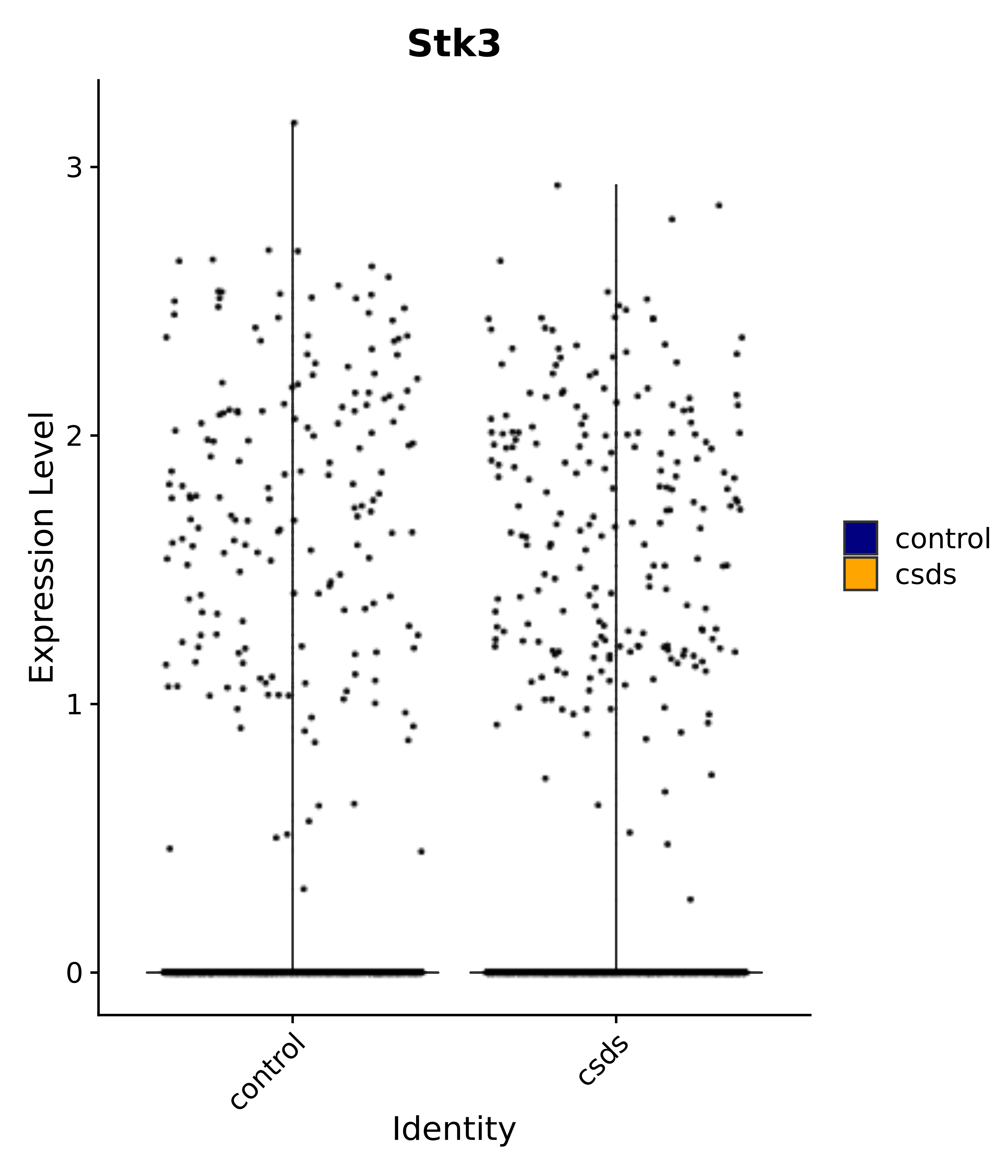
"Stk3",
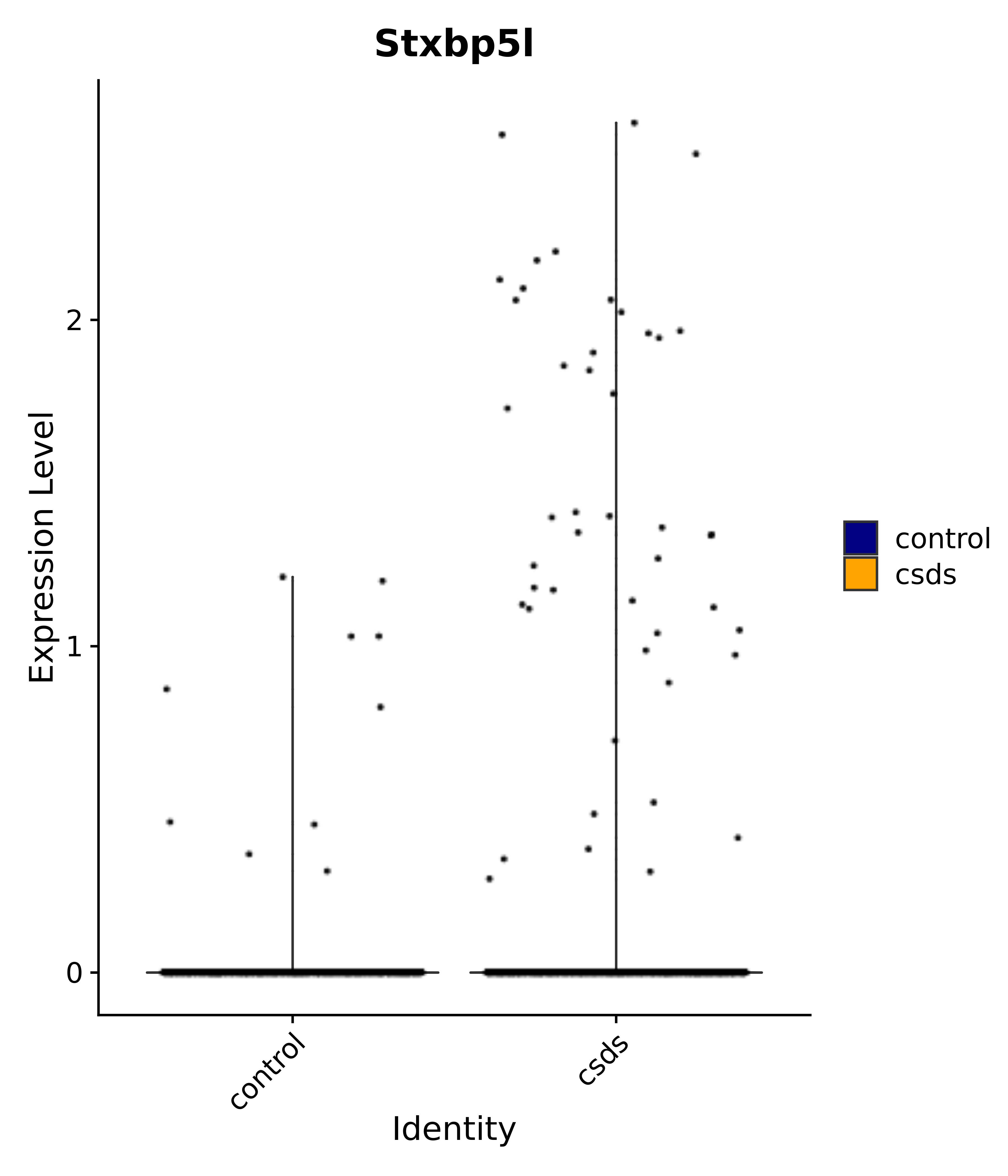
"Stxbp5l",
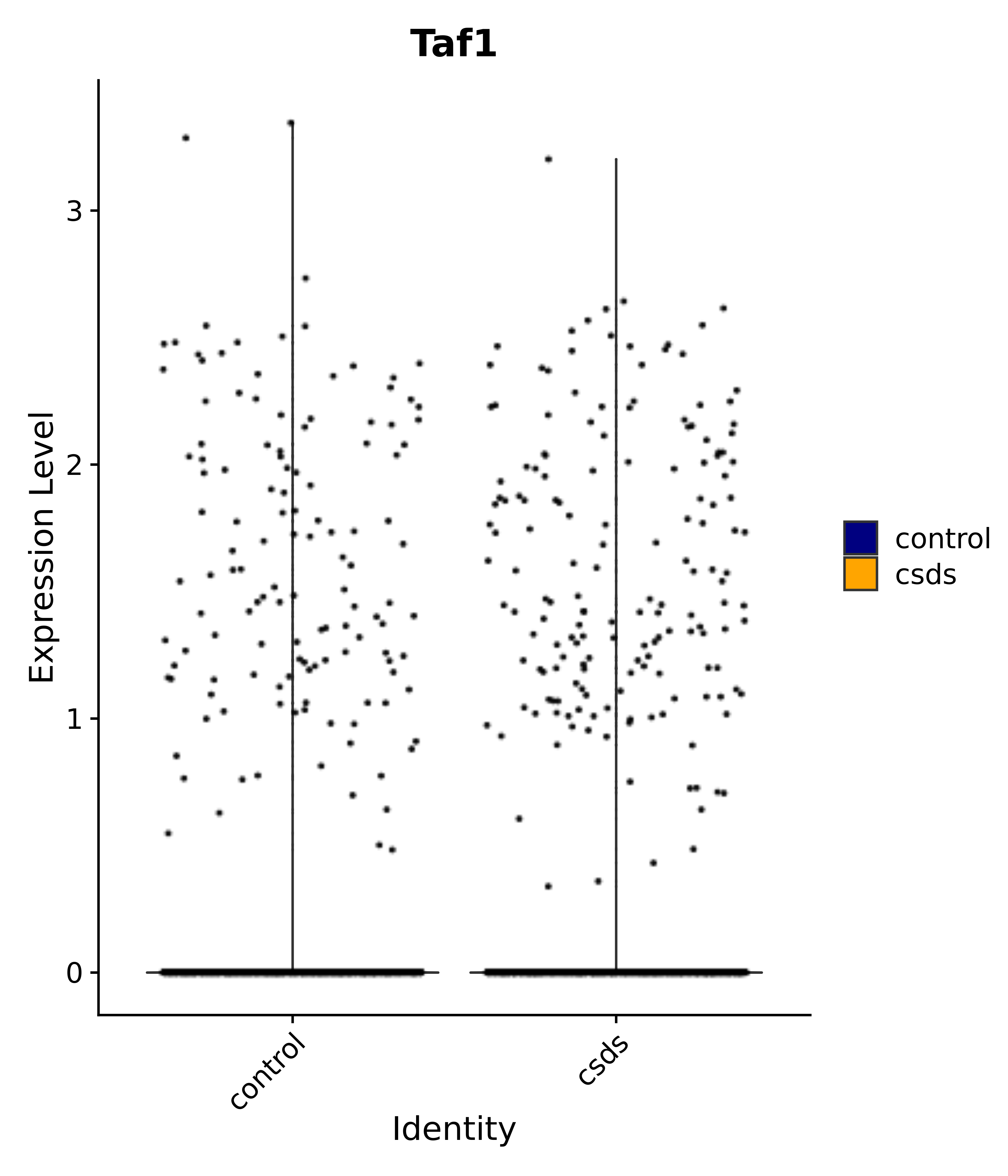
"Taf1",
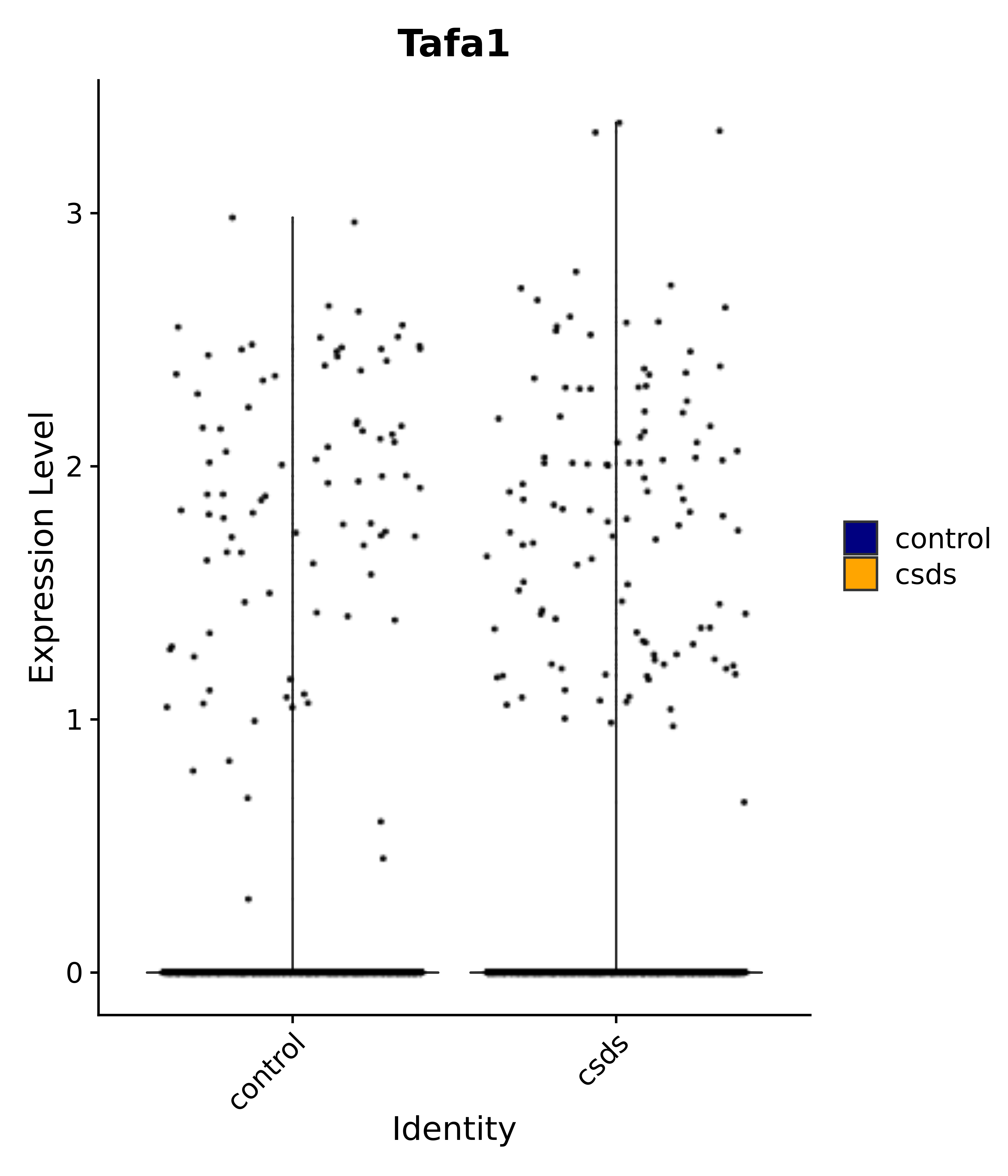
"Tafa1",
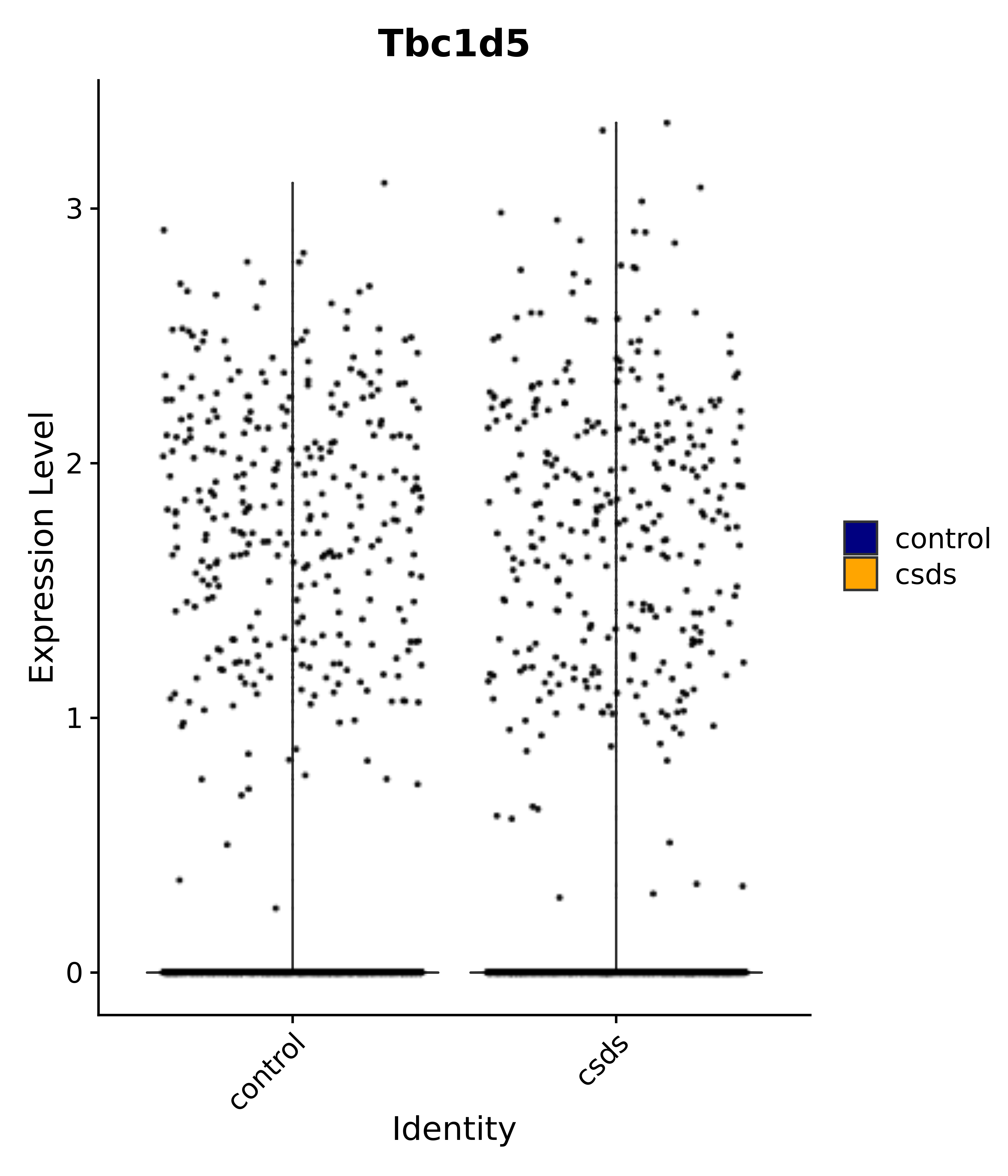
"Tbc1d5",
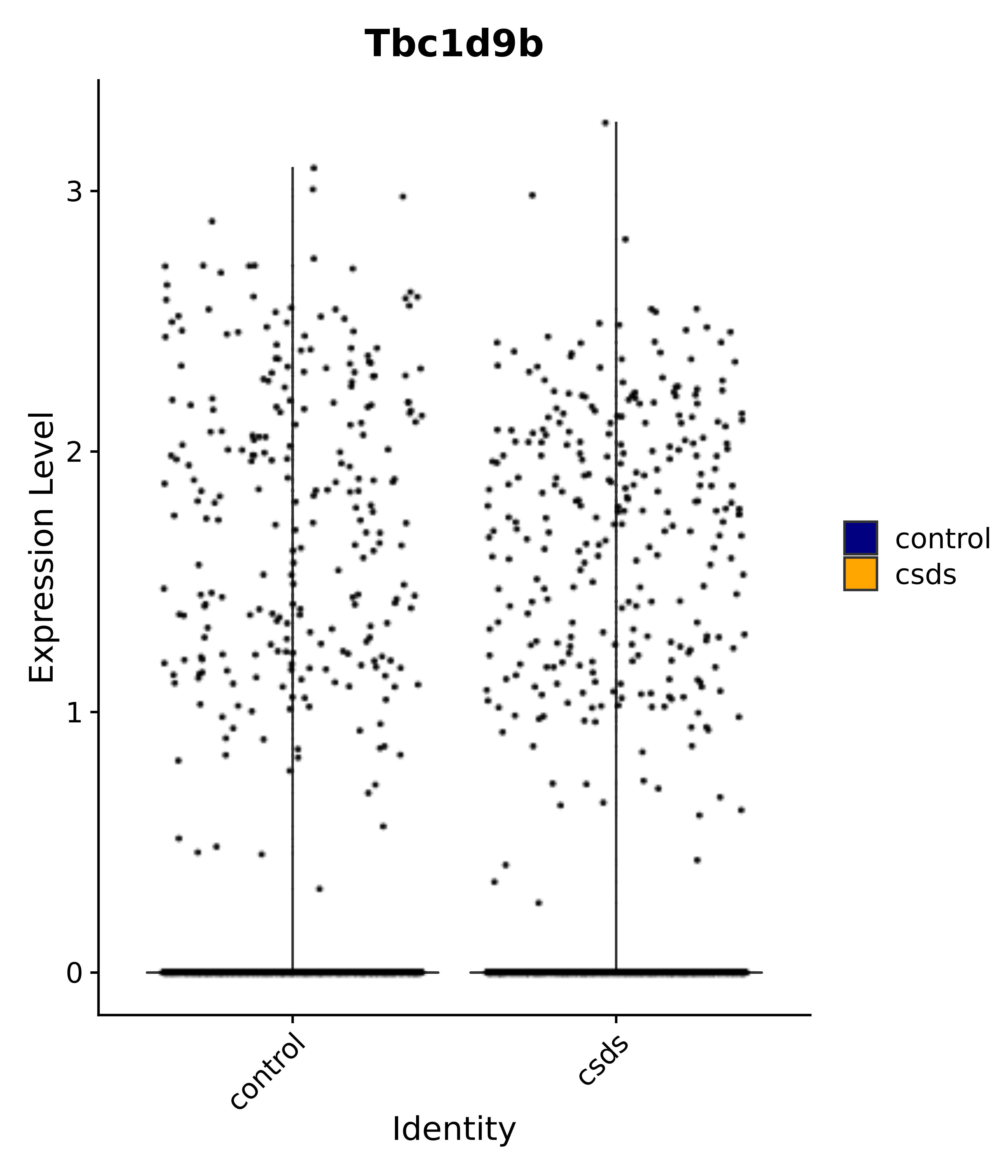
"Tbc1d9b",
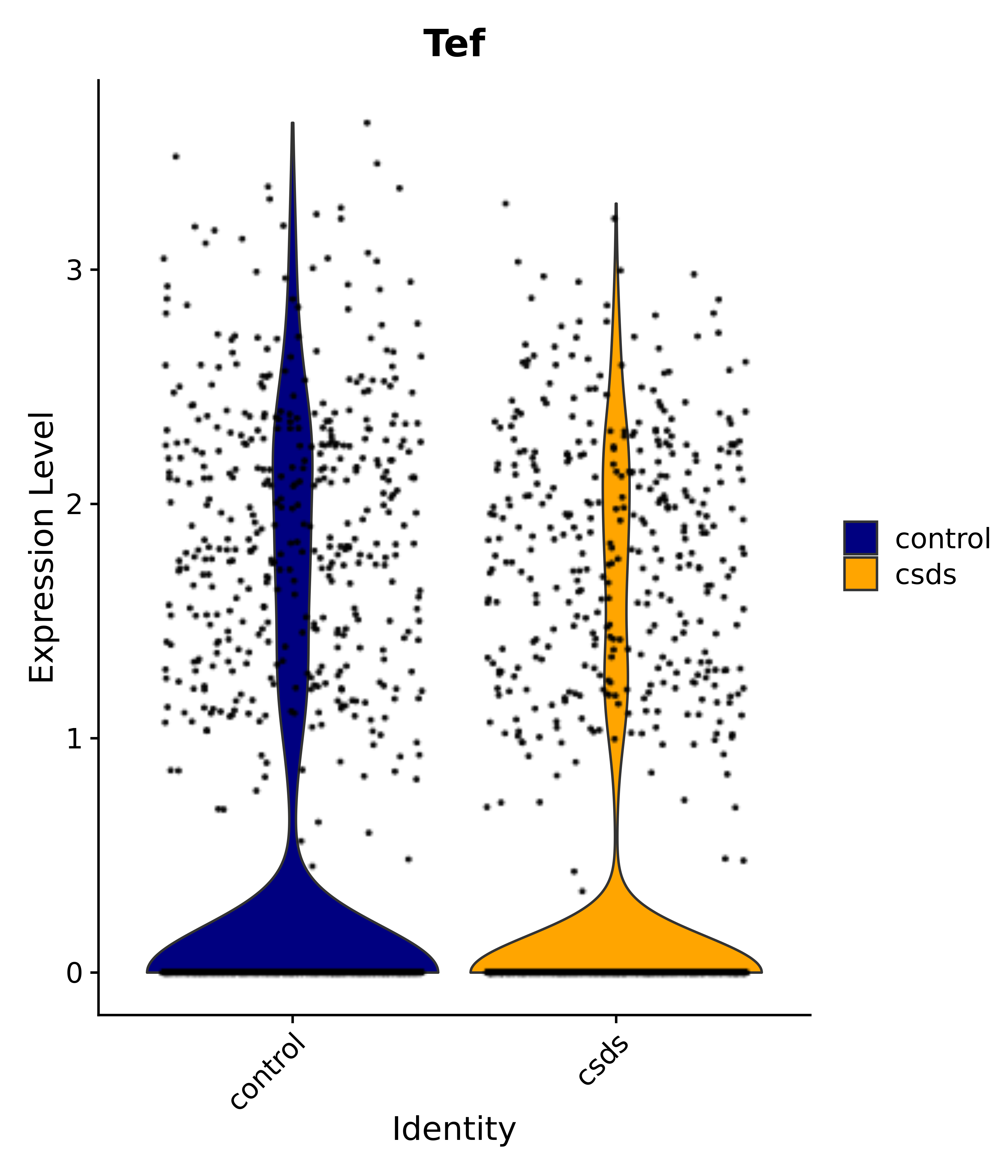
"Tef",
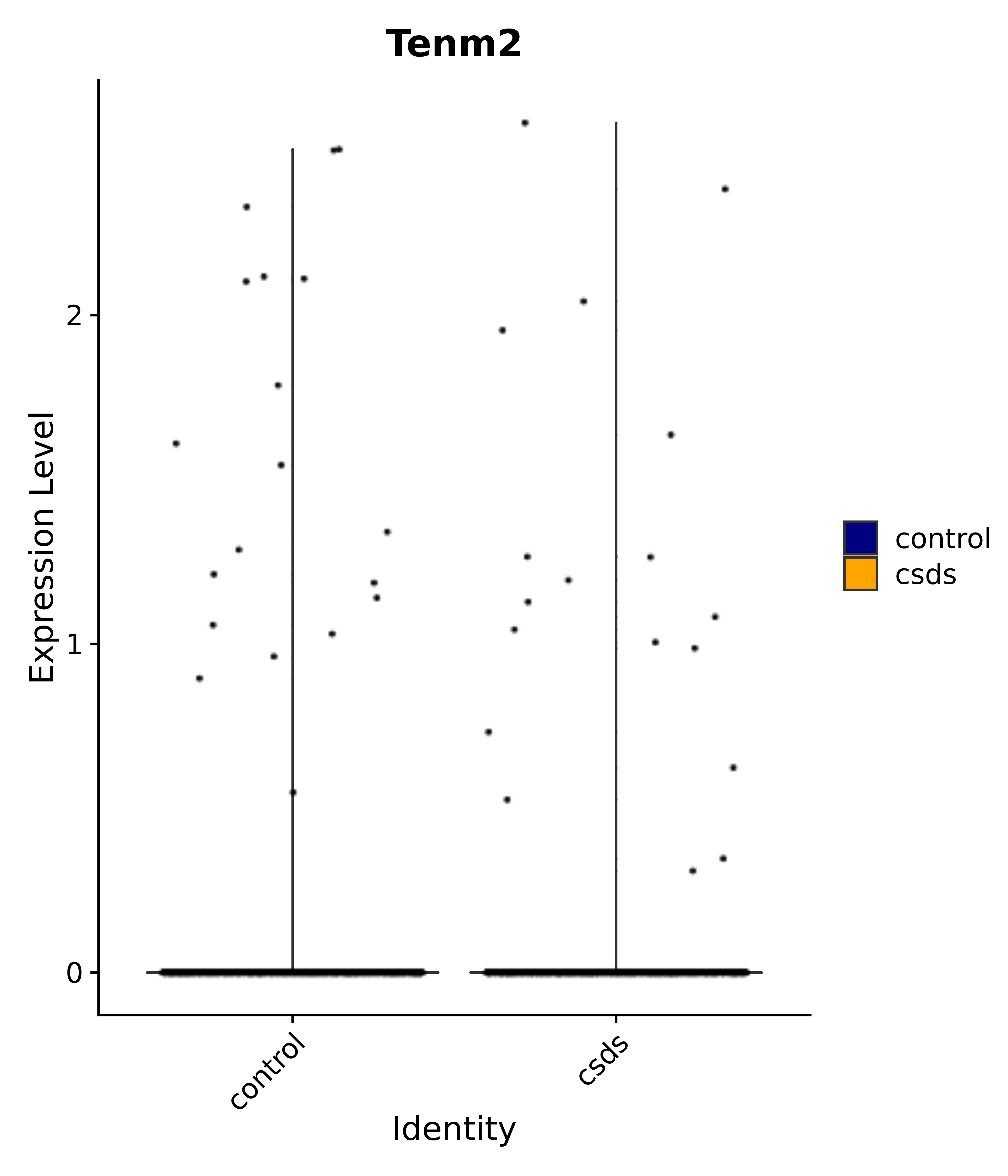
"Tenm2",
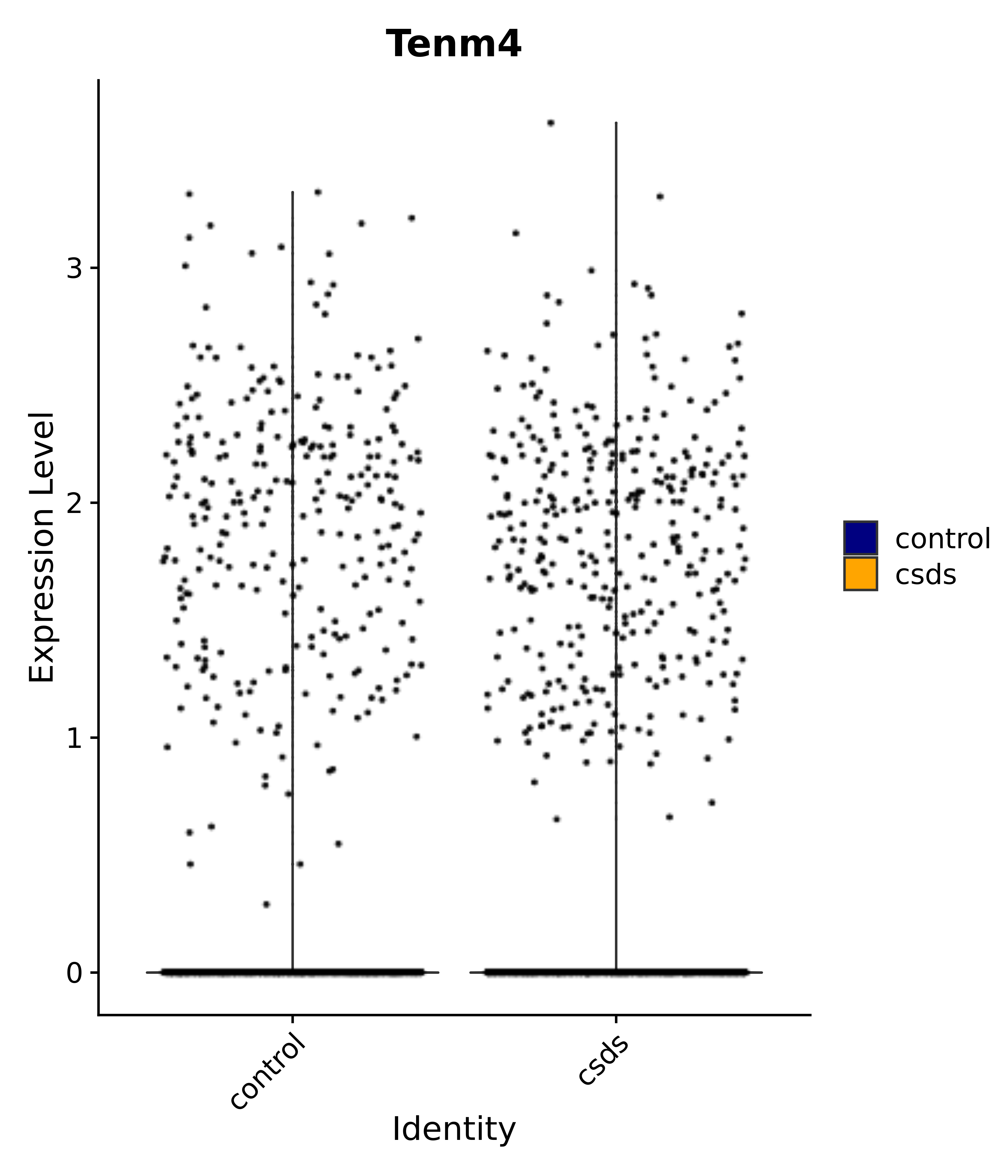
"Tenm4",
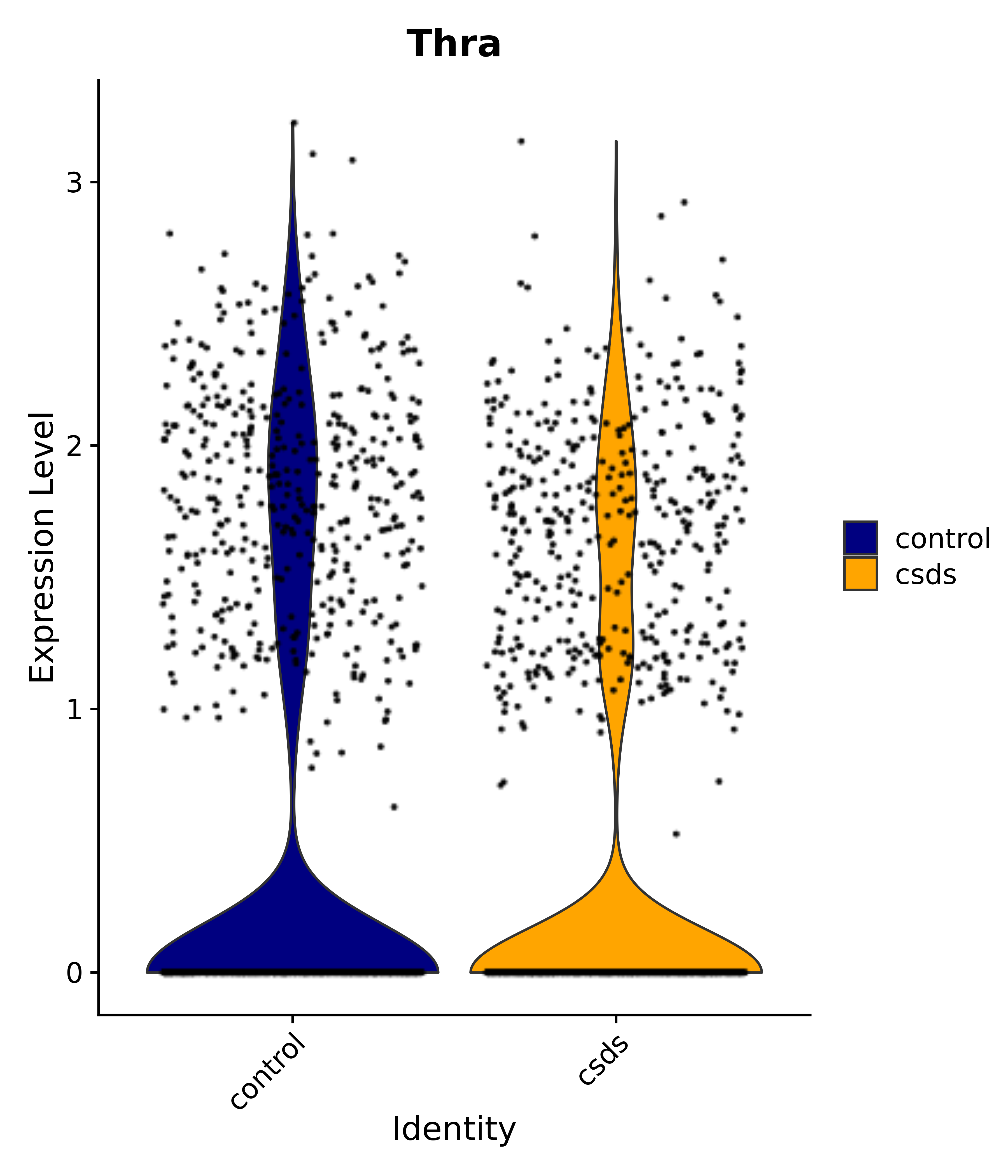
"Thra",
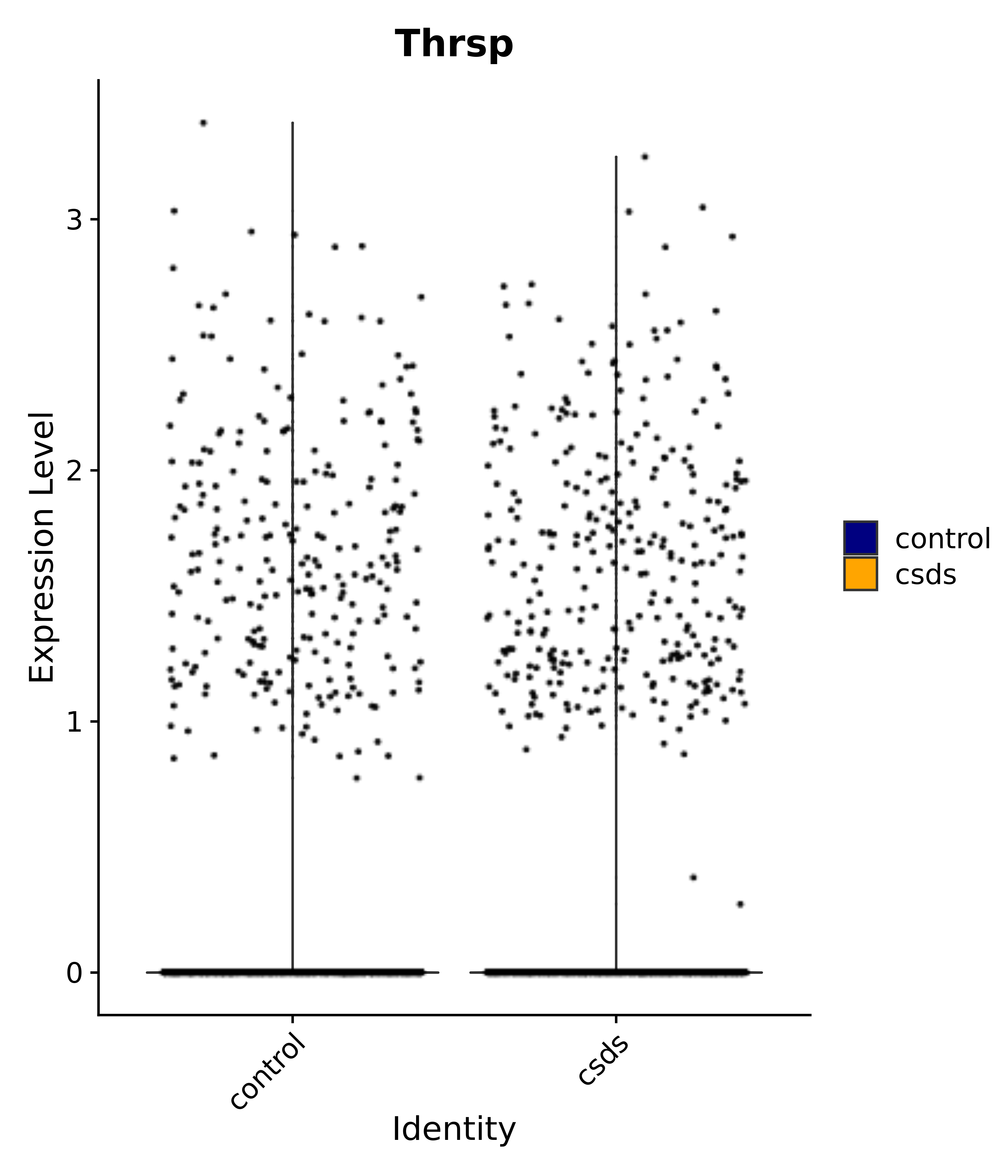
"Thrsp",
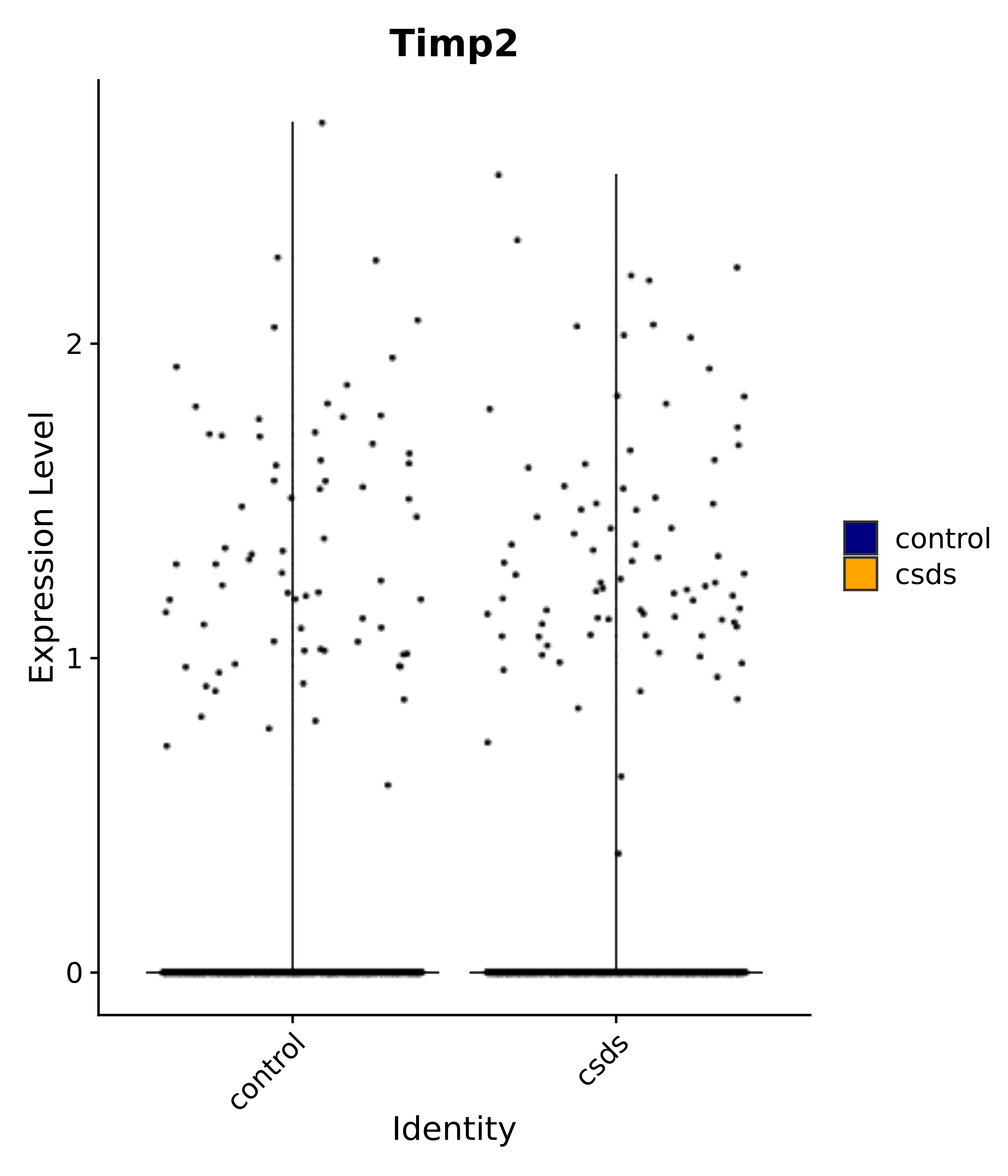
"Timp2",
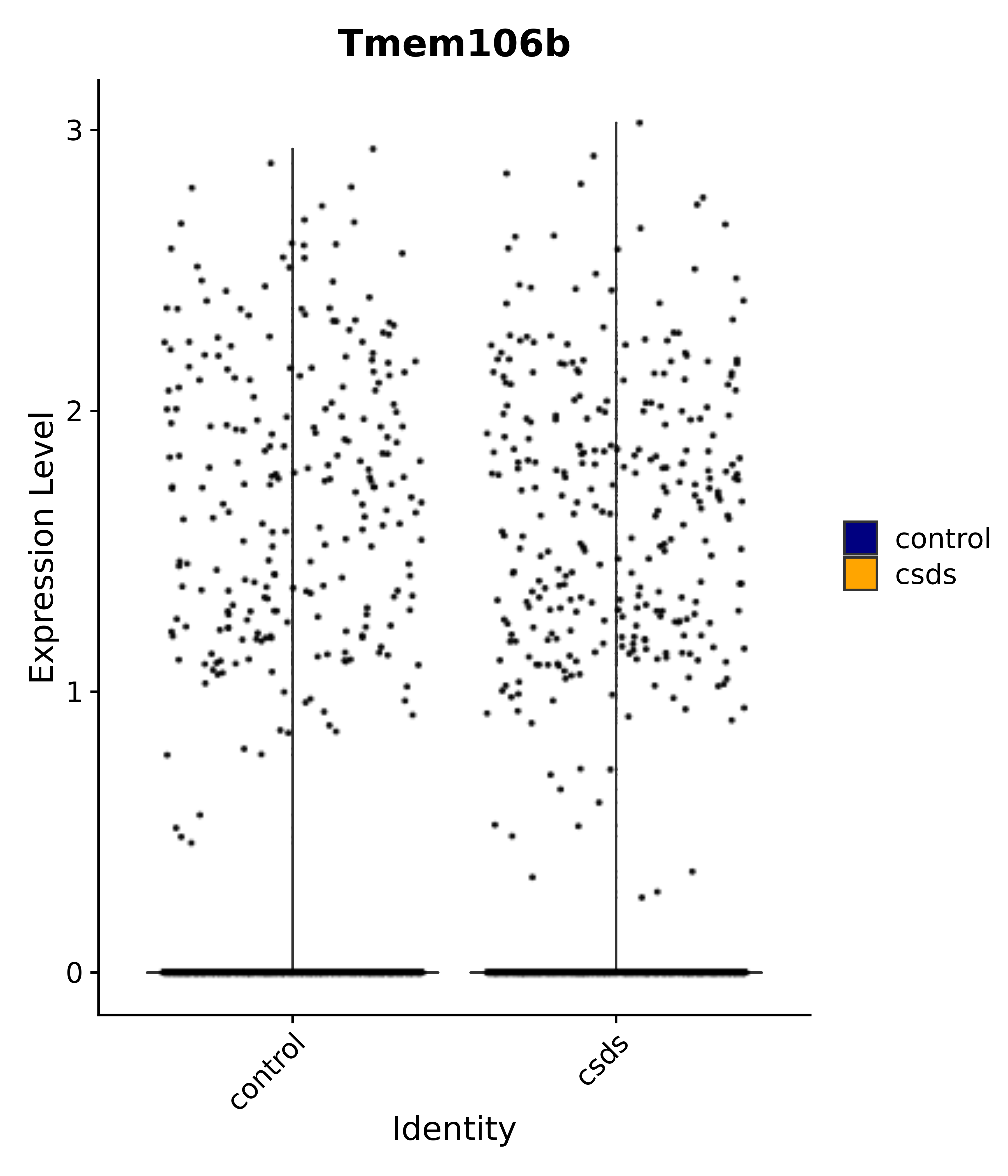
"Tmem106b",
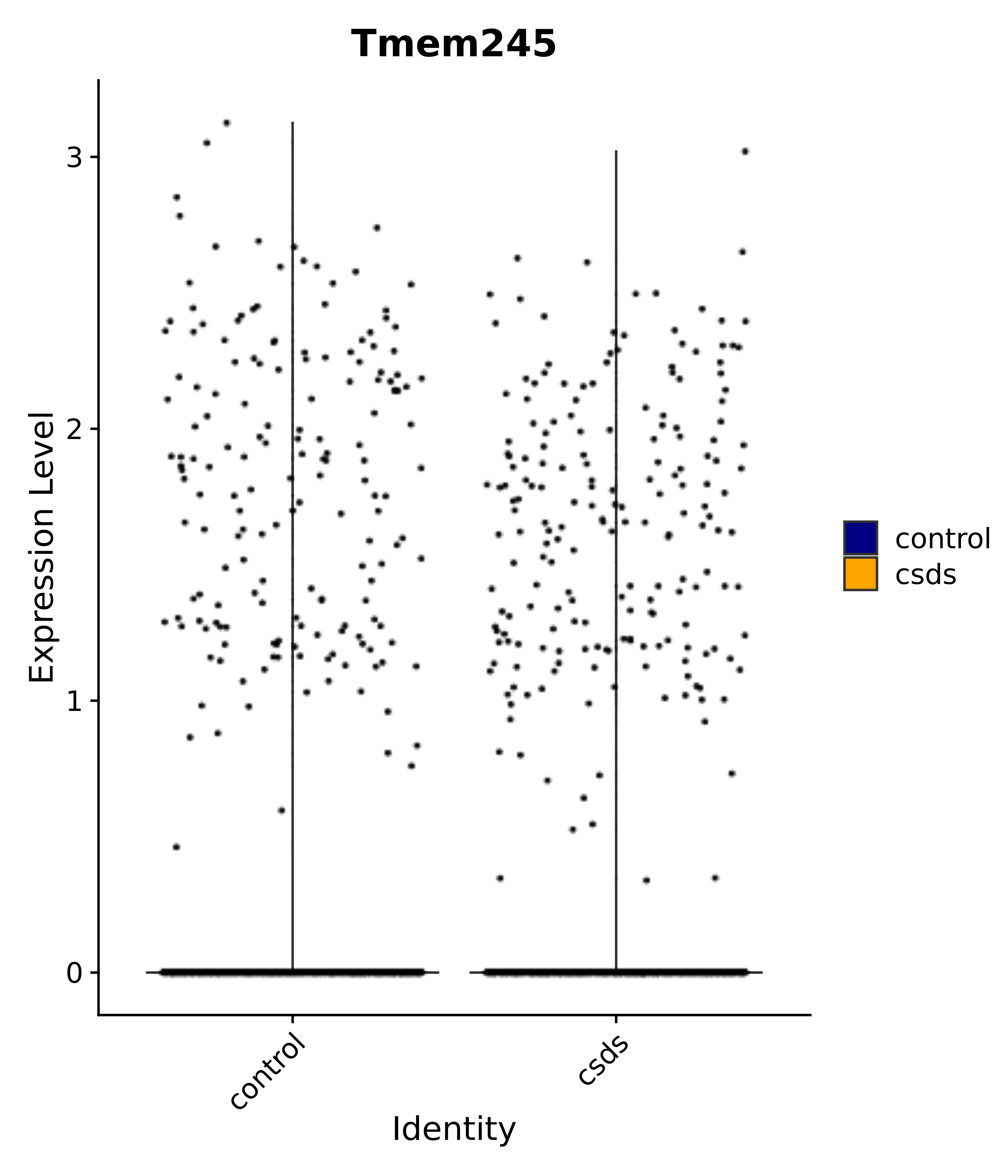
"Tmem245",
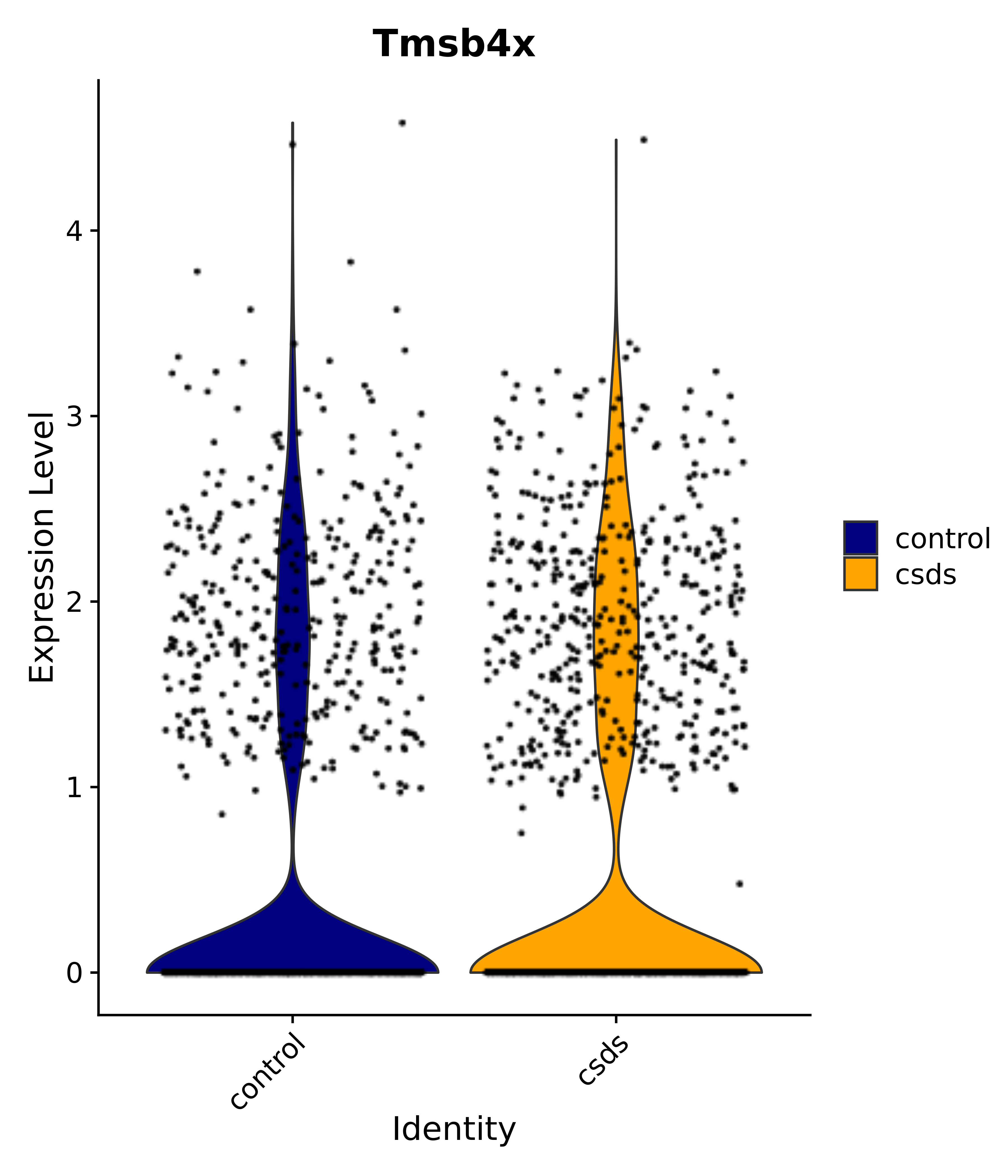
"Tmsb4x",
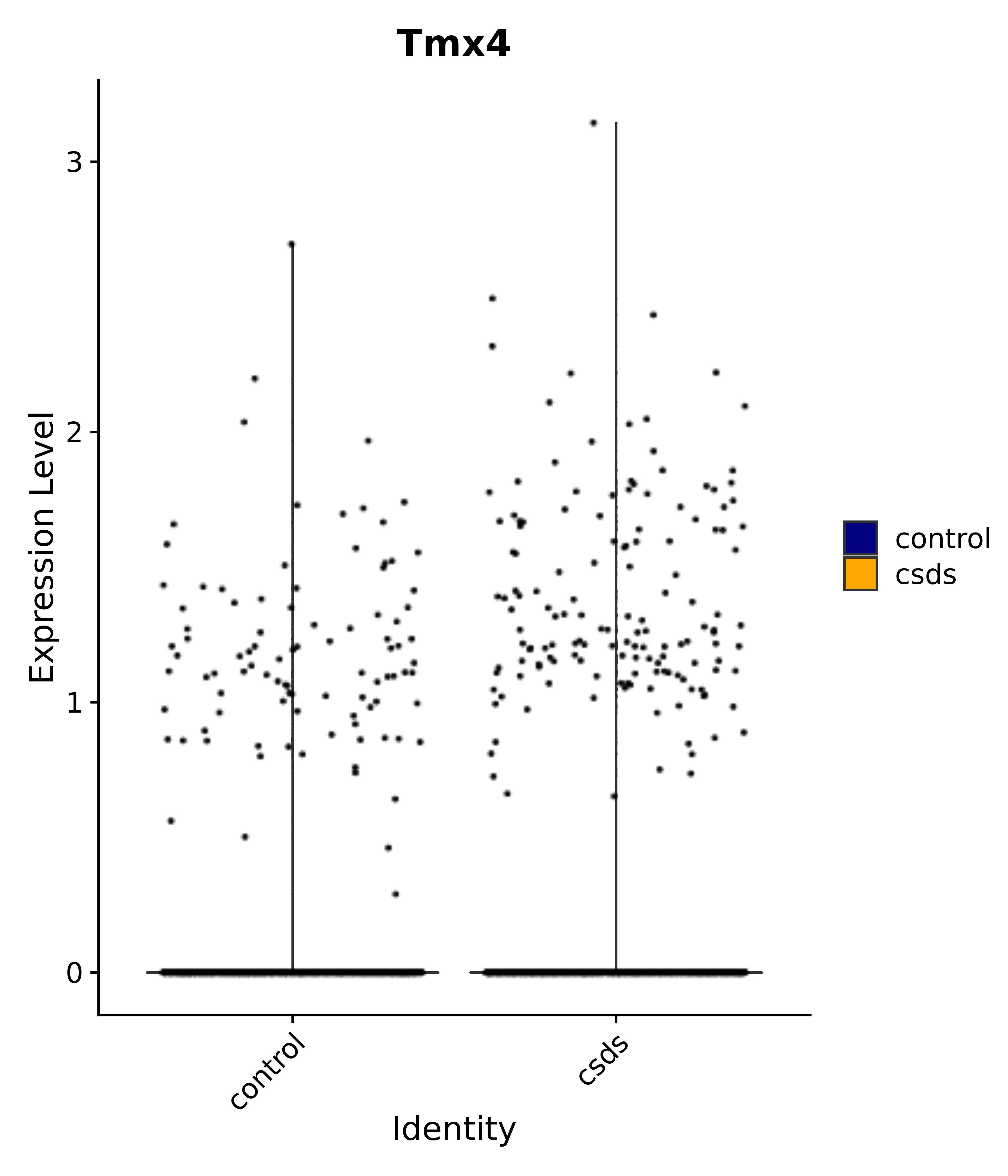
"Tmx4",
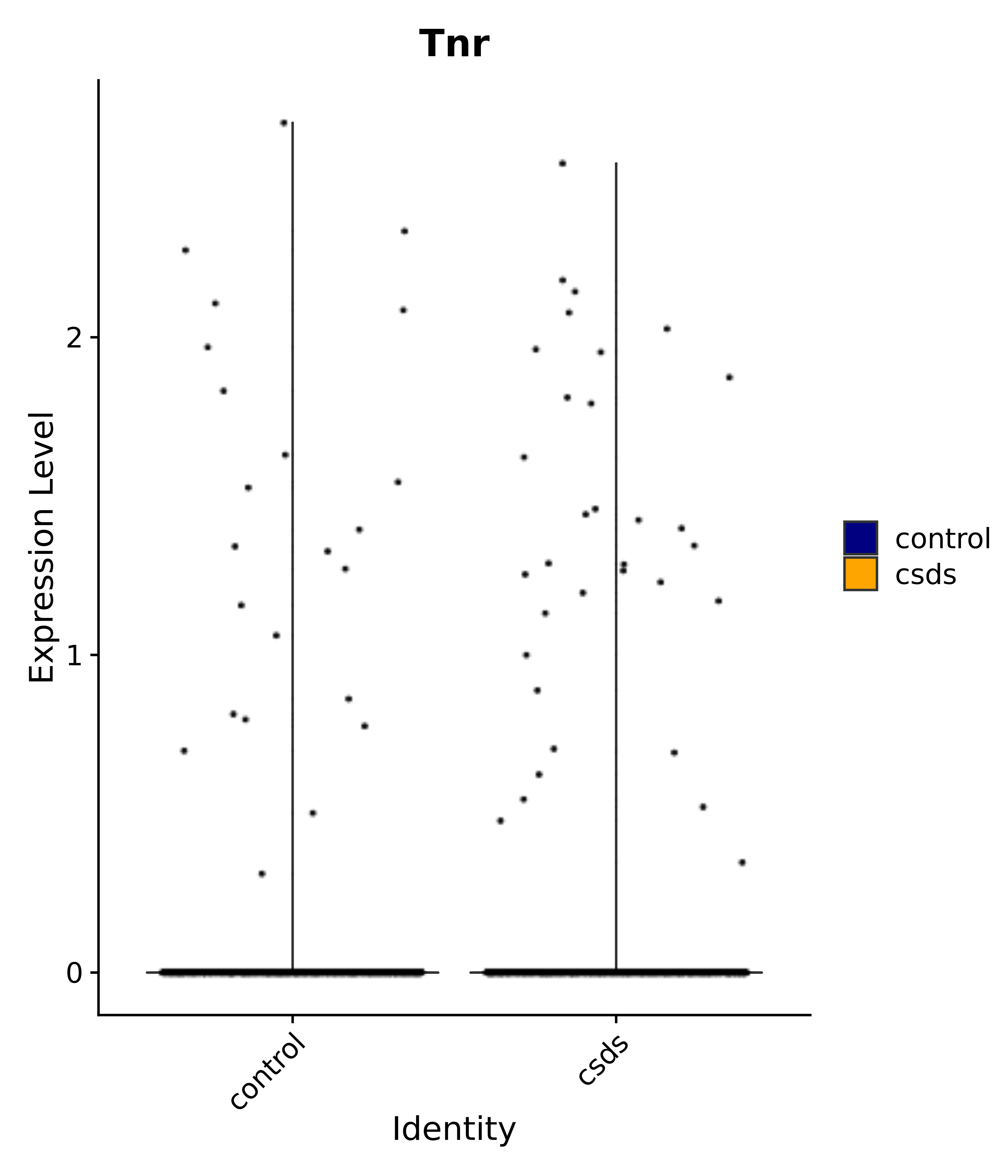
"Tnr",
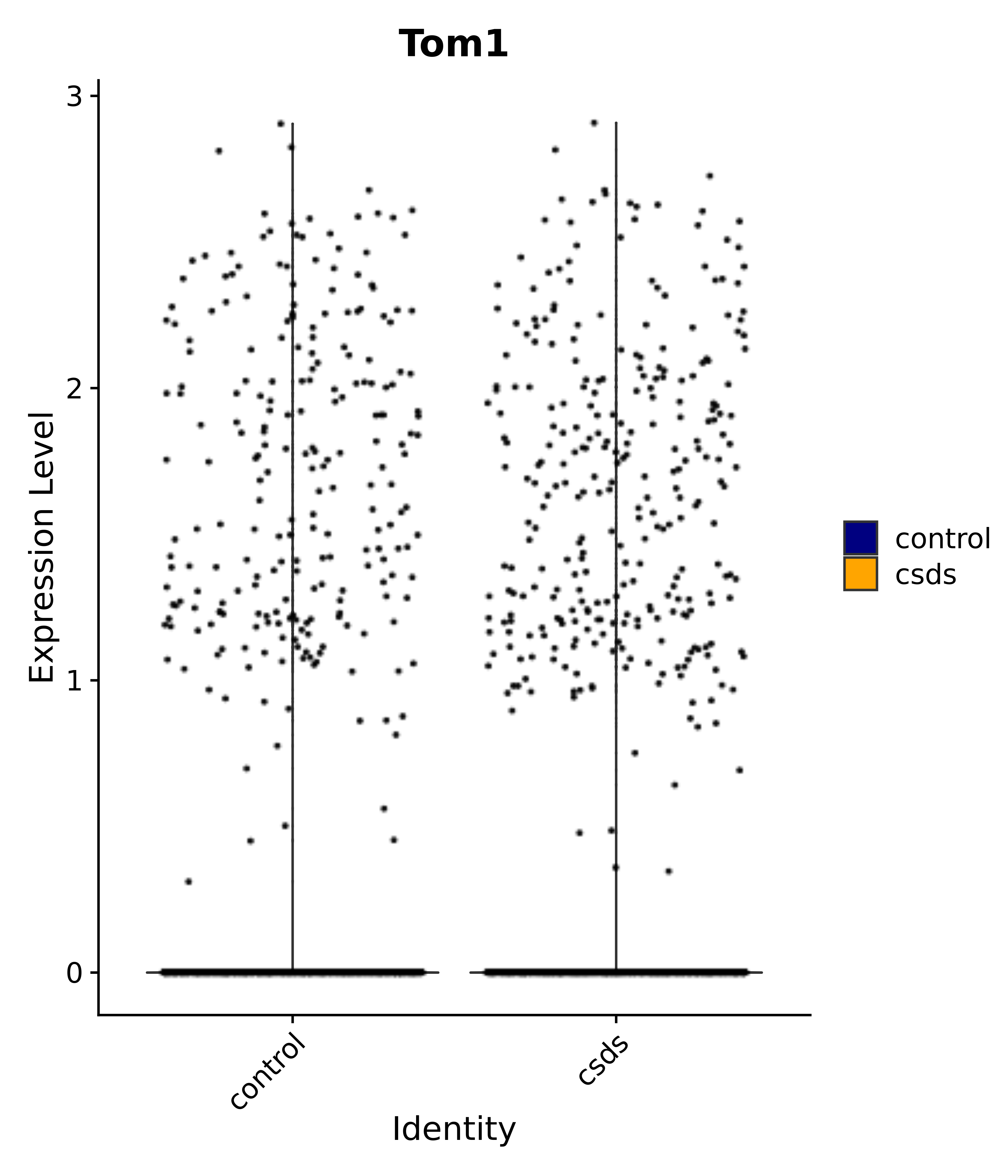
"Tom1",
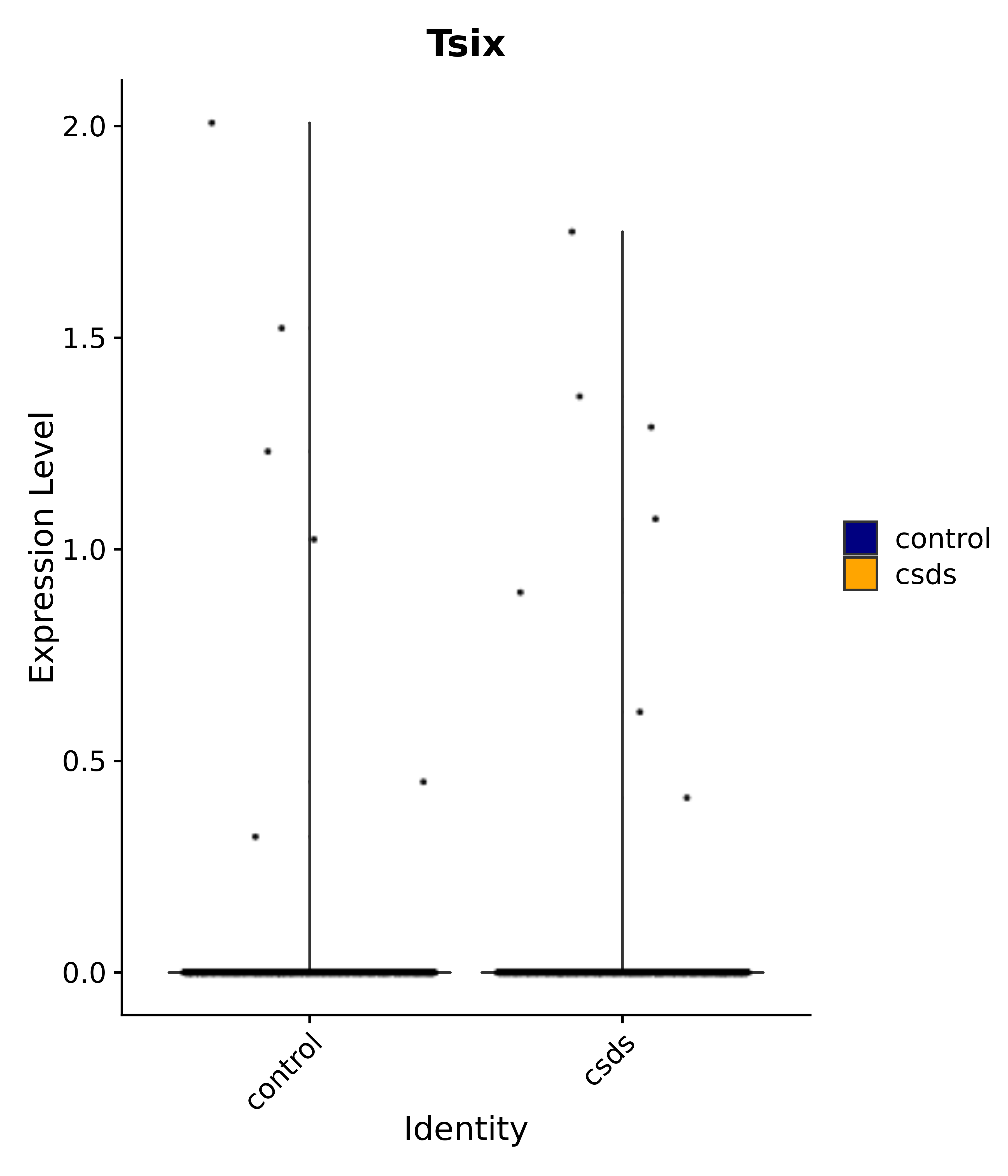
"Tsix",
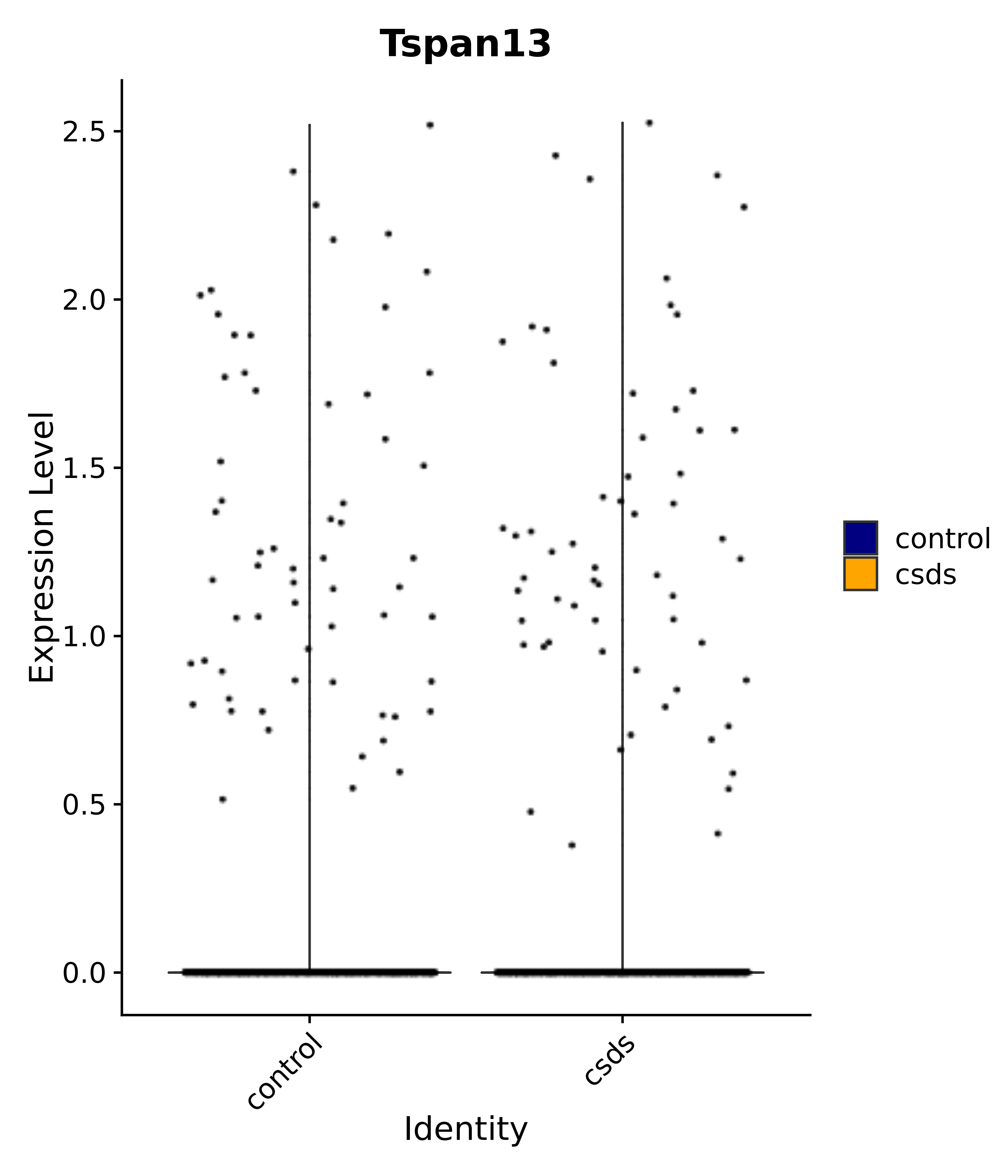
"Tspan13",
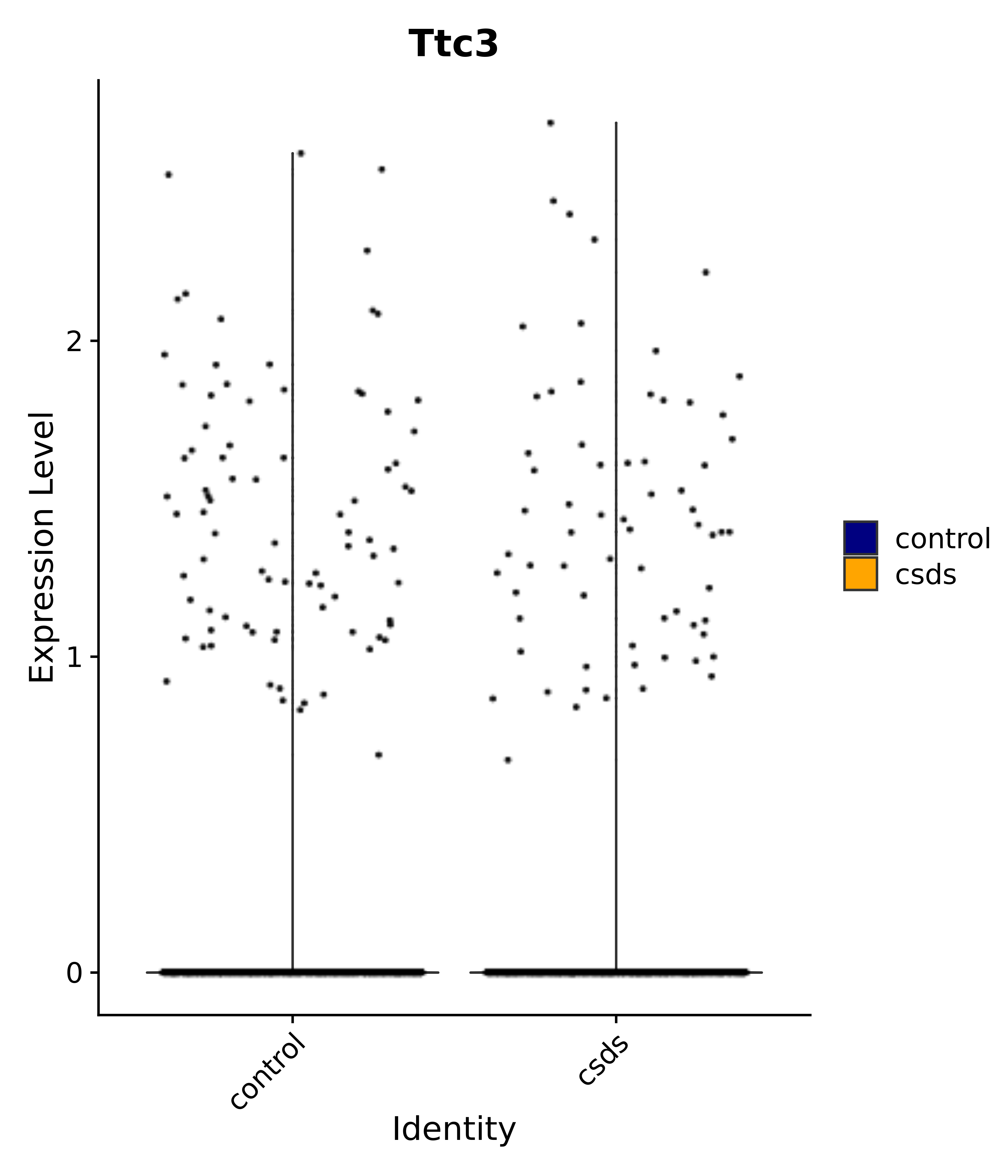
"Ttc3",
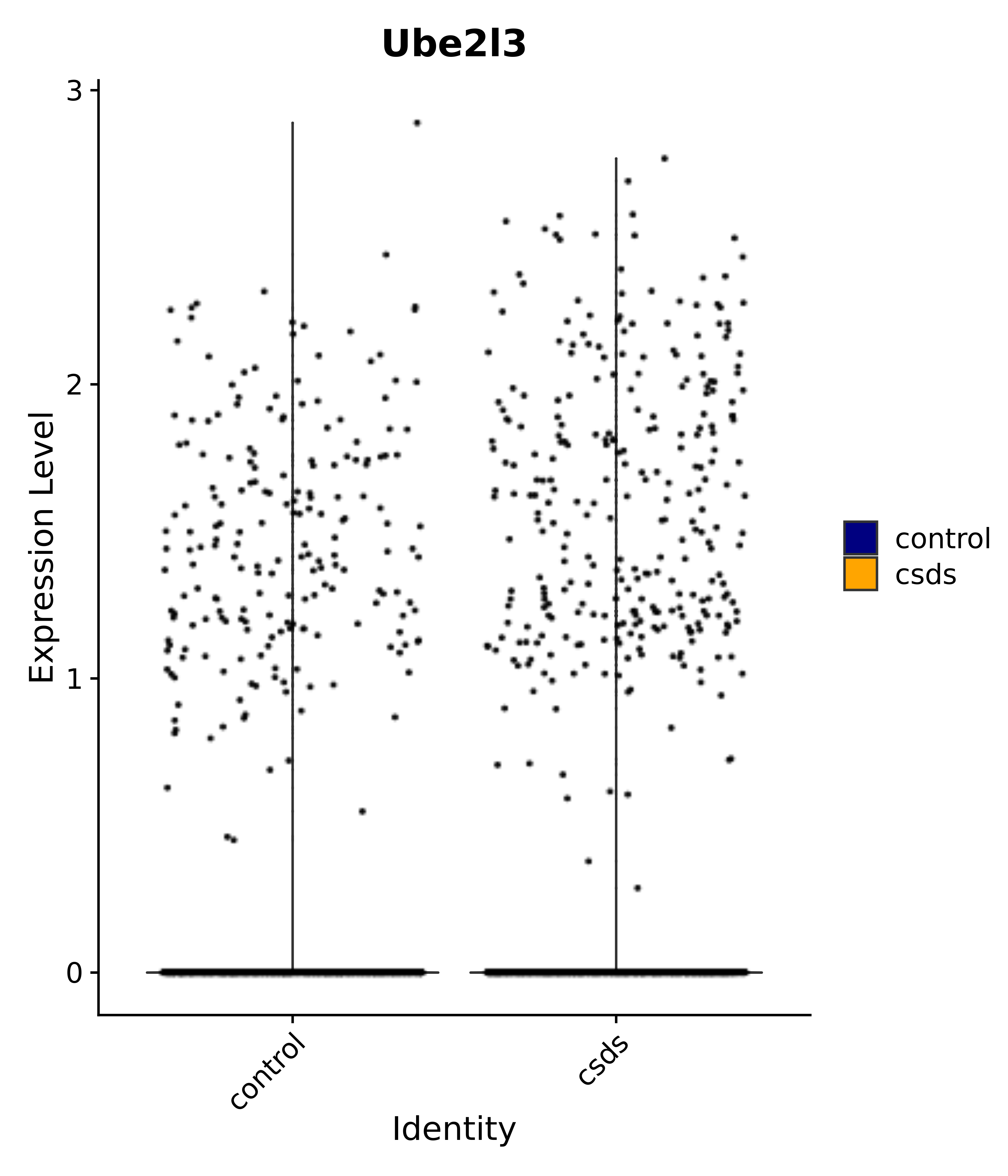
"Ube2l3",
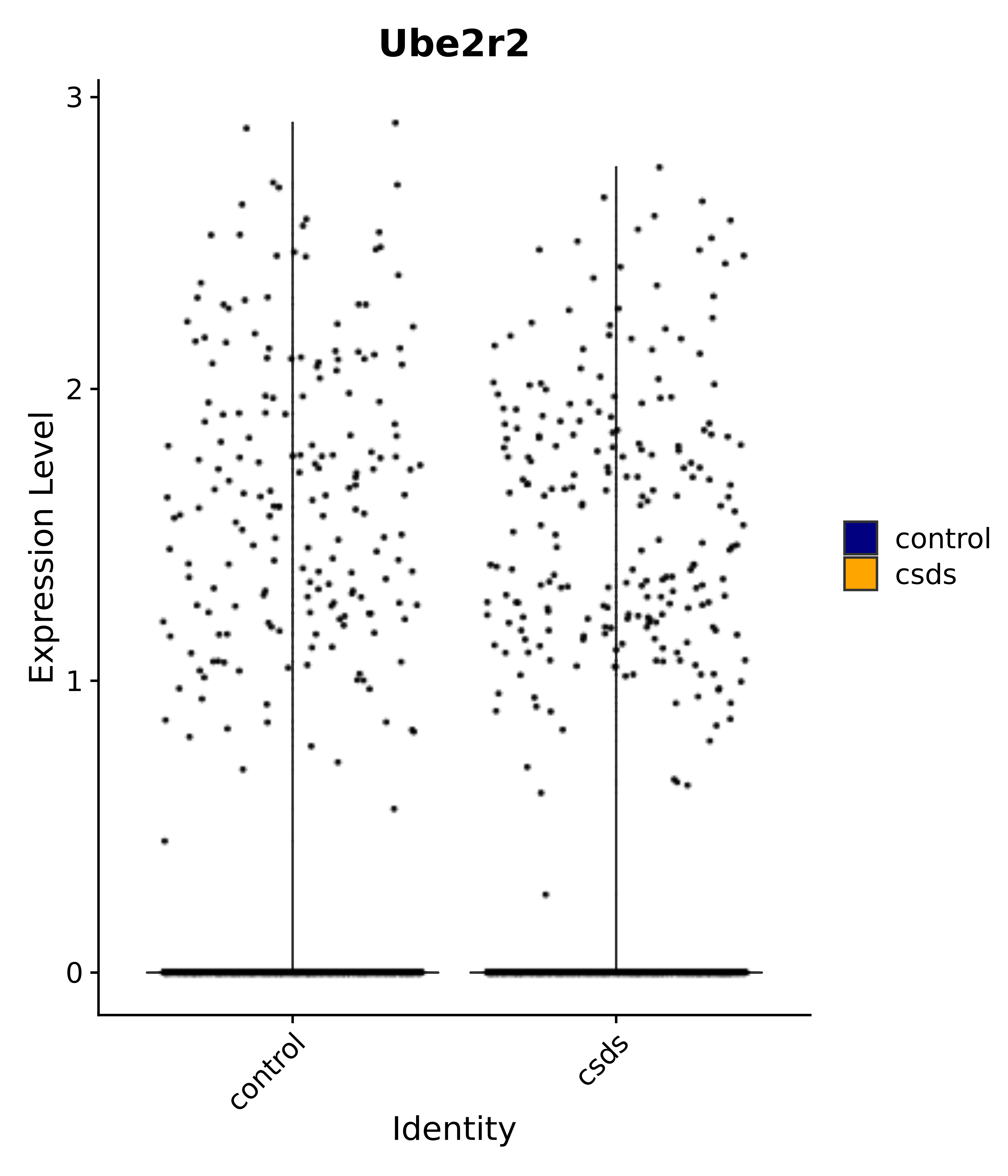
"Ube2r2",
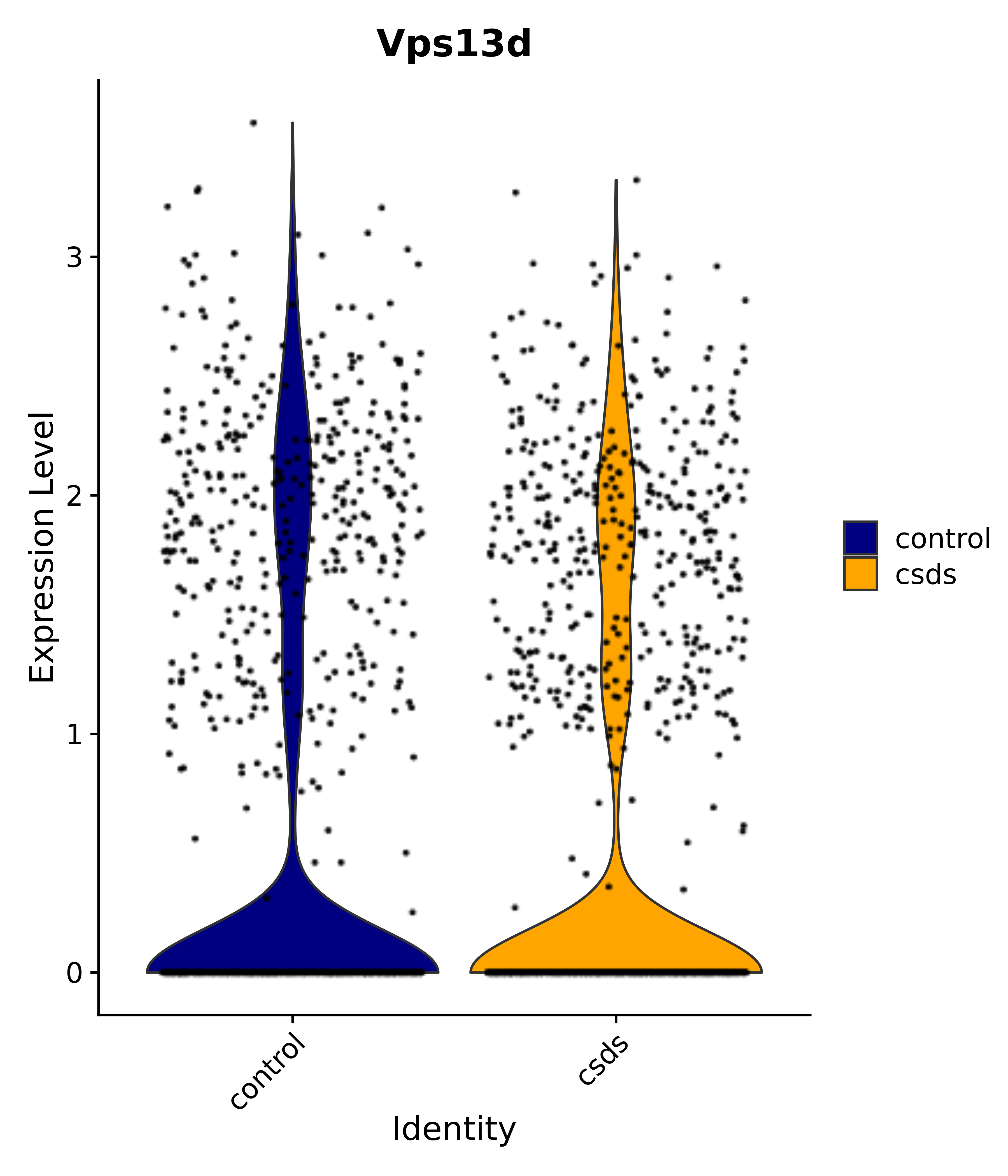
"Vps13d",
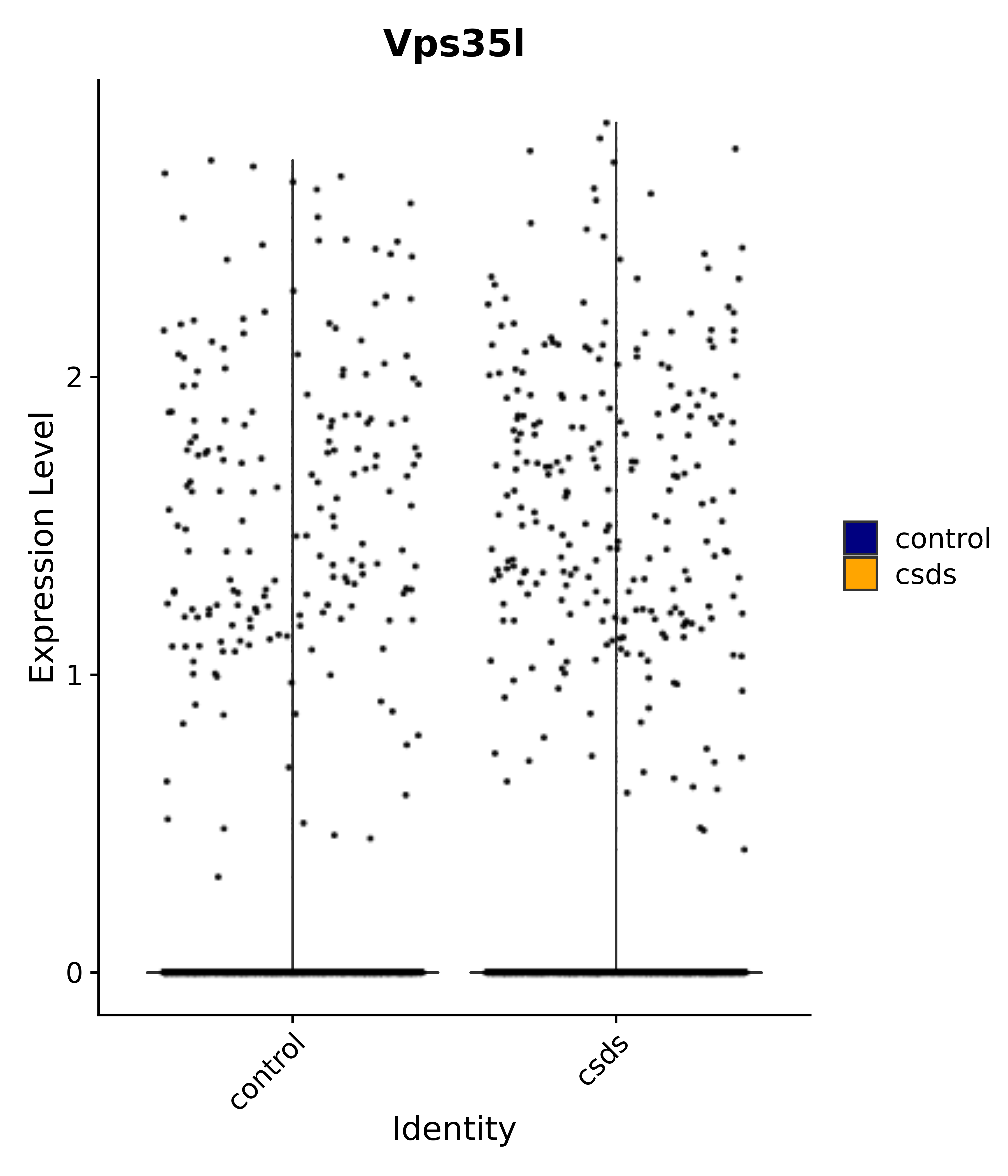
"Vps35l",
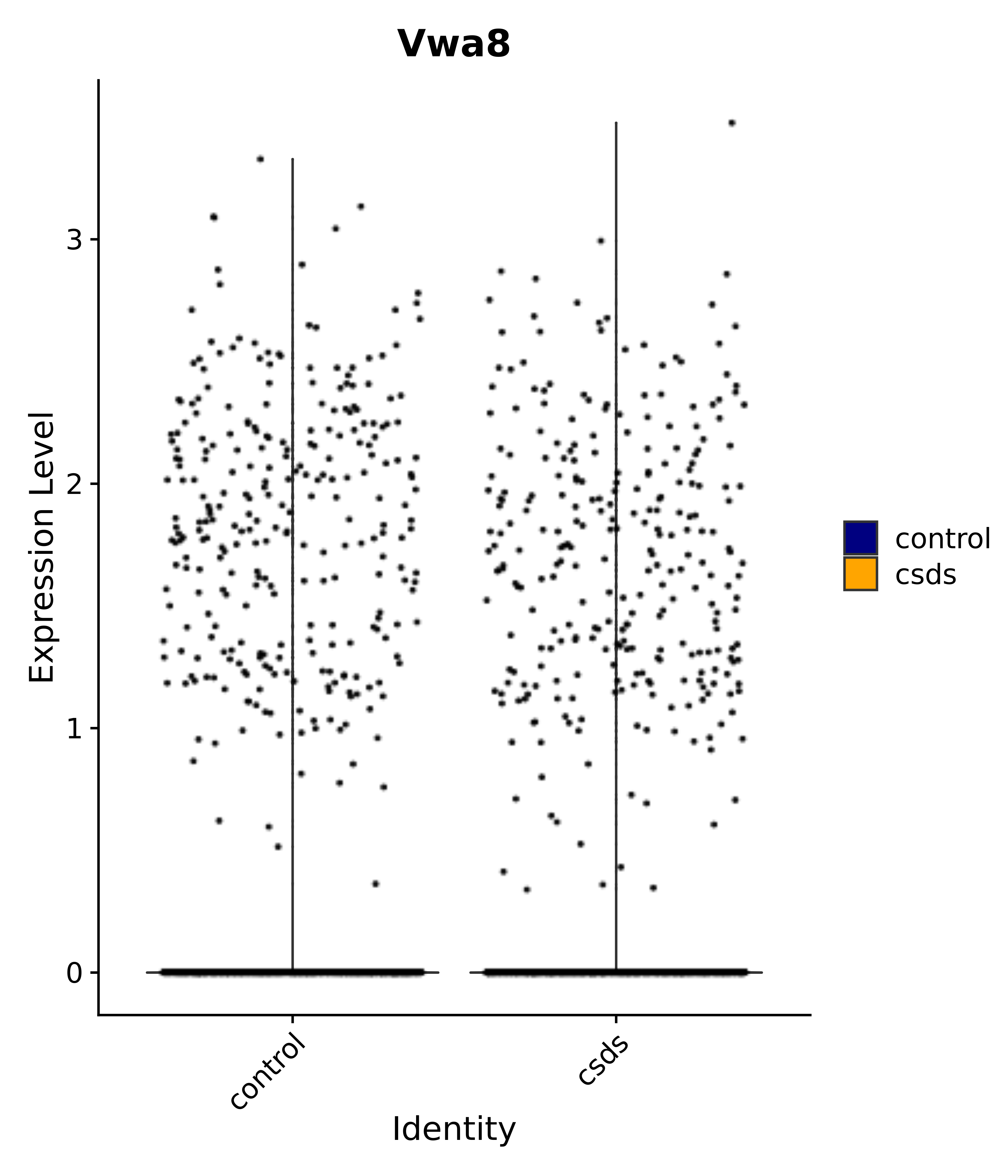
"Vwa8",
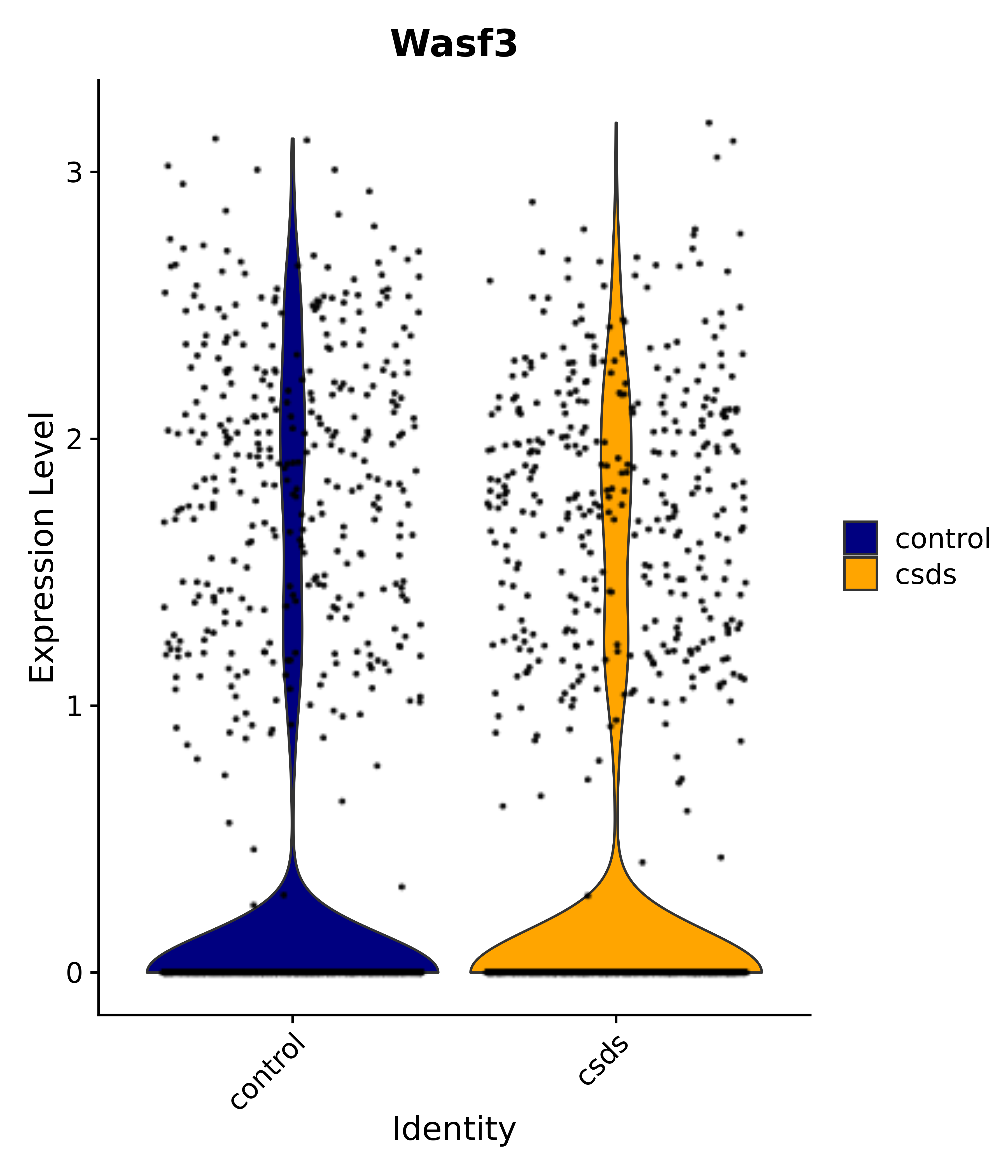
"Wasf3",
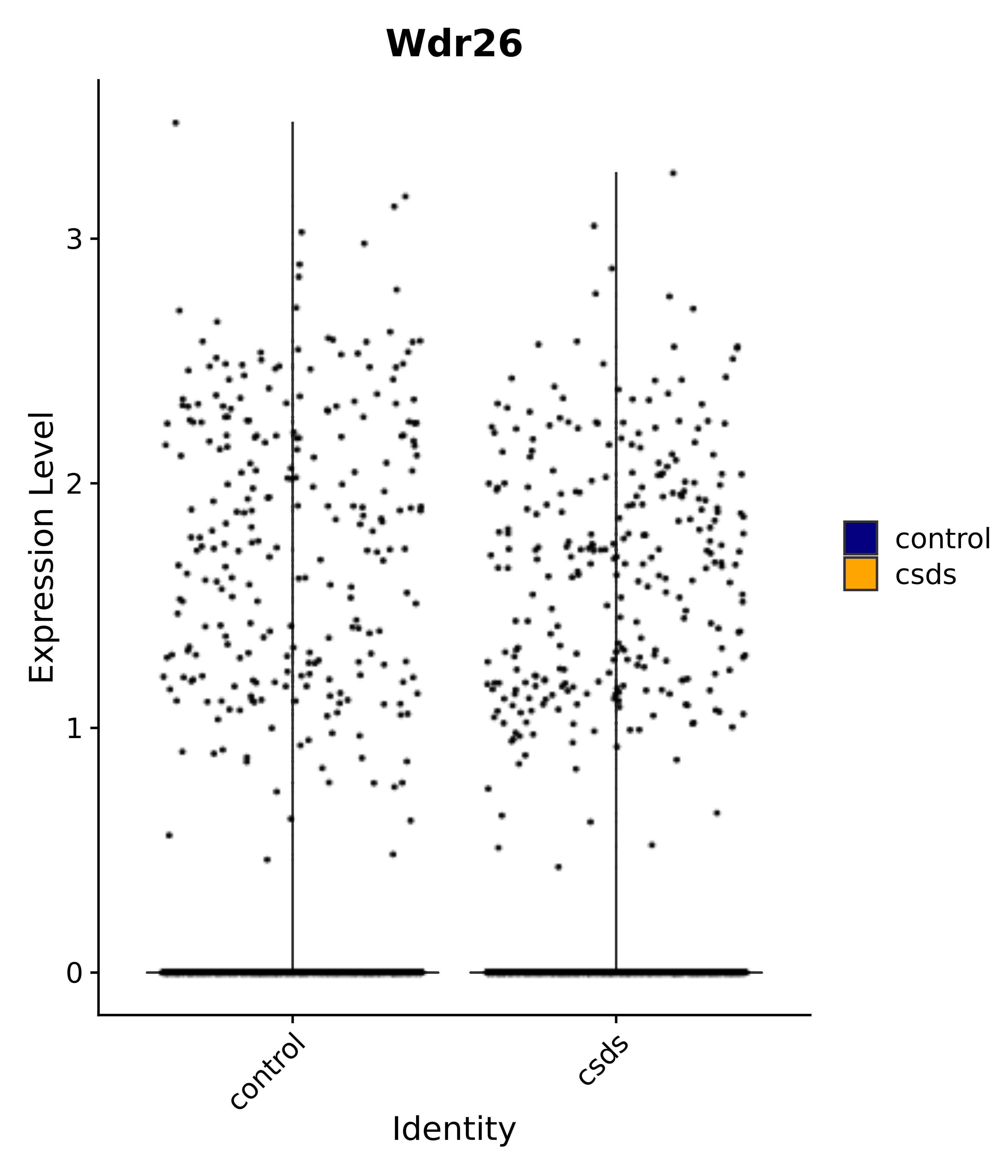
"Wdr26",
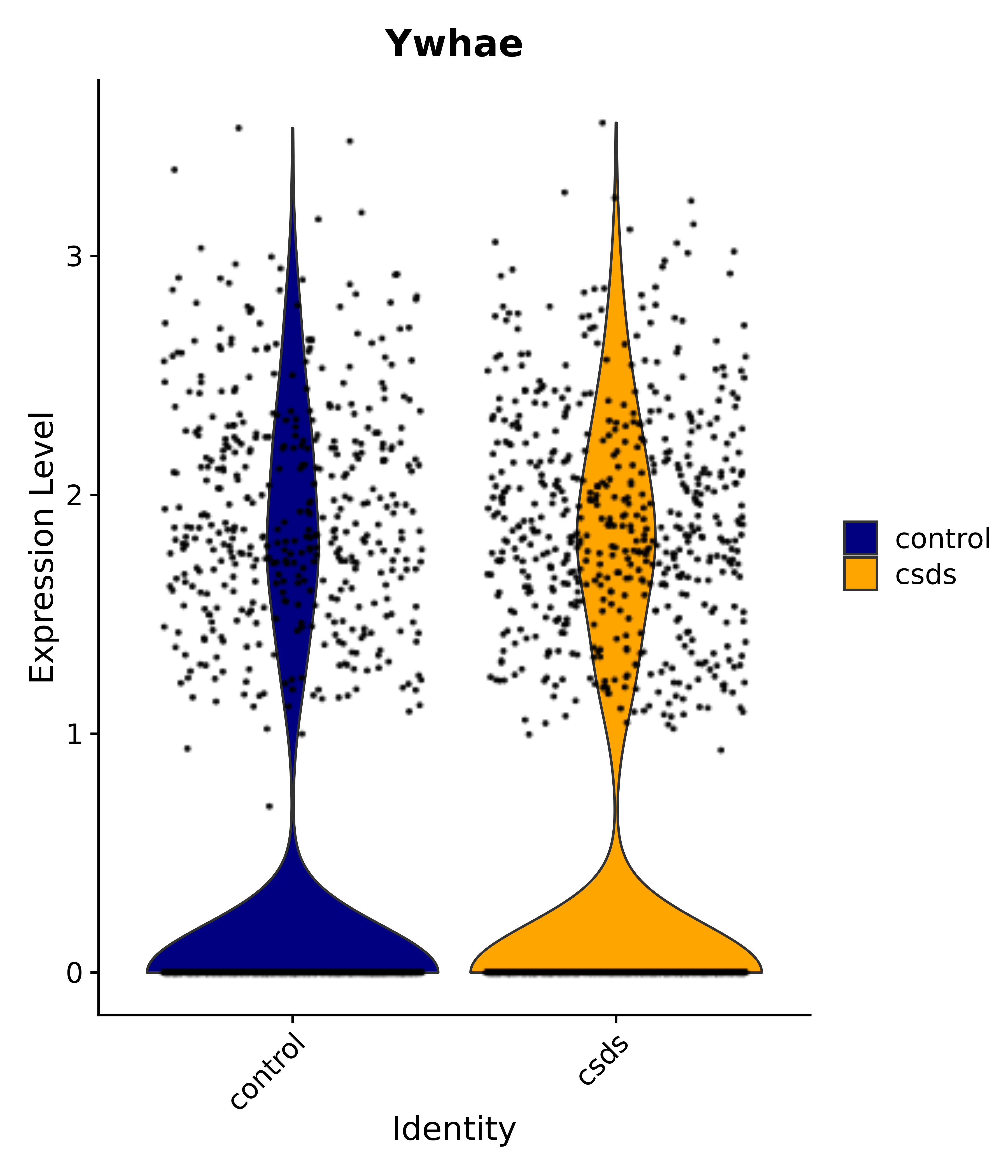
"Ywhae",
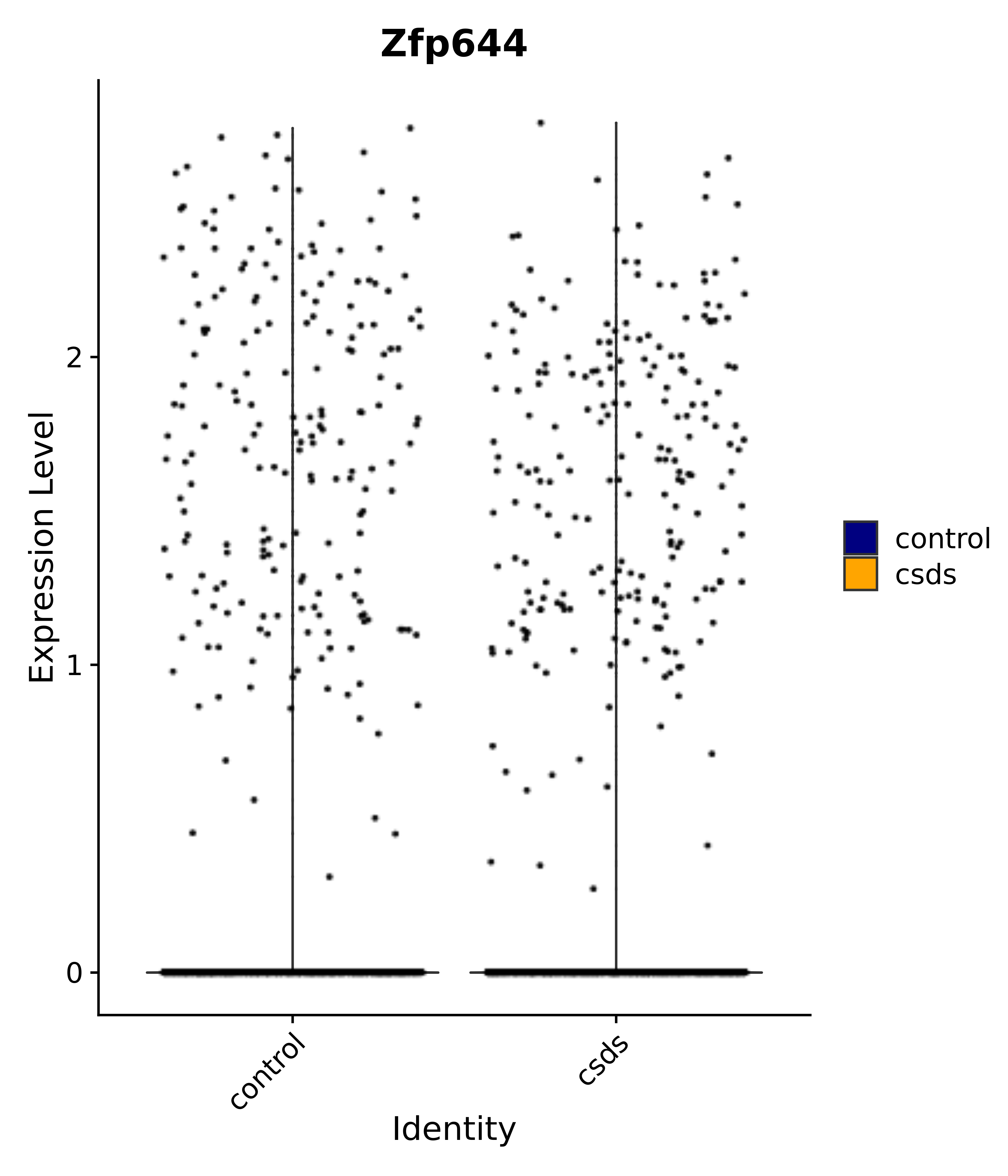
"Zfp644",
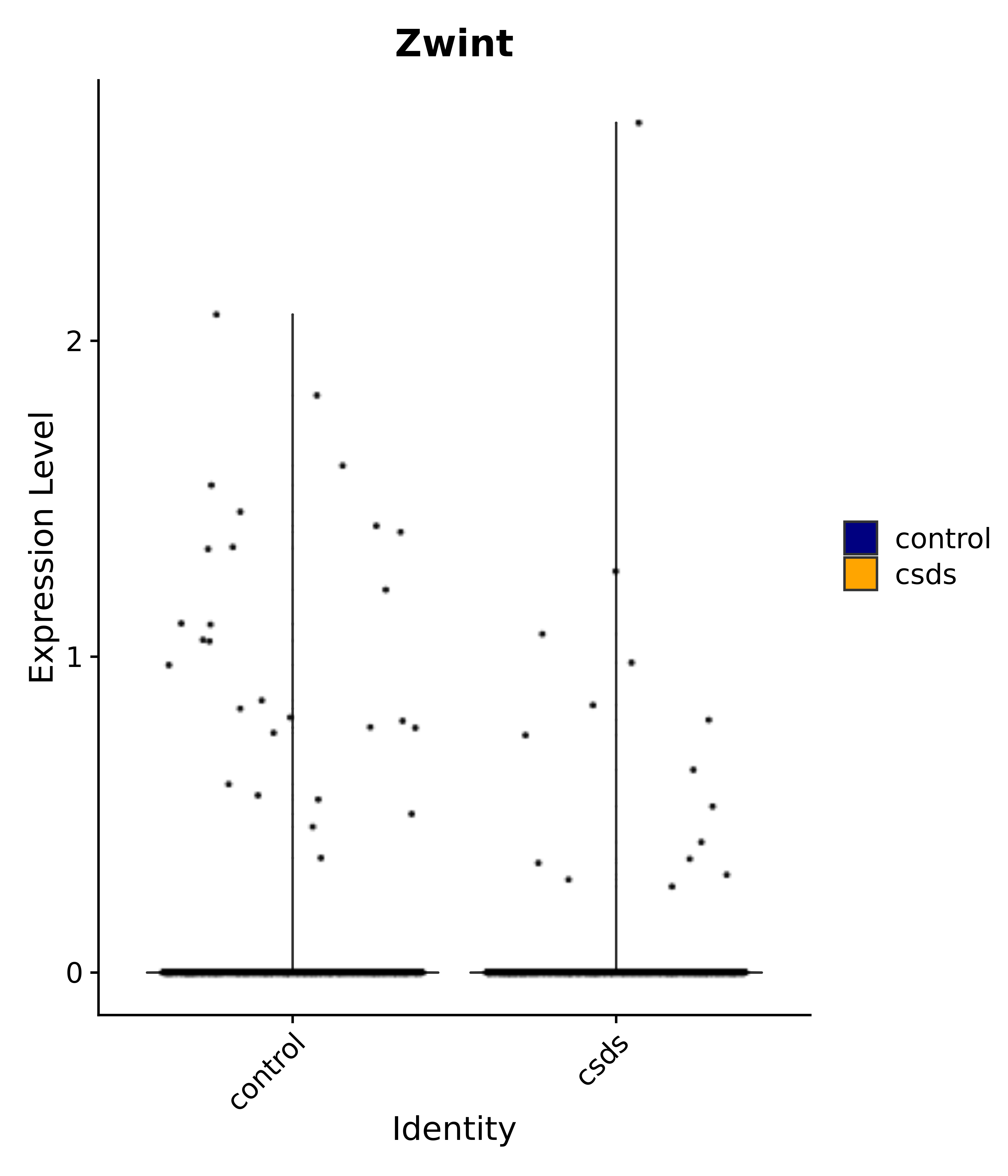
"Zwint"
)
selected_genes <- unique(c(selected_genes, at_top_50, at_top_50s, mitochondrial))
selected_genes <- selected_genes[selected_genes %in% rownames(GetAssayData(object = srt, slot = "data"))]